External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G20570 192 / 6e-65
HRT1, ROC1, RBX1, ATRBX1
REGULATOR OF CULLINS-1, RING-box 1 (.1.2.3)
AT3G42830 174 / 9e-58
RING/U-box superfamily protein (.1)
AT3G05870 69 / 2e-16
APC11
anaphase-promoting complex/cyclosome 11 (.1.2)
AT3G47180 40 / 8e-05
RING/U-box superfamily protein (.1)
AT4G12140 39 / 0.0001
RING/U-box superfamily protein (.1)
AT5G40250 40 / 0.0002
RING/U-box superfamily protein (.1)
AT3G16090 40 / 0.0002
AtHrd1A
homolog of yeast Hrd1, RING/U-box superfamily protein (.1)
AT5G17600 40 / 0.0002
RING/U-box superfamily protein (.1)
AT1G65040 39 / 0.0002
AtHrd1B
homolog of yeast Hrd1, RING/U-box superfamily protein (.1.2.3)
AT3G03550 39 / 0.0004
RING/U-box superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013153
192 / 5e-65
AT5G20570 206 / 2e-70
REGULATOR OF CULLINS-1, RING-box 1 (.1.2.3)
Lus10037680
134 / 2e-42
AT5G20570 122 / 9e-38
REGULATOR OF CULLINS-1, RING-box 1 (.1.2.3)
Lus10008115
117 / 2e-35
AT5G20570 129 / 2e-40
REGULATOR OF CULLINS-1, RING-box 1 (.1.2.3)
Lus10015126
67 / 6e-15
AT3G05870 156 / 2e-50
anaphase-promoting complex/cyclosome 11 (.1.2)
Lus10031549
64 / 2e-13
AT3G05870 158 / 6e-50
anaphase-promoting complex/cyclosome 11 (.1.2)
Lus10040359
39 / 0.0004
AT3G16090 652 / 0.0
homolog of yeast Hrd1, RING/U-box superfamily protein (.1)
Lus10023477
39 / 0.0005
AT3G16090 650 / 0.0
homolog of yeast Hrd1, RING/U-box superfamily protein (.1)
Lus10034587
38 / 0.0006
AT1G68070 489 / 6e-172
Zinc finger, C3HC4 type (RING finger) family protein (.1)
Lus10032161
38 / 0.001
AT5G40250 413 / 7e-144
RING/U-box superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10015667 pacid=23147194 polypeptide=Lus10015667 locus=Lus10015667.g ID=Lus10015667.BGIv1.0 annot-version=v1.0
ATGGACGTCGACGTTGCTATGGTACCGGCCGGCGAAGGAAGCTCCAGCGCTCCCGGACCTTCGACCTGCTGCAAGAAGGCCAAGAGCTTCGAGATAAAAA
AGTGGAATGCCGTCTCTCTTTGGGCATGGGATATTGTGGTGGATAACTGCGCTATTTGCAGGAATCATATTATGGATCTTTGCATTGAGTGCCAAGCTAA
TCAAGCGAGTGCCACTAGCGAGGAGTGTACTGTTGCTTGGGGTGTTTGTAACCATGCTTTCCACTTCCACTGCATTAGCCGTTGGTTGAAAACCCGCCAA
GTCTGCCCTCTTGATAATAGCGAATGGGAGTTCCAGAAGTACGGTCACTAG
AA sequence
>Lus10015667 pacid=23147194 polypeptide=Lus10015667 locus=Lus10015667.g ID=Lus10015667.BGIv1.0 annot-version=v1.0
MDVDVAMVPAGEGSSSAPGPSTCCKKAKSFEIKKWNAVSLWAWDIVVDNCAICRNHIMDLCIECQANQASATSEECTVAWGVCNHAFHFHCISRWLKTRQ
VCPLDNSEWEFQKYGH
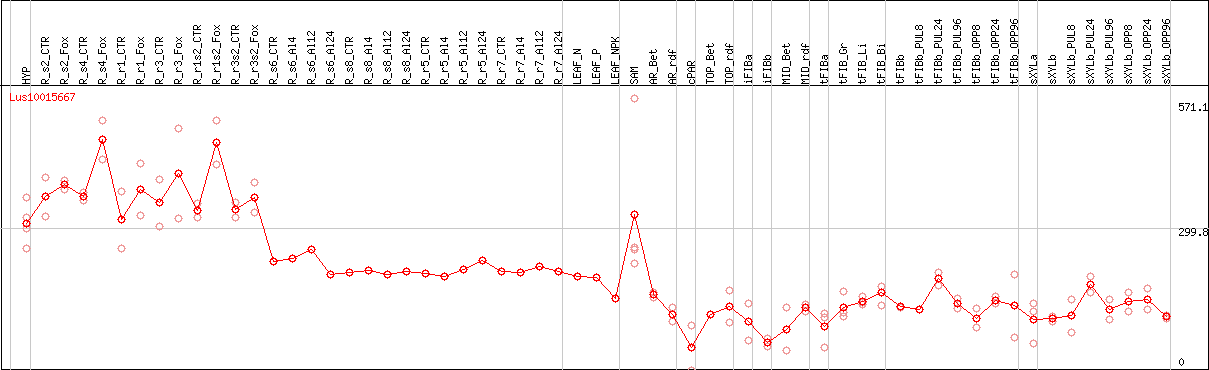
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015667 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.