External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G01970 110 / 7e-30
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G19520 80 / 9e-19
NFD5
NUCLEAR FUSION DEFECTIVE 5, pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
AT5G16640 52 / 5e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G50280 49 / 1e-07
EMB1006
embryo defective 1006, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G64580 46 / 1e-06
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G19890 45 / 1e-06
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT2G35130 45 / 2e-06
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT3G06430 44 / 4e-06
AtPPR2, EMB2750
pentatricopeptide repeat 2, embryo defective 2750, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63630 42 / 7e-06
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G63230 43 / 1e-05
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009299
154 / 3e-47
AT1G01970 430 / 1e-150
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10030026
81 / 1e-18
AT1G19520 660 / 0.0
NUCLEAR FUSION DEFECTIVE 5, pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
Lus10035298
79 / 3e-18
AT1G19520 658 / 0.0
NUCLEAR FUSION DEFECTIVE 5, pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
Lus10000428
54 / 1e-09
AT4G19890 407 / 1e-138
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10006083
54 / 1e-09
AT4G19890 844 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10042081
49 / 1e-07
AT5G50280 648 / 0.0
embryo defective 1006, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10022290
48 / 2e-07
AT2G31400 911 / 0.0
genomes uncoupled 1 (.1)
Lus10003657
45 / 2e-06
AT2G31400 902 / 0.0
genomes uncoupled 1 (.1)
Lus10023863
45 / 2e-06
AT5G12100 728 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G070500
107 / 1e-28
AT1G01970 469 / 7e-165
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G228200
89 / 1e-21
AT1G19520 649 / 0.0
NUCLEAR FUSION DEFECTIVE 5, pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
Potri.002G034900
69 / 1e-14
AT1G19520 296 / 2e-93
NUCLEAR FUSION DEFECTIVE 5, pentatricopeptide (PPR) repeat-containing protein (.1), pentatricopeptide (PPR) repeat-containing protein (.2)
Potri.015G121700
51 / 2e-08
AT4G19890 923 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.013G058900
45 / 2e-06
AT4G13650 1282 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G078900
45 / 2e-06
AT3G23020 1046 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.012G091800
44 / 3e-06
AT5G50280 845 / 0.0
embryo defective 1006, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G193900
43 / 9e-06
AT2G31400 1192 / 0.0
genomes uncoupled 1 (.1)
Potri.012G123600
42 / 3e-05
AT2G35130 852 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.005G074000
42 / 3e-05
AT4G39952 892 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10015864 pacid=23163311 polypeptide=Lus10015864 locus=Lus10015864.g ID=Lus10015864.BGIv1.0 annot-version=v1.0
ATGACTAAACAGTTTTTAGATCAGACTGATAGAGTAAAAGACTTTGATCTGTCTGACAAATTCTTCCCCTTGGGTTCAATGCTTGTTTATCAGGTGGCAA
AATTTGCTTTACCGGAAGAATCTTTCGAGGCGAATATCCGGGACTATACAAAGCTAATCCACTACTATGCGAAGAAACACCAGCTCGAAGATGCCGAAAG
GATCTTCTTATCAATGAAGGAAAGAGGCTTAGTTTACGATCAAGTAACAGTGACAGCCATGGTTCACATGTACAGCAAGGCAGGCTACCACAATAAGGCT
GAGGAAGCATTCTGCCGGCTCGCTCTGCTGGGCTAG
AA sequence
>Lus10015864 pacid=23163311 polypeptide=Lus10015864 locus=Lus10015864.g ID=Lus10015864.BGIv1.0 annot-version=v1.0
MTKQFLDQTDRVKDFDLSDKFFPLGSMLVYQVAKFALPEESFEANIRDYTKLIHYYAKKHQLEDAERIFLSMKERGLVYDQVTVTAMVHMYSKAGYHNKA
EEAFCRLALLG
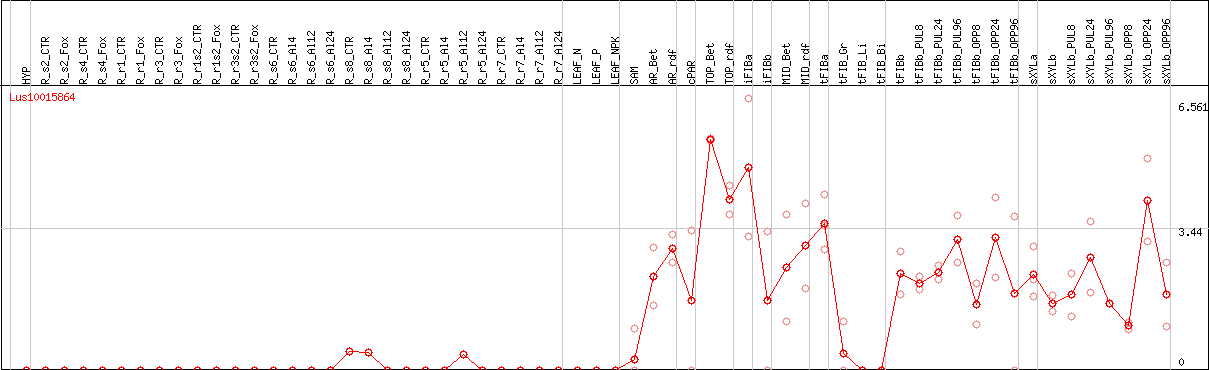
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015864 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.