Lus10015467 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10015467 pacid=23173692 polypeptide=Lus10015467 locus=Lus10015467.g ID=Lus10015467.BGIv1.0 annot-version=v1.0
ATGAACGCTGCAACTGGAGAGCGTACCTTGTTGACGACGCAGGAATTGAAGATCAGTCCAGATACGGCCATGGATACAGAGAAGATAGCTGCTGCAGTAG
AATCTCCGGCGTGGATCGGGTTTTCCAACCGGTTATGTACAAGCTGTTTAAGCAATTCAGGCGAAGAATTGAGATGTAATGAGAAGAAGCTGCCTAAAGT
GAATTGGTCCCAACATGCAAAGGCATACGATAAGTCATCATCAGTGCAGAAGGAAAATTTGAGCTCGAATTTCCTGTATTCTCTAGAATCTCAGAAACCT
CGTGTTGAAGTAGAAATGGCGAGAAGGTTAGCCTCTTGTCAAGTTCAGAGTTTCCAGAGACAGCTGAATCCACAAGTTGAAAAGGCTTGGCTTTCTCTTT
CCAATCTTCATATATCTACCAGAAGCTATTGTAGACCTGGGAAAACGGGAGCAATTAAGAATGTGCAAGCTAGTAAGGGACGTGCTCCCGTTGTACAGAG
CTCATCTAATGTACGTAAATATGGTGAAGGTGTGGTTTCCAAACAGACCTTTCATAAACCCGATGTTCAGAATGGTGACACTACAGAGCATACAAGATCG
TGCAATGAAGAAATGGGGAAAGGTGGGAACAAGGTGTCGTGGGCCAGTCAAACCATGCCTTCAGCTTCCTCTGGTCCCTTAGGCTATCAGAGTACGACTG
CACAAGAGTTAGCAGAAGGGTCAGCTGATTATATGGATGACGAGGAATTAATCCAGAGCATTGATCTCGATCAGATAGTTATGGATCATTACCAGTCTAC
TTGCACACCTCAACCTTCAGTTTCTAAGTTTCAACCAATAACTCCCTCCGTGAATAAGGCTAACTTCAGTGGGCCTGAAGATACATTTCTACCATCTGAG
TTATCCTCAAAGTGCAGCCATGGAATTCAGCTAGGGTTTTGTCCACAAGCAACGGTTCATATGCAGGAGATAAAGGACAAATTACTCAGCATATCTAACG
AATTACTTGATAATGCTAACCTTCATCCAGTAGAGATGGAGAAGCTTCGCCAAGATAGGTTGCATCTTAATAAACAGCTCCAGCAACTTGAGGGGCATAT
TAGTAATCAAGAGAGGCACAAATCAAACTTTTCCACATCCACTACAACTTGCAATGCTCAATATGAAACACCTCAGATGTCAGCTTTCAGAATGGAGCCA
AATCACTATCTTCAAAATGAATCCGGTGCAAGAGGGTATGAAAATTGGAACTCATCATCACTTCCATTCACTCCCATGGATGGATTTGGTGGTTCATATA
CTCCTGTGGTGAGGGAACCTTACATACCAAAGTTTGTTGAGGTGAATTACTCAGACGGCTCAAATGACCCACAGTGGAGCTCCACAAATTTCCAGTGGAC
AAAAAGCCTGGAGGAAAACAACAAGAAAGTGTTTGGAAATCATTCTTTTCGTCCCAATCAAAGAGAAGTCATTAATGCCACAATGAGCGGGTATGATGTC
TTTGTTCTAATGCCTACAGGTGGTGGAAAGAGTCTGACATATCAGCTACCTGCTTTAGTTTGTCCCGGGATAACATTGGTAATATCTCCACTGGTATCAC
TTATTCAAGATCAGATAATGCATCTTTTGCAGGCAAATATACCTGCTGCTTATTTGAGTGCAAATATGGAATGGTCTGAACAGCAGGAGATAATGAATGA
ACTTTGCTCCGACTATTGTAGATACAAGCTGTTATATGTTACACCAGAAAAAGTTGCAAAGAGCGATGCTCTTTTGCGAAATTTGGAGAGCTTAAATGCT
CGTGGAATGCTTGCCAGAATTGTAATTGACGAAGCTCATTGCGTGAGTCAGTGGGGACATGATTTTAGACCAGATTACAAGGAACTTGGAATCCTGAAAA
AGAAATTTGTGAAAACCCCAGTTTTGGCTTTAACTGCTACCGCAACAGCTAGCGTGAAAGAAGATGTTGTGCAAGCTCTTGGACTTGTTGATTGCATTAT
TTTTCGGCAGAGTTTTAATAGACCGAACCTGCAGTATATTGTGATTTCCAAGACAAAAAAGTGTTTGGATGACATGGACAACTTCATTAGGACGACCCAT
TTTGATGAATGTGGAATTGTTTACTGTCTCTCAAGAATGGACTGTGAAAAAGTTGCTGAAAAGTTGCAGCAATATGGACATAAGACGGCATTTTATCATG
GTAACATGGAGGCTTCTCAACGTGCTTTTGTCCAGAGCCAGTGGAGCAAAGATGAAGTTAATATAATTTGTGCCACAGTGGCATTTGGAATGGGTATCAA
CAAACCTGATGTTCGCTTTGTGATTCATCATTCTCTCCCAAAATCCATTGAAGGTTACCATCAGGAATGTGGTCGTGCTGGCCGAGATGGTCAACCCTCA
TCTTGTGTGCTATACTACAGCTACAGTGACTATATAAGGGTAAAGCATATGCTGTCGCAAGGACAGGCAGAGCAAAGTCCATGGACATCCGGGTATCGAG
CTAGCACAGTACAGCCAGATCGAATATATGAGAAAAATACTGAAAACCTCCTTCGAATGGTAAGCTATTGTGAAAATGATGTTGATTGTAGACGTTTCCT
ACAACTTGTGCATTTTGGAGAGAAGTTTAGCTCTGGAAACTGCAAGAAAACTTGTGATAACTGCAAAACTACTAAAAGCTTAGTGGAGAAGGATGTCACC
CAGATTGCAAAAGAATTGGTCGAGCTGGTAAAATTGGCAGGACAAAAATATTCGGCTTCCCATATTTTGGAAGTCTACAGGGGATCCATGAGCCAATTGG
TAAAGAAACATAGACATGAGACCTTGAGTCTTCATGGAGCCGGAAAACATCTGGCAAAAGAAGAGGCGTCTCGTGTATTGCGTCATCTTGTCATTGAGGA
GTACCTTGTGGAGGAAGTTAAGAAAAGTGAGGTCTACGGTTCTGTTTCATCAGTGTTAAAGGTGAATCAACGCAAAGCATCAAATCTGTATTCTGGAAAG
CAGATAATGCTGAGGTTCCTTTCCTCCGCAAAAGCGTCTAGACCCTGCAAATCTGATGCAACTCCTGCAAAAGGAGCACTGCTGCCTAATAAGCACAGTC
CTCCGCCGGCTGACACTCCAGCATCGACTCGACCTGAAATCAACTTGGATCTTAGTGCAAAGTTATATAATGCTTTACGACTGCTTCGAACCGCGATTCT
TAAAGAAGGCAATCAAGATTTGGGTGCGCACCATATATTTGCGAATGCTACACTACAAAACATAAGCAAAAGGATTCCGAGAACAAAGGAAGAACTGCTG
GAGGTTAACGGAATTGGAAAAGCAAAGTTGGCAAAGTATGGAGATCGGGTACTGGAGACCATTGAAGCCACAATCCGAAAATACTACAAAAATGGGGCAA
GTGGAAGCAGCAGCAACGACAGCAGCAGCGAAAAGGCAAAGAGGAGGCGAAACGGTGCTGCTGCAAATGGCGGTGGAGGGGATGATCGGGAGGAGGAGGA
TGACGAGTTTGTGAAAAGCACAGGTCGGCCTGTGAAGAAGAGGCTTGCTAAGAAGCTACATGACAACAATGGTGTTAATGAGTTTGGTAATCTTGGACTA
GAAGATTATGTGGAAGATATTCCGGATGATGATCTAGTTTTCTACGATTTGACTGGATTCCAATTCACTTGCCTTGCCTCAATTAGCAATTCGGCTGAAC
AGTGGGTAAAGAACATGAGCAGAAGCACGGACGATGACCCAGCTGCGGCAGAAGAACAGGGTCGATCTCTAAAGCTTGGACTCGGTGCAAGAGTAGCTCC
ACAATCCAACATTACACCTTCAAACGATCCGGCAGCACGGAGACTGCACAAGAAGTTGGAAGCTGGGAAGAAAAGAGCTTCCAATGTTGCTAATGAGACC
GGAGGACCTAGAGGTGACGAAAGTGATGACAGCGATGGAGAAATGGAAAGCAGAACCGGTGCATTTGCCAAGAAGAGAGCTAGACCCCCAGCTTTACCCT
CACAGCTTAACACCAAGAAGAAGAAGAGGAACCACAAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10015467 pacid=23173692 polypeptide=Lus10015467 locus=Lus10015467.g ID=Lus10015467.BGIv1.0 annot-version=v1.0
MNAATGERTLLTTQELKISPDTAMDTEKIAAAVESPAWIGFSNRLCTSCLSNSGEELRCNEKKLPKVNWSQHAKAYDKSSSVQKENLSSNFLYSLESQKP
RVEVEMARRLASCQVQSFQRQLNPQVEKAWLSLSNLHISTRSYCRPGKTGAIKNVQASKGRAPVVQSSSNVRKYGEGVVSKQTFHKPDVQNGDTTEHTRS
CNEEMGKGGNKVSWASQTMPSASSGPLGYQSTTAQELAEGSADYMDDEELIQSIDLDQIVMDHYQSTCTPQPSVSKFQPITPSVNKANFSGPEDTFLPSE
LSSKCSHGIQLGFCPQATVHMQEIKDKLLSISNELLDNANLHPVEMEKLRQDRLHLNKQLQQLEGHISNQERHKSNFSTSTTTCNAQYETPQMSAFRMEP
NHYLQNESGARGYENWNSSSLPFTPMDGFGGSYTPVVREPYIPKFVEVNYSDGSNDPQWSSTNFQWTKSLEENNKKVFGNHSFRPNQREVINATMSGYDV
FVLMPTGGGKSLTYQLPALVCPGITLVISPLVSLIQDQIMHLLQANIPAAYLSANMEWSEQQEIMNELCSDYCRYKLLYVTPEKVAKSDALLRNLESLNA
RGMLARIVIDEAHCVSQWGHDFRPDYKELGILKKKFVKTPVLALTATATASVKEDVVQALGLVDCIIFRQSFNRPNLQYIVISKTKKCLDDMDNFIRTTH
FDECGIVYCLSRMDCEKVAEKLQQYGHKTAFYHGNMEASQRAFVQSQWSKDEVNIICATVAFGMGINKPDVRFVIHHSLPKSIEGYHQECGRAGRDGQPS
SCVLYYSYSDYIRVKHMLSQGQAEQSPWTSGYRASTVQPDRIYEKNTENLLRMVSYCENDVDCRRFLQLVHFGEKFSSGNCKKTCDNCKTTKSLVEKDVT
QIAKELVELVKLAGQKYSASHILEVYRGSMSQLVKKHRHETLSLHGAGKHLAKEEASRVLRHLVIEEYLVEEVKKSEVYGSVSSVLKVNQRKASNLYSGK
QIMLRFLSSAKASRPCKSDATPAKGALLPNKHSPPPADTPASTRPEINLDLSAKLYNALRLLRTAILKEGNQDLGAHHIFANATLQNISKRIPRTKEELL
EVNGIGKAKLAKYGDRVLETIEATIRKYYKNGASGSSSNDSSSEKAKRRRNGAAANGGGGDDREEEDDEFVKSTGRPVKKRLAKKLHDNNGVNEFGNLGL
EDYVEDIPDDDLVFYDLTGFQFTCLASISNSAEQWVKNMSRSTDDDPAAAEEQGRSLKLGLGARVAPQSNITPSNDPAARRLHKKLEAGKKRASNVANET
GGPRGDESDDSDGEMESRTGAFAKKRARPPALPSQLNTKKKKRNHK
|
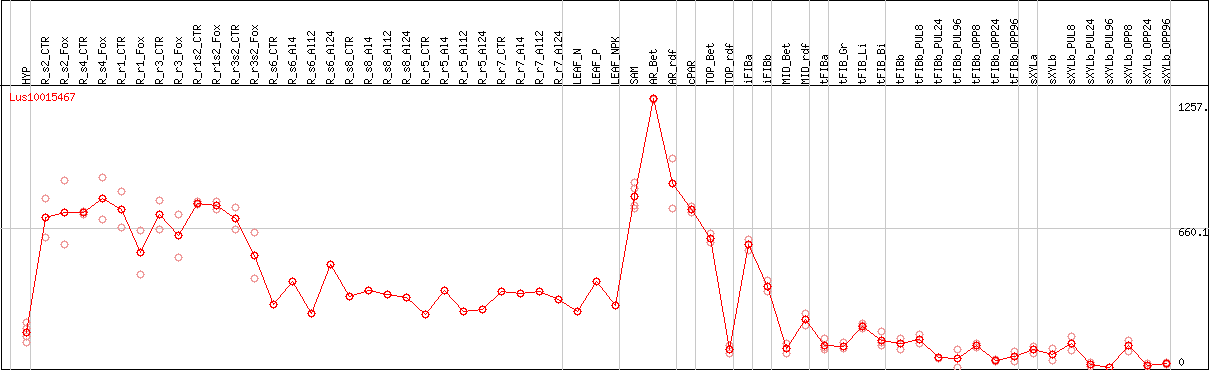
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015467 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
