Lus10016103 [FLAX]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||
| PFAM info | |||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016103 pacid=23171816 polypeptide=Lus10016103 locus=Lus10016103.g ID=Lus10016103.BGIv1.0 annot-version=v1.0
ATGGCAGTAGAAACTAGGGCTAGCAGTGCATCTAAAGTAGACGAAAGCACAAATTCGAGAACTCTGAGGTCCTCATCGCGGTTAGCTAGGGAGAAATTGT
TGAGGAGTGATACAAAGGCTAGTCCTTTGAACGTTCGGAAGTCGGTTAAGAGGAAACACAAGACCACTGTAAGGAAGTCTGAGAAGAGTAAGGAGGAATC
TGTACCGGGTTCTTCGAGGTTGAAGAAGCCAAGGATGGCAAAGTTGACTAGTAAAACCAAAGAGGTGAACAACAGTGAAGTGGAGTTGTCCAAGGGGAAG
ACGAGATATAATGCTCTTTCGTACAAAGCACTTTTCACGAAGAAGAAGAAGAAACAGAGAACTAATGTAGAAGCTGATTCGGTTAGTCCAACAACTTCAA
AGACGGTTGATGATCAGTCTAGCAAAGCTTCTGCGACAGAAGATCGTGCACAACCGCAGCTTGGGGATGACATAGTGCTCGAGGCTGGATCTGATAATGA
GATTGTTTCGGTGAGCATTGACAAATCCATGGCGTCGAAAAGAAAAAGAATAGAAGCCGGTTTAGTTGATGAGACTGCATTGGTGTCAAGTAAAGATGCC
GATATCTCGCCCACTGATGCTGCTGCTGCAGGGGAAGATCAATCTCGAGTGAAGCGTGTTTCGAATGATTTTGCTGGAGGCCTTAAGAGGCAGAGGGTTA
ATGATGAAGGAAAACAGCAAGAAGTTTGCTATTTGAACATCGAGTCAGATTGGGGCGATGCCTGTCACAAGTCGAAGTTAGATGGGAGTAATCCATCAGA
AGTTGAAGTAATTATTATCGACAAGTCCATTGAACCCGATTGGCTGCAGAATTCACAGAGCAATCTACATTGTGATCTGAAGCCGAAAATAGCATCGCTT
TGTGAAGTTCTAAGATTCCCGGATGATGTCAAATGTATAGTTGTAAGCTTTCTCGACTACATAGTGAACAATCAATATCATCAACTGGCTTCAAAGCGAG
AATCTGGATTGCAGGCTTTTCAGATTGCTTTGTGTTGGACTGCTGCTTCTTTGCTGAAGTACAAGATTGACCATATGGAATCTCTGGGACTTGCTAAGGA
GAGATTAAACTTCGGATGCAAGAAAGAACATGCTGACTTTGAGTATTCGATGTTACGATGTTTGAAGAAATTATTTCTATCCCATCATCAGCGAAGTAAA
TATGTTTCTGACGACCAGTTGCATGCTAGTCCTTCCAACAGATCAATTTCGTCAAGTCCTAAAGAAAACGGGGATGGAGTTCAAATCCAGCAACCACTTG
AATTGGCACATCAAGATTTTTCTAGAAGTATCAAAGACATCCACAAGAAGTGTGATAAGCAGTTTAGAAAACTGGTGGAGCAGCAAGCACAACAGAGAGA
GGACTTGCTTAAAAAGTACGAGGATCAAAAGGCTCAGCTTGAGAAGAAGCAGGAAATGGAGGCAGCTGTCATTTGCCTTTACAGCCCTGATTCAATGAGA
CCAGACAAGCTCGAGTTCATGGAAGCTGAATATGCCGAGAAATTCAAGAAACTTGAACGTGAGAGGGTCACAGAACTTAAAAATCTCGAGGCAATGCAGT
CTGCTTCAAGGAAAACATTGGAAGAGAATAAACTGAAATGGGTTGGAGGATTGGAAACCTGGGCGAAGGAAGAATTAATGCCTAGGCTGCCTCGAAAAGA
TACTGGGCAGACTTCAGGAACTGGTGTTTCTTCTGATGTTCATCCCAAGCAAGTTACTTGTCAGGCAAGTTTAACTGACCGTGGAGAAGAAGAAACAACC
CAAATGGTGAAGTCTGCTGCTGAAGTGATTCATGAAACACCATCTACTACGGTGCATGTTGAATCAACACAAGGTGAGGTTACTTCGGAACCATTGGCAT
CTACTGGTGAATCTGAAAGACCTCTGACAGATCAACAAGTCAAGGATAGTTCTCTTCAAAGTCGAGACGAGGAACCGGGTTCCACAATACCTGAAACAGC
TACCATTACTGGCTTTGGCTTAATAGATAATAATGCTGATGGAGAGATTGAAGAAGGCGAAATTTCCGATGCTAAGGAAGGAGACGGTGAGATTTTATCT
CCTGAATCTGAAAGACCTCTGACAGACCAACTTATGAAGCATAGTTCTCTTCAAAATCAAGACACCGGTACAGAGAAAGTTGGATTAGGGGTTCTAGACT
TACCACTTGGAGTTGACGCTGGTCTTCAAGGTGAACCGGTTTCTGAAATACCTGAAACAGATACCACTACGACTACTTGCTTAGTAGGTAATAATGCTGA
CGGAGAGATTGAAGAAGGTGAAATTTCTGGTGCTGAGGAAGGAGGCGGTGAGATTTTATCTTCTGAATCTGAAAGACCCATGACAGACCAACAAATGAAG
GATAGTTCTCTTCAAAGTCAAGACACCGGTACAGAGAATGTTGGATTAGGGGTTCCAGACTTACCACTTGGAGTTGACACTGGTCTTCAAGGTGAACCGG
TTTCTGCAATACCTGAAACAGCTACCACTACGACTACTTGCTTAGTAGATAATAATGTTGACGGAGAGATTGAAGAAGGTGAAATTTCTGGTACTGAGGA
AGGAGGTGGTGAGATTTTATCTGCTGAATCTGAAAGACCCGTGACAAAAAAACAAATGAAGGAAAGTTATCTTCAAAGTCAAGACACCGGTACAGAGAAT
GTTGGATTAGAAGTACTGGATTTGTGTCCCGAAGTTGATAAATTGGACGATCAGAATGATGGAAGCACAGAGCAGGATGTAGCAGCATCATCTATCCCTA
GTCGAAAAGACAGTAGCAAAGAACTTTTTGCGACAGATTCTCCTTTGATACAACCTGAAACCTCCATAACAGGAGTCTCTTCTAGTAGAATAGATCAGCA
GGATCCAGGGAGGATTCTTGCAGGCGTGTCTAATGTAAGCACTGATCTGGCAGATGGAAGCAGTCATCAGGAGGGAACGGTTACCGGAGTTCACAAGGAA
GCAACTGCAGAAGTTGTGGGTGTTGGGGTGAAAAATCTAGCCTCTGCTGAAGTATTAGAAGATCATAATGATGGAATCCTAATGCACGATCCGAAAGCTC
CTGTTACCAACCAAAGTGATCAATCCGAACAAATTGTTCACTCGATTTCAACTCCGGTGCAGGATGGTGTAGTTTCATGTGTCTCAGGCTCTGACACAAA
TACAGAAGTAGCAGATGTGGCAGACTTATGCATAGAGAACAATGGAACCAGAGTTGGTCAAGAGAGTCGTTCTTCTGAAGATACTGAAGTGCATCATCAG
GATGATCAATCCCAACATATTGTTCAAACAAGTTCAGCGCCAGTATCTATTGGAGTTGGTCAAGCGTATGGATCTTCTGAAGACACGGCAGTGCATCACC
AGGATGCAACCGCGATTCCACTAGTGGCCGAGACAGCCACTACTGCTAGAATAGCCATGGGGAACTCTTGTACAAAGAACAACATATCTTCTCAGAATCT
AGGAGCTTCTACTGCCAGGAGAGATGATGGTTCAGGAGAAAGTTCTATGACAAATTCAACCTCAATGCAGCCAGAAGTTGCGGTCTCTGCACAATCCAGC
TCATCGCTCCCGAATCAGCCTGTTCAAGATGAACTACGTCGACAGCTGCCTGCATCTACTAGAGCAAATGTCTCGGAAACAGTAACTGCACCTTCTACTT
CAGCAAACCGTGCCCGCCTTCCACAACCTGCACCTGTTTACCGAATGCCTCTCCCTTTTTTGGGCCCCGATCCTCTCCAGAACGAATTGGATAGAGTATC
CAGAGTTGTAGATCAGATAGTCAATATACACGAAAAGGAGAAGCTGCAGTTGAGATTCGACTGCGAAAAGGAAATGGAGGAAGTAGTTGCACAGATACGT
CGGAAGTACGAAGCTATAGCCAAGGAGTCAGAAACTGAATTCGCGCGAAACAAGTCGAAACTGGATGCAAAGCACTATAAAATCCGGATGAACAAGCTCC
TGGCTGATGTGTTCAGGTGCAAATGCGTGGACAGTCGAGCAAACAAATCTGTTGGAGTTGTGCAAGATGCTGCTGCTAGTTTTATGCAGCAATTCCAGAA
TCTATCTTCGCAGCCTACGCAACCATACACAGGACAGCGATTCTCAACTGCTGCTTCTCGGCCTTTGCAACCTCCACCAAATGGGTTAGTAGCGAATCAT
CAACCGACCGGTTCGCCTCATCTATCAATCTCTCGAGCACATCCTGTCGCTCATCGCTTCCCAACACCAACTCATATCCCTCAATCTTCAAGGCCACTCA
ACATCAGCACCCTCACCCCTACTACAGCAAGCAGCAATCTCCATCAATTCGGTAGCCAGTTCCGAGCGCCTGCTCCACACATGCAAGCCGTTCGAGCTAT
GACATCACATGGATTGCAAAGCCATCTATGTCCTCATAGTGTTGTCCGTCCCTCTACTGTAGTCTCCATAACTGCATCCTCGCCAGTTCCTCCTGCTTTG
GTCAATGCACCTGCAATCGTGCCTCCTGAAGTTCGGCCAAATGGTGCGCAAGGGAACTCGACGCAGCAAAACGAGGTCGTTTATTTATCAGACGATGATT
GA
|
||||||||||||||||||||
|
AA sequence
|
>Lus10016103 pacid=23171816 polypeptide=Lus10016103 locus=Lus10016103.g ID=Lus10016103.BGIv1.0 annot-version=v1.0
MAVETRASSASKVDESTNSRTLRSSSRLAREKLLRSDTKASPLNVRKSVKRKHKTTVRKSEKSKEESVPGSSRLKKPRMAKLTSKTKEVNNSEVELSKGK
TRYNALSYKALFTKKKKKQRTNVEADSVSPTTSKTVDDQSSKASATEDRAQPQLGDDIVLEAGSDNEIVSVSIDKSMASKRKRIEAGLVDETALVSSKDA
DISPTDAAAAGEDQSRVKRVSNDFAGGLKRQRVNDEGKQQEVCYLNIESDWGDACHKSKLDGSNPSEVEVIIIDKSIEPDWLQNSQSNLHCDLKPKIASL
CEVLRFPDDVKCIVVSFLDYIVNNQYHQLASKRESGLQAFQIALCWTAASLLKYKIDHMESLGLAKERLNFGCKKEHADFEYSMLRCLKKLFLSHHQRSK
YVSDDQLHASPSNRSISSSPKENGDGVQIQQPLELAHQDFSRSIKDIHKKCDKQFRKLVEQQAQQREDLLKKYEDQKAQLEKKQEMEAAVICLYSPDSMR
PDKLEFMEAEYAEKFKKLERERVTELKNLEAMQSASRKTLEENKLKWVGGLETWAKEELMPRLPRKDTGQTSGTGVSSDVHPKQVTCQASLTDRGEEETT
QMVKSAAEVIHETPSTTVHVESTQGEVTSEPLASTGESERPLTDQQVKDSSLQSRDEEPGSTIPETATITGFGLIDNNADGEIEEGEISDAKEGDGEILS
PESERPLTDQLMKHSSLQNQDTGTEKVGLGVLDLPLGVDAGLQGEPVSEIPETDTTTTTCLVGNNADGEIEEGEISGAEEGGGEILSSESERPMTDQQMK
DSSLQSQDTGTENVGLGVPDLPLGVDTGLQGEPVSAIPETATTTTTCLVDNNVDGEIEEGEISGTEEGGGEILSAESERPVTKKQMKESYLQSQDTGTEN
VGLEVLDLCPEVDKLDDQNDGSTEQDVAASSIPSRKDSSKELFATDSPLIQPETSITGVSSSRIDQQDPGRILAGVSNVSTDLADGSSHQEGTVTGVHKE
ATAEVVGVGVKNLASAEVLEDHNDGILMHDPKAPVTNQSDQSEQIVHSISTPVQDGVVSCVSGSDTNTEVADVADLCIENNGTRVGQESRSSEDTEVHHQ
DDQSQHIVQTSSAPVSIGVGQAYGSSEDTAVHHQDATAIPLVAETATTARIAMGNSCTKNNISSQNLGASTARRDDGSGESSMTNSTSMQPEVAVSAQSS
SSLPNQPVQDELRRQLPASTRANVSETVTAPSTSANRARLPQPAPVYRMPLPFLGPDPLQNELDRVSRVVDQIVNIHEKEKLQLRFDCEKEMEEVVAQIR
RKYEAIAKESETEFARNKSKLDAKHYKIRMNKLLADVFRCKCVDSRANKSVGVVQDAAASFMQQFQNLSSQPTQPYTGQRFSTAASRPLQPPPNGLVANH
QPTGSPHLSISRAHPVAHRFPTPTHIPQSSRPLNISTLTPTTASSNLHQFGSQFRAPAPHMQAVRAMTSHGLQSHLCPHSVVRPSTVVSITASSPVPPAL
VNAPAIVPPEVRPNGAQGNSTQQNEVVYLSDDD
|
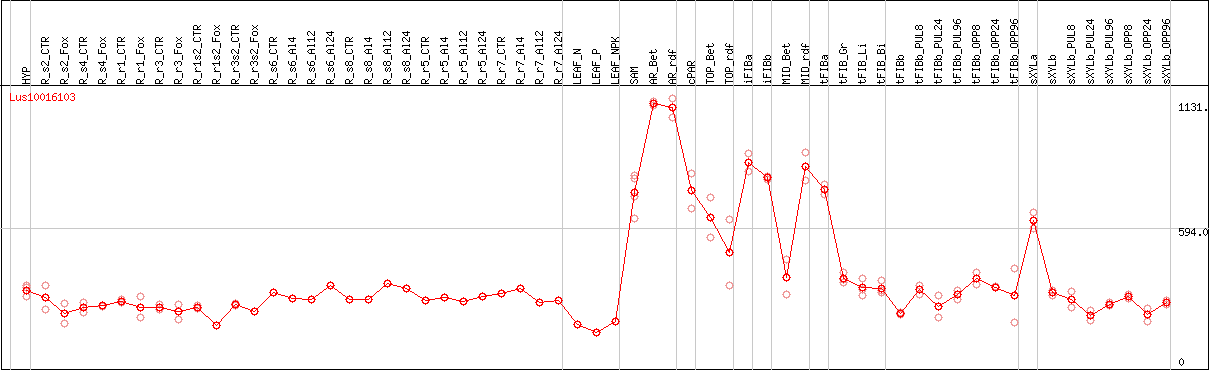
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016103 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
