External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G17710 301 / 3e-103
AtPEPC1
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
AT1G73010 299 / 2e-102
AtPPsPase1, ATPS2
pyrophosphate-specific phosphatase1, phosphate starvation-induced gene 2 (.1)
AT4G29530 229 / 1e-75
Pyridoxal phosphate phosphatase-related protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037652
379 / 9e-135
AT1G17710 255 / 2e-85
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
Lus10001479
330 / 8e-115
AT1G17710 381 / 1e-134
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
Lus10008479
327 / 1e-113
AT1G17710 378 / 3e-133
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
Lus10015632
315 / 5e-109
AT1G17710 372 / 1e-130
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
Lus10037650
308 / 4e-106
AT1G17710 359 / 1e-125
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
Lus10008573
219 / 8e-72
AT4G29530 285 / 5e-98
Pyridoxal phosphate phosphatase-related protein (.1)
Lus10032691
50 / 5e-07
AT5G53770 479 / 3e-164
Nucleotidyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G196800
333 / 3e-116
AT1G73010 397 / 2e-140
pyrophosphate-specific phosphatase1, phosphate starvation-induced gene 2 (.1)
Potri.003G034600
320 / 9e-111
AT1G17710 399 / 1e-141
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
Potri.018G054800
269 / 7e-91
AT1G17710 311 / 7e-107
Arabidopsis thaliana phosphoethanolamine/phosphocholine phosphatase 1, Pyridoxal phosphate phosphatase-related protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0137
HAD
PF06888
Put_Phosphatase
Putative Phosphatase
Representative CDS sequence
>Lus10015634 pacid=23147197 polypeptide=Lus10015634 locus=Lus10015634.g ID=Lus10015634.BGIv1.0 annot-version=v1.0
ATGTCGTCCGGAGGAGTGGTAGTACTCTACGACTTCGACGAGACAATCGTAGGAGTCGACACCGATTACTGGTTGATCGACGAATTCGGTTACACACATT
TGGCCGACCAACTCCGCCCCAATACGCCTTGGAACTCTATCATTGATATGATGCTGAAAGAGATACAAAAACATGGGAGAACCATTGATGACATTGTTGG
AGCTTTGGAAAGAATTCCAATCCTTCCTAGGGTTGTTTCCTCCATCAAATCTGCTTATGATTTGGGGTGCGAGCTGAGGATCGTAAGTGATTCGAACGTG
TTCTTCATCGAACAAGTGTTGGAGAACCTTGGAGTGAGGAGATTTTTCTCAGAGATCAACACAAATTCAGGGTTTGTTGATGATGAAGGGAGGTTGAGGG
TTGCTCCTTACCATGATTTCACCAAGGCTTCACATGGTTGTACCCTTTGCCCTCCAAACATGTGCAAGGGGTTGATAGTGGAGAGAATCCAAGCATCCAT
AGCAAAAAACAACAGCAAGAGGATCATCTACGTGGGAGATGGAATTGGGGATTACTGCCCAAGCTTGAAGCTCAAGGAAACTGATTTCCTGTTGCCCAGG
AAGAACTTCAAATTGTGGGATTTGATCAACAATAATAAGAACCCACTTCATAAAATAATCAAGGCTCAGATCCATGAATGGGGTGATGGAGATGAGCTCG
ACATTGCTGACATCGTTCAGAAAGTTAGTTAA
AA sequence
>Lus10015634 pacid=23147197 polypeptide=Lus10015634 locus=Lus10015634.g ID=Lus10015634.BGIv1.0 annot-version=v1.0
MSSGGVVVLYDFDETIVGVDTDYWLIDEFGYTHLADQLRPNTPWNSIIDMMLKEIQKHGRTIDDIVGALERIPILPRVVSSIKSAYDLGCELRIVSDSNV
FFIEQVLENLGVRRFFSEINTNSGFVDDEGRLRVAPYHDFTKASHGCTLCPPNMCKGLIVERIQASIAKNNSKRIIYVGDGIGDYCPSLKLKETDFLLPR
KNFKLWDLINNNKNPLHKIIKAQIHEWGDGDELDIADIVQKVS
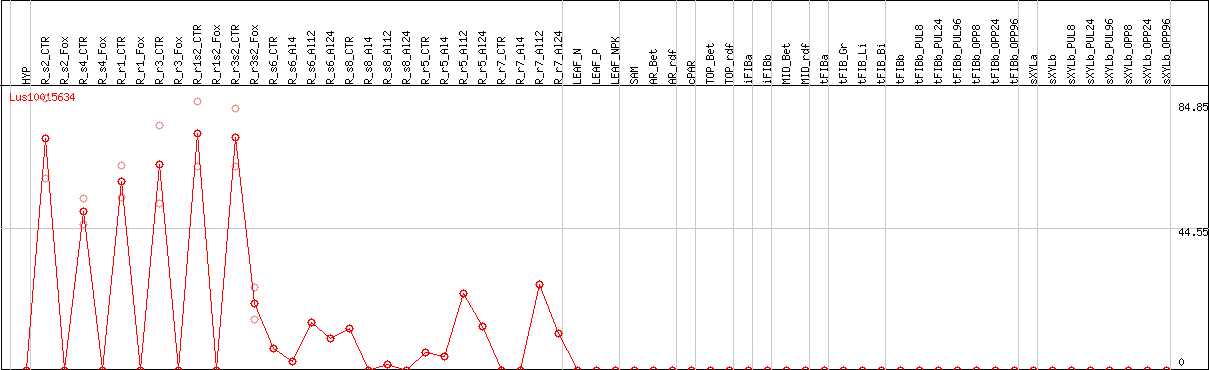
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015634 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.