External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G20630 108 / 1e-29
ATGER3, GLP3A, GLP3B, GLP3
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
AT1G72610 98 / 3e-25
ATGER1, GLP1
A. THALIANA GERMIN-LIKE PROTEIN 1, germin-like protein 1 (.1)
AT3G62020 86 / 6e-21
GLP10
germin-like protein 10 (.1.2)
AT1G02335 82 / 3e-19
GL22
germin-like protein subfamily 2 member 2 precursor (.1)
AT5G38960 80 / 2e-18
RmlC-like cupins superfamily protein (.1)
AT5G39160 77 / 1e-17
RmlC-like cupins superfamily protein (.1.2.3)
AT5G39190 77 / 1e-17
ATGER2, GLP2A
GERMIN-LIKE PROTEIN 2A, A. THALIANA GERMIN LIKE PROTEIN 2, germin-like protein 2 (.1.2)
AT5G38910 76 / 4e-17
RmlC-like cupins superfamily protein (.1)
AT5G38940 76 / 9e-17
RmlC-like cupins superfamily protein (.1)
AT5G38930 75 / 1e-16
RmlC-like cupins superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037663
211 / 9e-70
AT5G20630 209 / 2e-68
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10037661
185 / 2e-59
AT5G20630 182 / 6e-58
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10015643
183 / 2e-58
AT5G20630 182 / 4e-58
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10020851
178 / 1e-56
AT5G20630 181 / 2e-57
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10037662
175 / 1e-56
AT5G20630 124 / 2e-36
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10015644
176 / 1e-55
AT5G20630 192 / 6e-62
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10003652
144 / 2e-42
AT4G25300 206 / 3e-64
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10037664
133 / 6e-39
AT5G20630 198 / 3e-64
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
Lus10036296
128 / 4e-37
AT5G20630 213 / 3e-70
GERMIN-LIKE PROTEIN 3, ARABIDOPSIS THALIANA GERMIN 3, germin 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF00190
Cupin_1
Cupin
Representative CDS sequence
>Lus10015645 pacid=23147251 polypeptide=Lus10015645 locus=Lus10015645.g ID=Lus10015645.BGIv1.0 annot-version=v1.0
ATGTTGCTACTCCGCACACGCCGCGGATTTCTGCATATCCGATCTATCCCTCCAAACCCCGTCCGGCTATGCCTGCAAGGACCTCACCAAAGCCTCCTCC
AGCGACTTCGTCTTCACCGGAATATCCAAACAGTGCAACACTTCCAACCCCAACAAGGCAGCTGCTACCCTGGCCTTTGTCCACCAATGTCCAGCCCTAA
ACGGGCTGGGAATCTCCCTACCCGGGCCGATTTCGCCCTGGACGGGGTGATCCCGCTCCACACCCACCCGAGGGCCACCGAGATGGCCTACCTGGTGGAG
GGCACCCTCACGGTGGGGTTCGTCGCCGCGGGGACCAACAAGGTGTATGTGAATACAATCAGAAAGGGGGAAGTGACGGTGTTTCCTCAAGAGATGCCGC
ATTTTGTGGTGAATGATGATACCAAAAGGCCAGCCGTGGTGCTTGTGGCTTTGAGCAGTCTCGATCCGGAGAATCTGTTGGTGAGTTCTGCGCTGTTTGG
GAATGATCTCGCATCGGCTTTGGTGGAGAAGACGACCGGAATTGGTGAATCCGAAGCGAAAAGGCTTAAAGCTCTTTTCGGTGGTTCGGGTTAA
AA sequence
>Lus10015645 pacid=23147251 polypeptide=Lus10015645 locus=Lus10015645.g ID=Lus10015645.BGIv1.0 annot-version=v1.0
MLLLRTRRGFLHIRSIPPNPVRLCLQGPHQSLLQRLRLHRNIQTVQHFQPQQGSCYPGLCPPMSSPKRAGNLPTRADFALDGVIPLHTHPRATEMAYLVE
GTLTVGFVAAGTNKVYVNTIRKGEVTVFPQEMPHFVVNDDTKRPAVVLVALSSLDPENLLVSSALFGNDLASALVEKTTGIGESEAKRLKALFGGSG
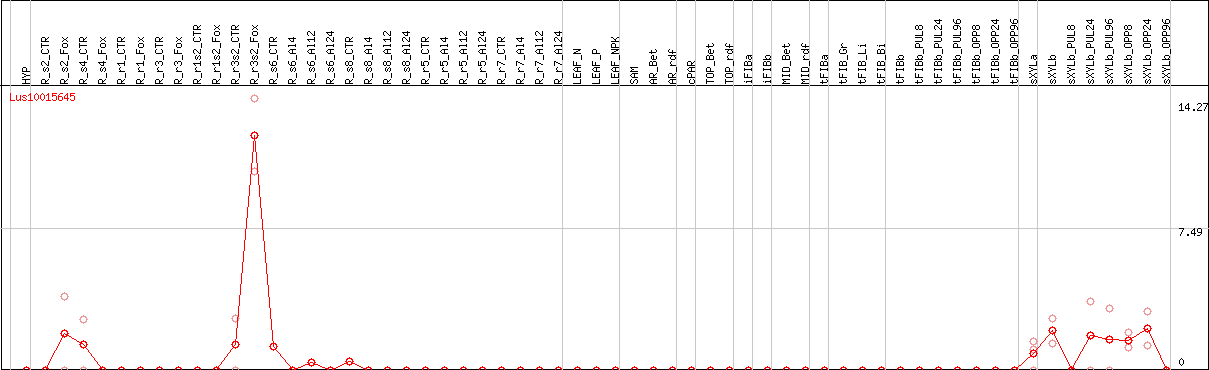
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015645 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.