Lus10015385 [FLAX]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Poplar homologues |
|
||||||||||
| PFAM info | |||||||||||
|
Representative CDS sequence |
>Lus10015385 pacid=23160861 polypeptide=Lus10015385 locus=Lus10015385.g ID=Lus10015385.BGIv1.0 annot-version=v1.0
ATGAGCAATCCAACTGCTGAACCCAGCGGTTCTAGTTCAGGATTCTCCTTGCAAGGGATTGGACTCTCCAACACAATACATTCGGAAGTTGCGCCATGCT
TGCCGCTACCTTTCCTTCCGGTGTTCTGCGGCGCTTCCGATCCGGAAATCCGACTTTTCGATGAGCGCAGTGGTGGTGGTGAAAGTCTTTCGTATTTGGA
TCGAAGTGAAATTGTAGCTCAGTCTGCTAGAATTGCTGAACTGCTTCGAGAAACTGATGTTTCGTACATTAATCTTCGAAACAATGCAAGGCCTATGACT
CTGGACTATGTGATACCTTTGGAACTTTGTGACCATGTTCTCCAGTGCAATCCTGAAGCTTTTGAGCACTCCACTCCCGGTAAGGCGCAACTTTCTGGAA
ATATAGCCTCTGAAAGCAGGCATGTTCGTGTTGAGACCAGTATACCTGTTCTGCTTAAGGTGCAAAATGACGATGTCAATCTAATGCGAAAGACGTTGCA
GGATATTGGTTCATCCTCTAGGAAACCAAAAGCCAAGAAAAAAGGTGTCGAAGATATCTCGGCACCGGTCCAGCTAGAACCGACTGACCTTCCTGGGCTT
CAGGATACCATTATTAGGAACTTCTGTGATGCATTGGAGGATTTCTGTGTCAAAGCTGAAGTTCTAAGTGATGAACATGAAGAAGTAGAGTGGTTGGTAT
TACCTGCCAGTGATGTTAGGTTGCTTGTCAATGAAATCATGTTGATTCGTGGAAAGAAACTTCTACATTTGGTTCCTCTTGAGGCACTTGTAAGATTGTT
ACGCATTCTAGATCATCAGATACATCGAGCAGAAGGCTTATCAGTTGATCAGTATGACCACTCAGACTCGGATGTCGTTTCAAACGTCTTTTCTGCCTTG
GAGTCTATTCATGCGGCTCTAGCTGTCATGGCTCACAATAACATGCCTAAACAGCTCTACAAAGAAGAGATCATTGAAAGAATTCTGGAGTTTTCTAAGC
ATCAGATGGTGGATATTCTGTCAGCTTATGATCCGTCATTTCGGGCTTTGCACAAGCCAAATGAACGCGAGACATATGAAGATGATGAAAACGAAGACCC
TCAGGCTGATTACGCTCCATCAGGCAAGAGATGGCATGTTCAAAAGAGTGTAAAAATGAAGAAATCAGCTTATAATAAGGTCACCAGTTCCATGAATGCA
ATTCTACAGAAACTGTGCACTATAATTGGCTTACTGAAGGAACTGTTATTAATAGAGAGGTTATCTGATAGTTGCATTTTACAATTAGTGAAGACAAGCT
TCACGACCCTTTTGGTTGATAATGTTCAACTCCTTCAGCTTAAGGCGATTGGTCTACTTGTTGGGATATTCTATTCTTACACACAACATCGAGTATATAT
CGTGGATGAGATGGTTCAGTTACTATGGAAGCTACCAGTCTCAAAACGTGCACTACGAGCCTATCACCTGCCTGATGAAGACCAAAAGCAGATTCAGATA
ATACCTGCCTTGCTAATTCAATTGGTACACAGTAGTGCAAACCTTCCTGAAGCGTTAAGGCAAGCATCAAGTGGTCATTCTATCTTGGAAGTCTCATTGG
ATGCAACCTATATCACTAAATGCCATGAAGCTGTAACAGACACATGCTGCCTTTTCTGGACCCGTGTACTTCAGCGGTTTACAACAGTGAAAGCACAGGA
TTCTTCTGAAATGAAGGTGATAATGGAAAACCTTGTTACAGACTTGCTGACAACTCTGAACTTGCCCGAGTACCCTGCTTCAGCTATAATTCTGGAGATC
CTCTGTGTCTTTCTACTGGAAAATGCTGGGCTCAAATCCAAGGATGTTGCAGTGCGTTCTATGGCAATTGATCTTCTTGGGATGATTGCTACTAGTTTGA
AACATAGTGCCGCCATTTGTAGCAGGAATACTTTCTGGATGTTACAGGCATTAGACGGTGGAGATGATGCTAATTGTAGTCTACAAACAGATGCATGTTG
TATTTGCTCTGATGGAAGAGTTGAGAAAGCACTGTTCGTATGTCAAGGTTGTAACAGATTGTTTCATGCTGATTGCATGGGTGCAAGAGAACGTGAAGTC
ACTCGGAACTGGTATTGTCAGATTTGTCTATGCAAGAAGCAGCTTCTTGTGTTGCAGTCATTCTCCAAGATGCAGTGCAAGGATGAAGGAAGAGGAAACA
AATCGCGATCTAGCAAATTCCCTGAAAATTCTGATCTGATTGTGCCAAAAGTTGAAATTATCCAACAGACACTCTTAAATTACCTCCAAGATTCTGTTTC
TGCTGATGATATGCATCACTTCATCTGCTGGTTCTATCTTTGCTTATGGTACAAGGACGACCCAAAATCTGAGCAGAGATTAACTTATGACCTCATGAGA
TTAAAATCAAATCTGATCGTGCGTGATTCTGGCACTAGCCATTCCACATTGACAAGGGAGTCGTTTAATAAGATTACAAGAGCATTGGGACAAAATAGTT
CTTTTGCTCGAGGTTTCAACAAAATTCTTCATCTGCTTCTGGCAAGTTTAAGGGAGAATTCCCCAGTAATCAGGGCCAAGGCATTGCGAGCTGTAAGCAT
AATAGTAGAAACTGATCCTGGGGTCCTGCGTGACAAAGGGGTTCAATTTGCAGTTGAAGGGAGATTTTGCGATTCTGCAATATCAGTAAGGGAGGCCGCC
TTGGAACTGGTAGGAAGGCATATTGCTTCGCATCCGGATGTAGGGCTACAGTATTTTGAGAAGGTTGCTGAGAGGATCAAAGACACTGGTGTGAGTGTCC
GGAAGAGGGCCATTAAAATTATCCGAGACATGTGTACTTCAAATGCCAACTTCTCTCAATTTACAACTGCTTGCATTGAGATAATCTCTCGTGTGAGTGA
TGACGAATCAAGTATTCAGGACCTTGTCTGCAAGACATTTTACGAGTTCTGGTTCGAGGAACCTTCTTGTGTGCAGACAACTTTTTCTGGGGATGGCAGT
TCTGTTCCTTTAGAAGTGGCTAAGAAGACCGACCAGCTTGTTGAAATGCTAAGGAGAATGTCAAGTCATAGCCTTCTCATTACTGTCATTAAACGCAACC
TCGCCCTTGACTTCTTTCCACAGTCTGCTAAAGCTGTTGGGATCAATCCCATGTCACTTGCATCTGTGAGGAAACGCTGTGAGCTAATGTGCAAGTGCAT
ACTCGAAAGGATATTGCAGGTGGAAGAAATGAACAACACTGAAGAGGAGGTGCAAACTCTGCCTTATGTGCTGGCATTGCATTCATTTTGTGTGGTGGAT
CCAACACTTTGTGTTCCCTCTTCAGACCCATCTCAGTTTGTTGTCACTCTGCAGCCTTATCTTAAAACTCAGGTTGATAACCGAGTTGTTGCAAACTTTC
TGGAGAATATAATCTTCATAATTGATTCAGCACTTCCCCTGATCCGAAAATTGACACCAGTTCTCGTTGAAGAACTTCAGCAAGACCTTAAACAAATGAT
CTTACGGCACTCCTTTTTTACCGTCGTGCATGCTTGCATCAAGTGTCTTTGCACGTTGAGTAGAGTTGGTGGAAAAGGGGCTAGTGTAGTTGAGTTCCTT
ATACAGATGTTCTTCAAACGTCTAGATGCTCAAACTCAAACATTTCAAAATAAACAGTTGGTAGTTCGTTCTCTGTTTTGCCTTGGGTTGCTTATTCGAT
ATGGAAGTCCTCTGGTCAGTGAGTTGGGTAACAAAAATATTGATATTGCCAGTAGTCTCAGCTTGTTTAAGAAATACCTCATAGTGGACGACTTCAGTGT
CAAAGTCAGATCTCTGCAGGCCTTAGGCTTTGTCCTAATTGCCAGACCCGAATATATGCTAGAGAAAGATATTGGAAAAATATTGGAAGCAACTTTATCA
CCGAGTTCTCATGTTCATATGAAGATGCAAGCACTGCAAAATCTGTATGAGTATCTTCTGCATGCGGAAAACGAGTTGGAAACAGATAAAACTAGCAATG
ATGCTCAAAGTTCTGTTGGAGCTGGTCAAAGGGTACCGGTTGCTGCAGGAGCTGGGGATACTAACATCTGTGGTGGTATCGTCCAGTTATATTTTGATCG
CATCATAGGAAGGTGCTTAGATTTCAATGCCCAAGTTCGCAAGACTACACTTCAGATTGTGGAGGTGGTGCTACGGCAAGGTCTTGTTCATCCTATTAGT
TGTGTTCCCCATCTCATAGCTCTAGAAACTGACCCATTAGAGTCTAACTCAAAGTTGGCACATCATCTTCTGATGAATATGAATGAGAAGTACCCAGCTT
TCTTTGAAAGTCGATTAGGGGATGGTCTGCAGCTATCATTTTCATTCGTGCATTCTCTTTCTGGCGTCTCCCATGAAAACTTGAATCAAATGTTCCCACA
AAAGGCTGCCGGAAGTTTGAAAGGAAAGTCTGAAGTCGGTAATCTAGCTCAGGCTAGGCTTGGGGTTTCTCGGATCTACAAGCTTATTCGAGGAAATCGT
ATTTCGCGGAATAAGTTTATGTCTTCAATCATTCGCAAATTTGATAATCCGAGCCGTATTGACTCGGTGATTTCCTTTTTGATGTACTGTACCGAAGTGC
TTGCTCTACTGCCATTCACTATTCCCGATGAACCACTCTATCTTGTTTATTCTATAAATCGGGTAATCCAAGTTAGGGCTGGGGTGCTTGAGGCAAACAT
GAAGGGCCTCATATTGCATCTGTCACAAAGAAGTGCCCGTAGGCCACACCATGATACTGAATTAGTTCACCCGGAAGCAAATCAGCTTCATTTCCAGCAA
GTGGAGAAGATGGACTTTGATGGATTGGTTAATGAACAGCATGTTGGTTTGCCATTATTTGATTTGAATGGAACGGCAAAAGAGGAGCCGTCTGACCAGT
CAGTTTTGACCTCAAGAGATTCGAAAGCAGGGTATAGGAAGTCAGTTGAGCCTTGCAGCATCTCCGAGGATGATGTCAAAAGAATCCAGGTTGAGTGTCT
GTCAGCCACTGCATTGCAGCTTCTTCTAAAGCTGAAGAGGCACCTAAAAATTGTGTATGGCCTGAATGATGCCCGCTGTCAGGCATTTTCTCCGAATGAA
CCCCCAAAACCGGGAGAGAATCTATCTAGGCAGAATGTTCCGTTTGACATAAGTGAAACGCGGACTGGTTTGGCTTCAACTTACCAAGATTTGGTACAAG
GATATCAGGATCTGAAAAATGCACTTAAAGAAGACACTGTAGATTATTCTACGTACACAGCTAACATCAAGAGGAAGAGGCCAACACCAAGAAAGAAAAC
GAATCGTCGTGTGATGGCAAATGATGAAGGTATGGAGGATGACGACGACGAAGATGATGATGCAGGGTGGACAGGCGGGAGAAGGAAGTTAAGTGGTAGC
AGCAGTAGGAGAGCCGAAAGCACCAGAAGAGGCAGACAACGATGA
|
||||||||||
|
AA sequence
|
>Lus10015385 pacid=23160861 polypeptide=Lus10015385 locus=Lus10015385.g ID=Lus10015385.BGIv1.0 annot-version=v1.0
MSNPTAEPSGSSSGFSLQGIGLSNTIHSEVAPCLPLPFLPVFCGASDPEIRLFDERSGGGESLSYLDRSEIVAQSARIAELLRETDVSYINLRNNARPMT
LDYVIPLELCDHVLQCNPEAFEHSTPGKAQLSGNIASESRHVRVETSIPVLLKVQNDDVNLMRKTLQDIGSSSRKPKAKKKGVEDISAPVQLEPTDLPGL
QDTIIRNFCDALEDFCVKAEVLSDEHEEVEWLVLPASDVRLLVNEIMLIRGKKLLHLVPLEALVRLLRILDHQIHRAEGLSVDQYDHSDSDVVSNVFSAL
ESIHAALAVMAHNNMPKQLYKEEIIERILEFSKHQMVDILSAYDPSFRALHKPNERETYEDDENEDPQADYAPSGKRWHVQKSVKMKKSAYNKVTSSMNA
ILQKLCTIIGLLKELLLIERLSDSCILQLVKTSFTTLLVDNVQLLQLKAIGLLVGIFYSYTQHRVYIVDEMVQLLWKLPVSKRALRAYHLPDEDQKQIQI
IPALLIQLVHSSANLPEALRQASSGHSILEVSLDATYITKCHEAVTDTCCLFWTRVLQRFTTVKAQDSSEMKVIMENLVTDLLTTLNLPEYPASAIILEI
LCVFLLENAGLKSKDVAVRSMAIDLLGMIATSLKHSAAICSRNTFWMLQALDGGDDANCSLQTDACCICSDGRVEKALFVCQGCNRLFHADCMGAREREV
TRNWYCQICLCKKQLLVLQSFSKMQCKDEGRGNKSRSSKFPENSDLIVPKVEIIQQTLLNYLQDSVSADDMHHFICWFYLCLWYKDDPKSEQRLTYDLMR
LKSNLIVRDSGTSHSTLTRESFNKITRALGQNSSFARGFNKILHLLLASLRENSPVIRAKALRAVSIIVETDPGVLRDKGVQFAVEGRFCDSAISVREAA
LELVGRHIASHPDVGLQYFEKVAERIKDTGVSVRKRAIKIIRDMCTSNANFSQFTTACIEIISRVSDDESSIQDLVCKTFYEFWFEEPSCVQTTFSGDGS
SVPLEVAKKTDQLVEMLRRMSSHSLLITVIKRNLALDFFPQSAKAVGINPMSLASVRKRCELMCKCILERILQVEEMNNTEEEVQTLPYVLALHSFCVVD
PTLCVPSSDPSQFVVTLQPYLKTQVDNRVVANFLENIIFIIDSALPLIRKLTPVLVEELQQDLKQMILRHSFFTVVHACIKCLCTLSRVGGKGASVVEFL
IQMFFKRLDAQTQTFQNKQLVVRSLFCLGLLIRYGSPLVSELGNKNIDIASSLSLFKKYLIVDDFSVKVRSLQALGFVLIARPEYMLEKDIGKILEATLS
PSSHVHMKMQALQNLYEYLLHAENELETDKTSNDAQSSVGAGQRVPVAAGAGDTNICGGIVQLYFDRIIGRCLDFNAQVRKTTLQIVEVVLRQGLVHPIS
CVPHLIALETDPLESNSKLAHHLLMNMNEKYPAFFESRLGDGLQLSFSFVHSLSGVSHENLNQMFPQKAAGSLKGKSEVGNLAQARLGVSRIYKLIRGNR
ISRNKFMSSIIRKFDNPSRIDSVISFLMYCTEVLALLPFTIPDEPLYLVYSINRVIQVRAGVLEANMKGLILHLSQRSARRPHHDTELVHPEANQLHFQQ
VEKMDFDGLVNEQHVGLPLFDLNGTAKEEPSDQSVLTSRDSKAGYRKSVEPCSISEDDVKRIQVECLSATALQLLLKLKRHLKIVYGLNDARCQAFSPNE
PPKPGENLSRQNVPFDISETRTGLASTYQDLVQGYQDLKNALKEDTVDYSTYTANIKRKRPTPRKKTNRRVMANDEGMEDDDDEDDDAGWTGGRRKLSGS
SSRRAESTRRGRQR
|
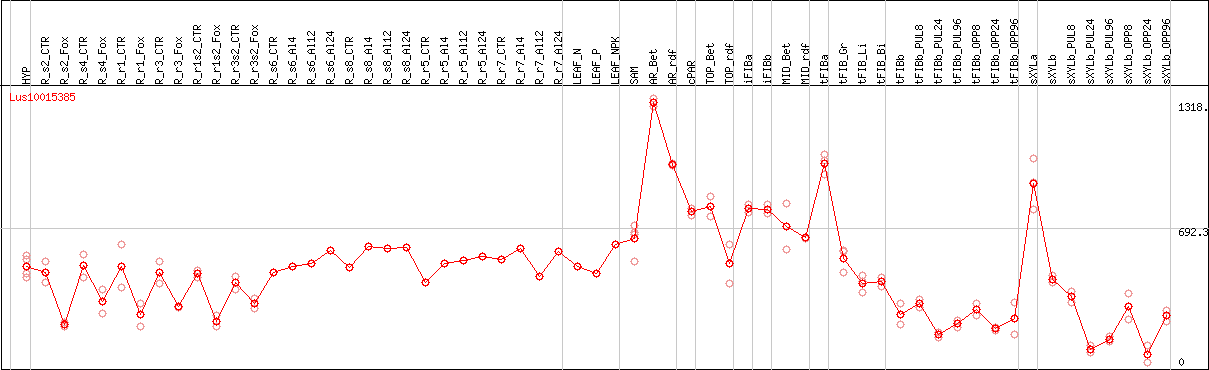
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015385 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
