External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45430 171 / 3e-51
AT-hook
AHL22
AT-hook motif nuclear-localized protein 22 (.1)
AT4G22810 167 / 1e-49
AT-hook
Predicted AT-hook DNA-binding family protein (.1)
AT4G12050 165 / 1e-48
AT-hook
Predicted AT-hook DNA-binding family protein (.1)
AT3G60870 160 / 1e-47
AT-hook
AHL18
AT-hook motif nuclear-localized protein 18 (.1)
AT2G35270 156 / 9e-46
AT-hook
GIK, 2-ATH, AHL21
GIANT KILLER, Predicted AT-hook DNA-binding family protein (.1)
AT4G14465 152 / 2e-44
AT-hook
AHL20
AT-hook motif nuclear-localized protein 20 (.1)
AT2G42940 145 / 5e-42
AT-hook
Predicted AT-hook DNA-binding family protein (.1)
AT4G35390 144 / 3e-41
AT-hook
AGF1
AT-hook protein of GA feedback 1 (.1)
AT1G76500 144 / 4e-41
AT-hook
SOB3, AHL29
SUPPRESSOR OF PHYB-4#3, AT-hook motif nuclear-localized protein 29, Predicted AT-hook DNA-binding family protein (.1)
AT3G04570 143 / 1e-40
AT-hook
AHL19
AT-hook motif nuclear-localized protein 19 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009301
317 / 2e-108
AT3G60870 223 / 4e-72
AT-hook motif nuclear-localized protein 18 (.1)
Lus10020763
157 / 2e-46
AT2G42940 294 / 1e-100
Predicted AT-hook DNA-binding family protein (.1)
Lus10006577
146 / 8e-42
AT3G04570 270 / 1e-89
AT-hook motif nuclear-localized protein 19 (.1)
Lus10031460
142 / 1e-41
AT2G42940 214 / 1e-70
Predicted AT-hook DNA-binding family protein (.1)
Lus10000519
139 / 4e-39
AT2G35270 205 / 1e-64
GIANT KILLER, Predicted AT-hook DNA-binding family protein (.1)
Lus10031458
133 / 4e-38
AT2G42940 212 / 1e-69
Predicted AT-hook DNA-binding family protein (.1)
Lus10002711
135 / 2e-37
AT4G14465 211 / 1e-66
AT-hook motif nuclear-localized protein 20 (.1)
Lus10042551
116 / 1e-30
AT3G04570 202 / 4e-64
AT-hook motif nuclear-localized protein 19 (.1)
Lus10030966
114 / 2e-30
AT2G35270 199 / 1e-63
GIANT KILLER, Predicted AT-hook DNA-binding family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G070800
202 / 2e-63
AT2G45430 222 / 5e-71
AT-hook motif nuclear-localized protein 22 (.1)
Potri.002G149300
199 / 1e-62
AT2G45430 226 / 1e-72
AT-hook motif nuclear-localized protein 22 (.1)
Potri.003G116900
184 / 1e-56
AT4G22810 258 / 5e-85
Predicted AT-hook DNA-binding family protein (.1)
Potri.001G115200
182 / 1e-55
AT4G22810 254 / 1e-83
Predicted AT-hook DNA-binding family protein (.1)
Potri.005G257200
164 / 7e-49
AT1G76500 192 / 2e-59
SUPPRESSOR OF PHYB-4#3, AT-hook motif nuclear-localized protein 29, Predicted AT-hook DNA-binding family protein (.1)
Potri.010G074201
156 / 6e-46
AT4G14465 256 / 3e-85
AT-hook motif nuclear-localized protein 20 (.1)
Potri.010G200100
157 / 2e-45
AT3G55560 209 / 2e-65
AT-hook motif nuclear-localized protein 15, AT-hook protein of GA feedback 2 (.1)
Potri.008G058700
154 / 1e-44
AT3G55560 192 / 5e-59
AT-hook motif nuclear-localized protein 15, AT-hook protein of GA feedback 2 (.1)
Potri.001G142800
153 / 1e-44
AT4G17800 300 / 6e-102
Predicted AT-hook DNA-binding family protein (.1)
Potri.002G003800
152 / 2e-44
AT1G76500 186 / 6e-57
SUPPRESSOR OF PHYB-4#3, AT-hook motif nuclear-localized protein 29, Predicted AT-hook DNA-binding family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0615
ALDC
PF03479
PCC
Plants and Prokaryotes Conserved (PCC) domain
Representative CDS sequence
>Lus10015862 pacid=23163288 polypeptide=Lus10015862 locus=Lus10015862.g ID=Lus10015862.BGIv1.0 annot-version=v1.0
ATGGATCATCATCCAGTTACAGCCGGCCGGGGACCCCTACTTCCACCTCCTTTCAACATTCGCTCAGATCTCCACCAAAACCAATTTCAACAGTTTCAGG
ATCAACGACAAAGCATGATGCTGATGATGATGAATCGCGGCCAGAAGCGAGACCACGACGAAGTCCTCAGCCCGGATAATAATTCCAACAACGAATTATT
AACGCCGTCCTCCTCCGGTCAAACCAAATCCAATATAAAGAGACCCAGAGGAAGACCGGCCGGTTCCAAGAACAAGCCCAAGCCGCCCGTAATAATCACC
CGAGACTCCGGAAACTCCCTCCGATGCCACGTCATCGAGATTGCCGACTCCGCCGACATTATACACAGCGTCTCCACTTTCGCCGCCCGCCGCCAGCGGG
GGGTCTGCATCCTCAGCGGCACCGGCGCGGTCAACAACGTCACCCTCCGACAGCCCGCCGGATCTGTCGTGAACCTCCAAGGCCGGTTCGAGATCCTCTC
CCTCTCAGGCTCTTTCTTACCTCCGCCTGCGCCGCCTGCTGCCTCCGGGTTGACTATCTACCTCACCGGCGGACAAGGCCAGGTGGTCGGCGGCAGCGTA
GTTGGACCCTTGCTTACATCCGGTCCGGTTGTTGTAATGGCTGCTTCGTTTGGGAATGCAGCTTATGAGAGGCTACCGCTTCAGGAAGAGGAGGATGAAG
TGGGCCAGCCACTGGCTGGTGCTGCTACGCCGCCGCAGCAGCAGGTGATTGGAGATGCTCCTCCCAATGGAATGTCTGATGAGAATGTGATGAGATCGGT
GCAAATGGGCGGCGGAGGAGGAGGAGGAGAAGGGTATTGGGGATCCAGCAGCGCCGCATCTGGTCCTTGTCCACCACCTCCGTTCTAG
AA sequence
>Lus10015862 pacid=23163288 polypeptide=Lus10015862 locus=Lus10015862.g ID=Lus10015862.BGIv1.0 annot-version=v1.0
MDHHPVTAGRGPLLPPPFNIRSDLHQNQFQQFQDQRQSMMLMMMNRGQKRDHDEVLSPDNNSNNELLTPSSSGQTKSNIKRPRGRPAGSKNKPKPPVIIT
RDSGNSLRCHVIEIADSADIIHSVSTFAARRQRGVCILSGTGAVNNVTLRQPAGSVVNLQGRFEILSLSGSFLPPPAPPAASGLTIYLTGGQGQVVGGSV
VGPLLTSGPVVVMAASFGNAAYERLPLQEEEDEVGQPLAGAATPPQQQVIGDAPPNGMSDENVMRSVQMGGGGGGGEGYWGSSSAASGPCPPPPF
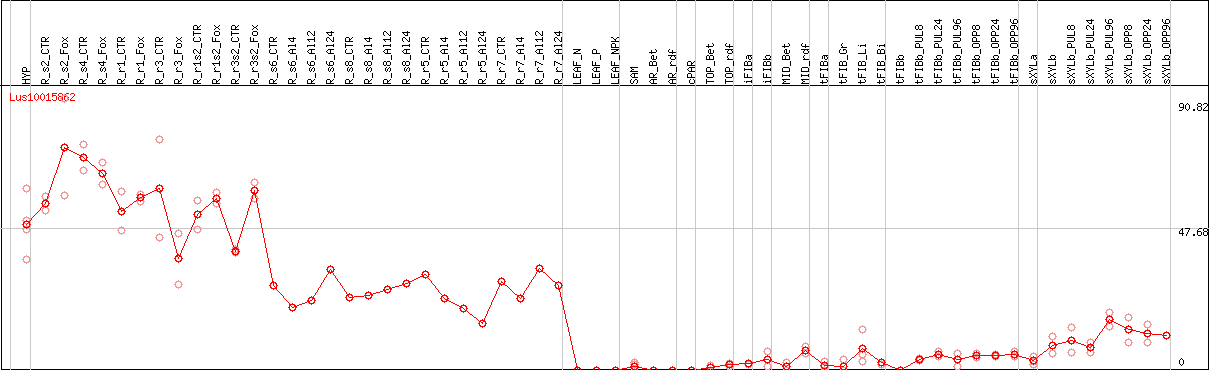
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015862 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.