External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45140 260 / 5e-88
PVA12
plant VAP homolog 12 (.1)
AT3G60600 246 / 6e-82
(AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, (AT)VAP, VAP27-1, VAP27, (AT)VAP, (AT)VAP, (A
VAMP/SYNAPTOBREVIN-ASSOCIATED PROTEIN 27-1, vesicle associated protein (.1.2.3)
AT4G00170 195 / 3e-62
Plant VAMP (vesicle-associated membrane protein) family protein (.1)
AT1G51270 190 / 1e-57
structural molecules;transmembrane receptors;structural molecules (.1.2.3.4)
AT2G23830 154 / 2e-47
PapD-like superfamily protein (.1)
AT1G08820 138 / 1e-38
VAP27-2
vamp/synaptobrevin-associated protein 27-2 (.1.2)
AT5G47180 131 / 2e-37
Plant VAMP (vesicle-associated membrane protein) family protein (.1), Plant VAMP (vesicle-associated membrane protein) family protein (.2)
AT5G54110 45 / 2e-05
ATMAMI
membrane-associated mannitol-induced (.1)
AT4G05060 44 / 5e-05
PapD-like superfamily protein (.1)
AT4G21450 43 / 0.0001
PapD-like superfamily protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009279
398 / 1e-127
AT4G00060 1034 / 0.0
maternal effect embryo arrest 44, Nucleotidyltransferase family protein (.1)
Lus10008512
227 / 1e-74
AT2G45140 296 / 7e-102
plant VAP homolog 12 (.1)
Lus10007463
227 / 4e-74
AT3G60600 306 / 4e-105
VAMP/SYNAPTOBREVIN-ASSOCIATED PROTEIN 27-1, vesicle associated protein (.1.2.3)
Lus10028937
225 / 7e-71
AT3G60600 305 / 2e-101
VAMP/SYNAPTOBREVIN-ASSOCIATED PROTEIN 27-1, vesicle associated protein (.1.2.3)
Lus10010360
189 / 8e-60
AT4G00170 311 / 5e-108
Plant VAMP (vesicle-associated membrane protein) family protein (.1)
Lus10036495
188 / 2e-59
AT4G00170 316 / 4e-110
Plant VAMP (vesicle-associated membrane protein) family protein (.1)
Lus10033837
129 / 2e-34
AT1G08820 256 / 2e-80
vamp/synaptobrevin-associated protein 27-2 (.1.2)
Lus10000591
120 / 4e-33
AT5G47180 277 / 5e-95
Plant VAMP (vesicle-associated membrane protein) family protein (.1), Plant VAMP (vesicle-associated membrane protein) family protein (.2)
Lus10000032
114 / 3e-32
AT2G45140 148 / 7e-46
plant VAP homolog 12 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0556
PapD-like
PF00635
Motile_Sperm
MSP (Major sperm protein) domain
Representative CDS sequence
>Lus10015885 pacid=23163295 polypeptide=Lus10015885 locus=Lus10015885.g ID=Lus10015885.BGIv1.0 annot-version=v1.0
ATGAGCTCCGACGGCCTCCTCACTATCCAGCCTCAAGATCTGCAGTTTCCATTCGAATTGAAGAAGCAGATCTCCAGCTCGCTGCAGCTATCCAACAATA
CCGACAATTATGTGGCGTTCAAGGTTAAGACGACGAATCCGAAGAAGTACTGTGTGAGGCCTAACACTGGCGTTGTGCAGCCTCGGTCTGCATGTGATGT
CATAGTTACTATGCAGGCACAAAAGGAGGCTCCTGTTGATATGTTGCAGTGCAAAGACAAGTTTTTATTACAGAGCGTTGTGACGAGTCCTGGGGCTACC
GCCAAGGATGTCTCCGTAGAGATGTTCAATAAGGAGGCAGGGCATCATGTCGAGGAGAACAAGTTGAGGGTTGTCTATGTTGCTCCACCAAGACCACCAT
CACCAGTTCGTGAAGGATCTGAGGAAGGTGAAGAACCTATAGCTTCTCTTTCCGACAATGGCGGTGTGCATCCAGTCGAGCATACATCGGTTTCCAAGCC
GGAGTTTGGTCGTAGAGTTGAGCCCCGTGATAATTCCTTGGAGGCACGTGCTCTTATCTCCAAGCTAACTGACGAGAAAGATTCTGTTGTCAAGCAAAAC
AAGCTGTTACAGAGGGAACTGGAGCAACTACGGAATCAAACAAGCAGAAACCGCATGGGCATCCCGATCACATATGTGGTTTTCGTGGCCCTTCTAGCAA
TCCTTTTGGGGTACTTGCTGAAAAGGGCATGA
AA sequence
>Lus10015885 pacid=23163295 polypeptide=Lus10015885 locus=Lus10015885.g ID=Lus10015885.BGIv1.0 annot-version=v1.0
MSSDGLLTIQPQDLQFPFELKKQISSSLQLSNNTDNYVAFKVKTTNPKKYCVRPNTGVVQPRSACDVIVTMQAQKEAPVDMLQCKDKFLLQSVVTSPGAT
AKDVSVEMFNKEAGHHVEENKLRVVYVAPPRPPSPVREGSEEGEEPIASLSDNGGVHPVEHTSVSKPEFGRRVEPRDNSLEARALISKLTDEKDSVVKQN
KLLQRELEQLRNQTSRNRMGIPITYVVFVALLAILLGYLLKRA
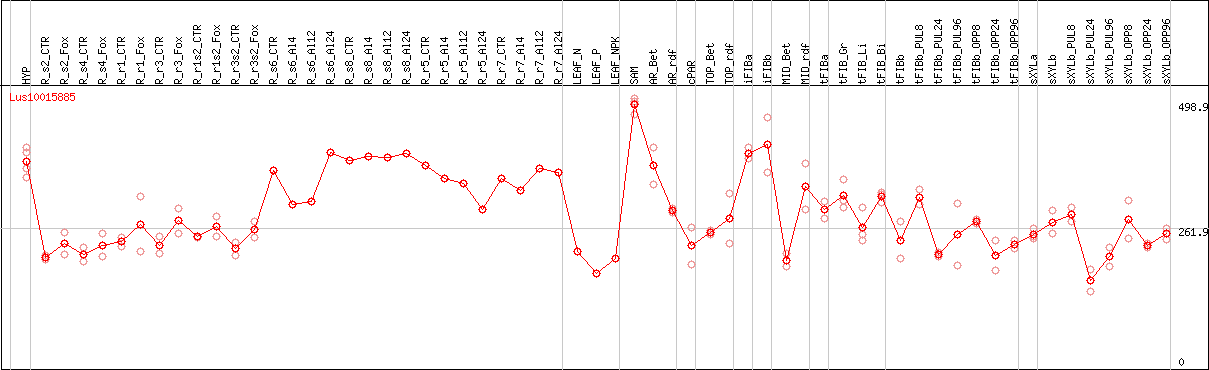
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015885 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.