External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G15560 189 / 2e-56
AtCLA1, DXS, DXPS2, DEF, CLA1
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
AT3G21500 185 / 2e-55
DXPS1
1-deoxy-D-xylulose 5-phosphate synthase 1 (.1.2)
AT5G11380 99 / 2e-24
DXPS3
1-deoxy-D-xylulose 5-phosphate synthase 3 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015517
390 / 2e-133
AT4G15560 1020 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10019992
388 / 2e-133
AT4G15560 504 / 7e-171
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10019991
383 / 2e-130
AT4G15560 1018 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10001322
255 / 1e-81
AT4G15560 1021 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10006984
251 / 4e-80
AT4G15560 1023 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10012724
212 / 4e-65
AT4G15560 1030 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10002663
209 / 6e-64
AT4G15560 1026 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10033987
190 / 5e-57
AT4G15560 1134 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Lus10012789
189 / 3e-56
AT4G15560 1229 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G171700
247 / 3e-78
AT4G15560 1037 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Potri.007G058500
209 / 4e-64
AT4G15560 999 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Potri.010G015200
189 / 2e-56
AT4G15560 1227 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Potri.008G196500
185 / 5e-55
AT4G15560 1207 / 0.0
1-DEOXY-D-XYLULOSE 5-PHOSPHATE SYNTHASE 2, CLOROPLASTOS ALTERADOS 1, Deoxyxylulose-5-phosphate synthase (.1)
Potri.006G249700
131 / 2e-35
AT5G11380 949 / 0.0
1-deoxy-D-xylulose 5-phosphate synthase 3 (.1.2)
Potri.018G031800
113 / 3e-29
AT5G11380 958 / 0.0
1-deoxy-D-xylulose 5-phosphate synthase 3 (.1.2)
PFAM info
Representative CDS sequence
>Lus10015518 pacid=23173619 polypeptide=Lus10015518 locus=Lus10015518.g ID=Lus10015518.BGIv1.0 annot-version=v1.0
ATGACTTCTTCAGTTCTGAGATCAACTTTCCTTCCTAATTTGATCCATTCTCAAGATGCTCAATCTTTCTCAAGCTTCCCTCCAAAACCCACCTCAGCCA
CTCTCCGACCCAGAAAGAAGAGCAGAGGAGGCCTGGTGGCTGCAATGGAGAGTGACGATGTCAATGACAATAAGGTAGAGAAATACAATGAAATTACTAA
GGAAAAGAGAACGCTTGACTTCTCAGGGAACAAGCCTTCCACTCCAATTCTGGACACCATTAACTATCCAAATCACATGAACAATCTCTCAACTGAGGAG
CTTGAAACATTGGCTGATGAGTTGAGGGAAGAGATTGTCTACTCTGTTTCCAAGACAGGTGGTCACCTTAGCTCAAGCTTGGGAGTTGCTGAACTCACTG
TCGCCCTCCACCATGTTTTCAACACCCCCGAGGATAAGATCATCTGGGACGTTGGTCATCAGTCATACCCGCACAAGATATTAACCGGAAGAAGATCCAA
GATGGGGTCCATCCGCCAGACATTCGGATTAGCAGGATTCCCAAAGAGGGACGAGAGTGAGCACGATGCATTGTGA
AA sequence
>Lus10015518 pacid=23173619 polypeptide=Lus10015518 locus=Lus10015518.g ID=Lus10015518.BGIv1.0 annot-version=v1.0
MTSSVLRSTFLPNLIHSQDAQSFSSFPPKPTSATLRPRKKSRGGLVAAMESDDVNDNKVEKYNEITKEKRTLDFSGNKPSTPILDTINYPNHMNNLSTEE
LETLADELREEIVYSVSKTGGHLSSSLGVAELTVALHHVFNTPEDKIIWDVGHQSYPHKILTGRRSKMGSIRQTFGLAGFPKRDESEHDAL
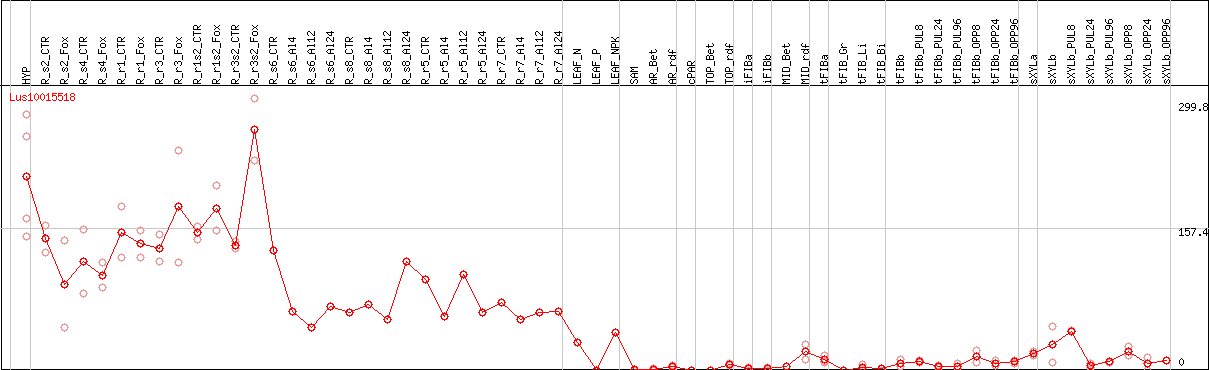
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015518 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.