External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G06850 364 / 2e-121
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT5G48060 364 / 6e-119
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT5G12970 346 / 1e-114
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
AT3G57880 339 / 9e-112
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
AT1G51570 338 / 2e-111
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
AT4G11610 335 / 3e-108
NTRB, ATNTRB
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT1G22610 314 / 3e-100
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT4G00700 296 / 2e-93
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT1G74720 274 / 5e-85
QKY
QUIRKY, C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
AT3G61300 271 / 1e-84
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037479
434 / 4e-152
AT5G48060 866 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10021030
372 / 3e-124
AT5G06850 1383 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10023823
371 / 5e-124
AT5G06850 1382 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10012026
344 / 1e-113
AT3G57880 1496 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Lus10016280
344 / 2e-113
AT3G57880 1499 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Lus10000605
333 / 2e-107
AT4G11610 1634 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10018839
323 / 8e-104
AT4G11610 1348 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10009459
300 / 1e-102
AT1G22610 498 / 2e-170
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Lus10001538
308 / 2e-97
AT1G22610 1425 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G065600
399 / 1e-134
AT5G06850 1308 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.001G271400
399 / 4e-132
AT5G48060 1481 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.006G058700
377 / 2e-126
AT5G06850 1321 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.016G049300
376 / 6e-126
AT5G06850 1355 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.016G049100
339 / 6e-112
AT3G57880 1425 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.003G125900
341 / 2e-110
AT4G11610 1543 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.006G058900
334 / 6e-110
AT3G57880 1372 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.001G105400
338 / 4e-109
AT4G11610 1567 / 0.0
C2 calcium/lipid-binding plant phosphoribosyltransferase family protein (.1)
Potri.T085601
319 / 5e-104
AT5G12970 1315 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
Potri.003G210801
318 / 6e-104
AT5G12970 1300 / 0.0
Calcium-dependent lipid-binding (CaLB domain) plant phosphoribosyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0484
Peroxisome
PF08372
PRT_C
Plant phosphoribosyltransferase C-terminal
Representative CDS sequence
>Lus10015710 pacid=23147250 polypeptide=Lus10015710 locus=Lus10015710.g ID=Lus10015710.BGIv1.0 annot-version=v1.0
ATGGGCAAGCTTCATCTCGTGATTCGGTTCACTACATTGTCACTAGCCAATTTGATATCCGCGGATGGACAACCGATCCTGCCGAGGATGCACTACCTGC
ATCCATTAACGGTCAACCAAGTCGAAAGCCTAAGGTACCAAGCGATGAACATCGTAGAAGTGAGGCTAGGTAGGGCAGAGCCACAGCTTCAAAAAGAAGT
CGTTGAGTACATGCTAGATGTCGATTCCCACATGTGGAGCATGAGGAGGAGCAAGGTTAACTTCTTCCGGATCATGTCACTAGTTTCAGGGGCGGTCTCG
ATGAGCAAGTGGTTAGACGACATTTGTCAATGGAGGAAACCTGTGACATCGGTACTAGTACACGCCCTGTTCCTGATACTAATCTGGTACCCGGAGCTGA
TCCTCCCGACAATGTTCCTCTACATGTTCCTGATCGGGATCAAGAACTATAGGTTCCGCCCCAGGCACCTGCACCACATGGACAATAAGCTGTCCTGGGC
GGAGGCGGTGAGCCCCGACGAGCTCGACGAGGAGTTCGACATTTTCCCTACTTCGAGGGCTCACGATCTCGTCGGGATGAGGTACGACAGGCTGAGGAGC
ATCGCTGGTAGGATACAGTATGAGGTGGGGGATGTAGCGACGCAAGGTGAGAGTTTTCAGTCACTGTTGAGTTGGAGGGACCCCAGAGCTACTAGTTTGT
TTCGTAGTGTTCCGTCTTTGTGCGGCTATAGTGCTTTATGTGACGCCGTTTCGAGTGGTTGCTATGGGCTCGGATTTGTACTACTTGAGGCACCCTAA
AA sequence
>Lus10015710 pacid=23147250 polypeptide=Lus10015710 locus=Lus10015710.g ID=Lus10015710.BGIv1.0 annot-version=v1.0
MGKLHLVIRFTTLSLANLISADGQPILPRMHYLHPLTVNQVESLRYQAMNIVEVRLGRAEPQLQKEVVEYMLDVDSHMWSMRRSKVNFFRIMSLVSGAVS
MSKWLDDICQWRKPVTSVLVHALFLILIWYPELILPTMFLYMFLIGIKNYRFRPRHLHHMDNKLSWAEAVSPDELDEEFDIFPTSRAHDLVGMRYDRLRS
IAGRIQYEVGDVATQGESFQSLLSWRDPRATSLFRSVPSLCGYSALCDAVSSGCYGLGFVLLEAP
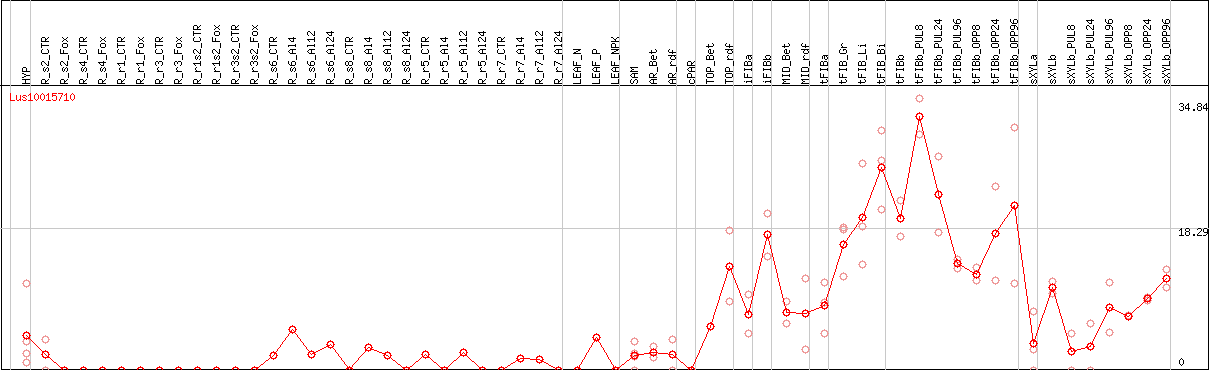
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015710 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.