External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G22500 154 / 5e-45
UCP5, ATPUMP5, DIC1
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
AT4G24570 150 / 1e-43
DIC2
dicarboxylate carrier 2 (.1)
AT5G09470 118 / 5e-31
DIC3
dicarboxylate carrier 3 (.1)
AT3G54110 83 / 4e-18
ATUCP1, UCP1, UCP2, UCP, ATPUMP1
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
AT5G58970 72 / 1e-14
ATUCP2
uncoupling protein 2 (.1.2)
AT1G14140 72 / 2e-14
Mitochondrial substrate carrier family protein (.1)
AT5G19760 62 / 5e-11
Mitochondrial substrate carrier family protein (.1)
AT4G03115 61 / 2e-10
Mitochondrial substrate carrier family protein (.1)
AT1G34065 51 / 3e-07
SAMC2
S-adenosylmethionine carrier 2 (.1)
AT5G66380 48 / 3e-06
ATFOLT1
folate transporter 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009271
190 / 5e-61
AT4G24570 138 / 7e-41
dicarboxylate carrier 2 (.1)
Lus10009270
164 / 2e-51
AT2G22500 204 / 8e-67
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Lus10025725
154 / 1e-45
AT2G22500 310 / 2e-106
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Lus10035936
147 / 6e-43
AT4G24570 209 / 5e-67
dicarboxylate carrier 2 (.1)
Lus10024540
142 / 4e-38
AT5G09470 91 / 3e-19
dicarboxylate carrier 3 (.1)
Lus10015570
98 / 5e-26
AT4G24570 100 / 5e-27
dicarboxylate carrier 2 (.1)
Lus10043336
91 / 4e-23
AT2G22500 74 / 4e-17
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Lus10027755
76 / 1e-15
AT3G54110 515 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10017187
74 / 3e-15
AT3G54110 501 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G157300
184 / 2e-56
AT2G22500 412 / 2e-145
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.002G104400
175 / 5e-53
AT2G22500 427 / 1e-151
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G047800
159 / 8e-47
AT4G24570 435 / 2e-154
dicarboxylate carrier 2 (.1)
Potri.017G045400
155 / 1e-45
AT4G24570 355 / 4e-123
dicarboxylate carrier 2 (.1)
Potri.017G045150
155 / 4e-45
AT2G22500 457 / 3e-163
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.007G113000
154 / 6e-45
AT4G24570 404 / 4e-142
dicarboxylate carrier 2 (.1)
Potri.017G045200
149 / 6e-45
AT2G22500 286 / 3e-98
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G045332
154 / 7e-45
AT2G22500 410 / 1e-144
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G045100
153 / 2e-44
AT2G22500 451 / 6e-161
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.010G165900
78 / 2e-16
AT1G14140 447 / 1e-159
Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Lus10015897 pacid=23163316 polypeptide=Lus10015897 locus=Lus10015897.g ID=Lus10015897.BGIv1.0 annot-version=v1.0
ATGGGTGTTAAAGGATTCGCTGAAGGCGGAATCGCTTCAATCATCGCCGGTTGCTCCACCCACCCGCTGGATCTGATTAAGGTCCGAATGCAACTCCAGG
GCGAAGTCTCTGCTCCTAAACCTATTCATAACCTCCGCCCCTCTTTCGCTTTCCATCCCGGCACTGGCGCCGCTGTCTCCGCTATCCACCTCCCTCCTAC
TCCACCTCCTTCAGTCGGTCCCGTCGCCGTCGGAATGAACATACTCAAGACCGAAGGGGTCGCCGCTCTGTTCTCCGGCGTCTCCGCCACCGTCCTCCGC
CAGACTCTCTACTCCACCACAAGGATGGGGCTCTACGACGTCCTCAAGACCAAGTGGACCAATCCTCAGACGGGGAACATGCCCCTCGTCAGCAAAATCA
CCGCCGGACTAATTGCCGGCGGGATCGGAGCGGCGGGGGGGGGGGGGGCCGGCNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNTTTTTGTGGCGTCGGTG
GCGTCGAATCCGTTGGATGTGATTAAGACTAGGGTAATGAATAGGGCTGTGGAGGCTGGGAAGGCGGCGCCGTACAGCGGAGCGATGGATTGTGCGATTA
AGACGGTGAAGGCGGAAGGGCCGATGGCGTTGTATAAGGGATTCATCCCTACGATATCGAGGCAAGGTCCTTTCACTGTTGTGCTGTTTGTGACGCTTGA
GCAGGTGAGGAAGCTGCTCAAGGATTTCTGA
AA sequence
>Lus10015897 pacid=23163316 polypeptide=Lus10015897 locus=Lus10015897.g ID=Lus10015897.BGIv1.0 annot-version=v1.0
MGVKGFAEGGIASIIAGCSTHPLDLIKVRMQLQGEVSAPKPIHNLRPSFAFHPGTGAAVSAIHLPPTPPPSVGPVAVGMNILKTEGVAALFSGVSATVLR
QTLYSTTRMGLYDVLKTKWTNPQTGNMPLVSKITAGLIAGGIGAAGGGGAGXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXFVASV
ASNPLDVIKTRVMNRAVEAGKAAPYSGAMDCAIKTVKAEGPMALYKGFIPTISRQGPFTVVLFVTLEQVRKLLKDF
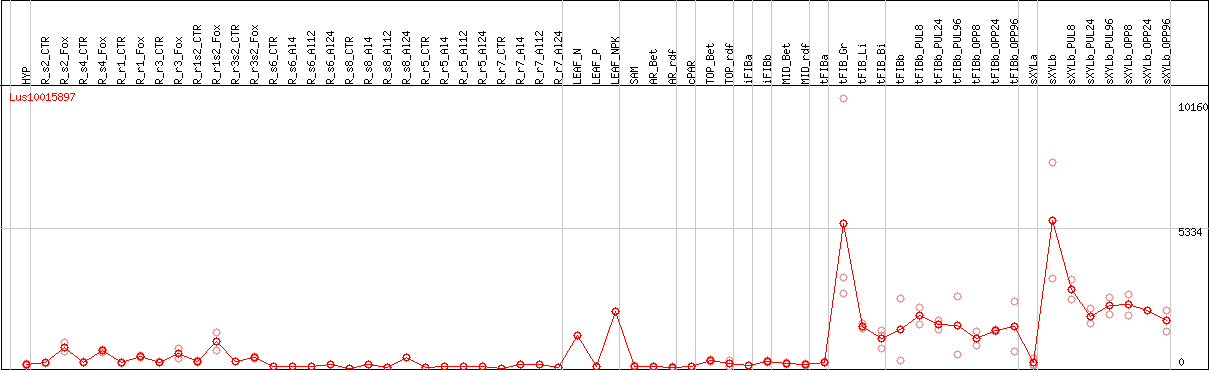
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10015897 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.