External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G66654 221 / 1e-72
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
AT2G47320 177 / 9e-56
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT3G44600 89 / 4e-20
AtCYP71, CYP71
cyclophilin 71, cyclophilin71 (.1)
AT2G36130 82 / 2e-19
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT1G01940 75 / 9e-17
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT5G67530 74 / 6e-15
ATPUB49
plant U-box 49 (.1)
AT2G15790 66 / 2e-12
CYP40, SQN
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
AT5G13120 62 / 3e-11
Pnsl5, ATCYP20-2
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
AT2G29960 61 / 4e-11
CYP19-4, ATCYP5, CYP5
CYCLOPHILIN 19-4, ARABIDOPSIS THALIANA CYCLOPHILIN 5, cyclophilin 5 (.1.2)
AT4G33060 62 / 5e-11
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012005
380 / 1e-134
AT3G66654 259 / 5e-87
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Lus10021052
319 / 2e-111
AT3G66654 282 / 1e-96
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Lus10004174
316 / 4e-110
AT3G66654 272 / 8e-93
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Lus10006271
92 / 2e-21
AT3G44600 1110 / 0.0
cyclophilin 71, cyclophilin71 (.1)
Lus10020573
92 / 2e-21
AT3G44600 1115 / 0.0
cyclophilin 71, cyclophilin71 (.1)
Lus10004731
76 / 1e-15
AT5G67530 916 / 0.0
plant U-box 49 (.1)
Lus10007794
74 / 6e-15
AT5G67530 868 / 0.0
plant U-box 49 (.1)
Lus10011330
71 / 1e-14
AT5G13120 336 / 2e-117
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Lus10003138
71 / 2e-14
AT5G13120 333 / 2e-115
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G105800
233 / 1e-77
AT3G66654 306 / 2e-106
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Potri.010G144500
222 / 3e-73
AT3G66654 311 / 2e-108
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1.2.3)
Potri.002G194250
100 / 9e-26
AT2G47320 125 / 1e-35
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.009G146400
89 / 3e-20
AT3G44600 1095 / 0.0
cyclophilin 71, cyclophilin71 (.1)
Potri.016G075600
81 / 8e-19
AT2G36130 278 / 3e-97
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.002G149700
74 / 3e-16
AT1G01940 293 / 3e-103
Cyclophilin-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.005G148900
75 / 3e-15
AT5G67530 806 / 0.0
plant U-box 49 (.1)
Potri.001G060200
66 / 1e-12
AT5G13120 297 / 7e-102
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Potri.003G167700
64 / 3e-12
AT5G13120 277 / 5e-94
Photosynthetic NDH subcomplex L 5, ARABIDOPSIS THALIANA CYCLOPHILIN 20-2, cyclophilin 20-2 (.1.2)
Potri.005G135500
64 / 7e-12
AT2G15790 553 / 0.0
SQUINT, CYCLOPHILIN 40, peptidyl-prolyl cis-trans isomerase / cyclophilin-40 (CYP40) / rotamase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0475
Cyclophil-like
PF00160
Pro_isomerase
Cyclophilin type peptidyl-prolyl cis-trans isomerase/CLD
Representative CDS sequence
>Lus10016265 pacid=23142679 polypeptide=Lus10016265 locus=Lus10016265.g ID=Lus10016265.BGIv1.0 annot-version=v1.0
ATGGCGAGGATCAAACCTCAAGCTCTGTTGCAACAGAGCAAGAAGAAGAAAGCGCCGAGCCGTATAAGCGCCACAACTGTCATAATCTGCAGTTTGATCG
CTGTTTTGGCTTTGTTCTTCCTCTACTCTACTTACAAGCACCGTTCCATCAGGTCTAGCATCGAGAATTCCAGCTCCGAGGATGGTCGTAAGCTTCAGGA
ATTGGTCGAATCTGATATCCCTGGATATGCGATCTTTACTACATCAAAAGGGCCTCTGACTGTGGAGCTTTTAAAGGATAGCTCTCCTCAAATCATCGAG
AAGTTCATTGACTTATGTAAGAAAGGGCATTTCATGGGGATGCAATTTCTTCGCGTGGTAAAGCACTATGTGATTCAAGCAGGGAACACTGATAATCTGG
GCAATACAGAGGATTGGACTGTGAAAGGAAACGATAATGCACAACTTCATTCAAGCGTGAAACACGAATCGTTAATGGTTGGAACTTCGAAGGGTAAACA
TGACGACAAAGGATTCGACCTTTTCATCACAACTGCACCAATGCCAGACCTTAACGAGAAGCTGGTGGTTTTTGGACGAGTTGTCAAGGGAGAAGATGTA
GTTCAGGAAATCGAAGAGGTGGATACCGACGAACACTACCAGCCTAAGAGTCGAATAGAAATCACTAGTGTCGCTCTGCAACAGAAGGCTTAG
AA sequence
>Lus10016265 pacid=23142679 polypeptide=Lus10016265 locus=Lus10016265.g ID=Lus10016265.BGIv1.0 annot-version=v1.0
MARIKPQALLQQSKKKKAPSRISATTVIICSLIAVLALFFLYSTYKHRSIRSSIENSSSEDGRKLQELVESDIPGYAIFTTSKGPLTVELLKDSSPQIIE
KFIDLCKKGHFMGMQFLRVVKHYVIQAGNTDNLGNTEDWTVKGNDNAQLHSSVKHESLMVGTSKGKHDDKGFDLFITTAPMPDLNEKLVVFGRVVKGEDV
VQEIEEVDTDEHYQPKSRIEITSVALQQKA
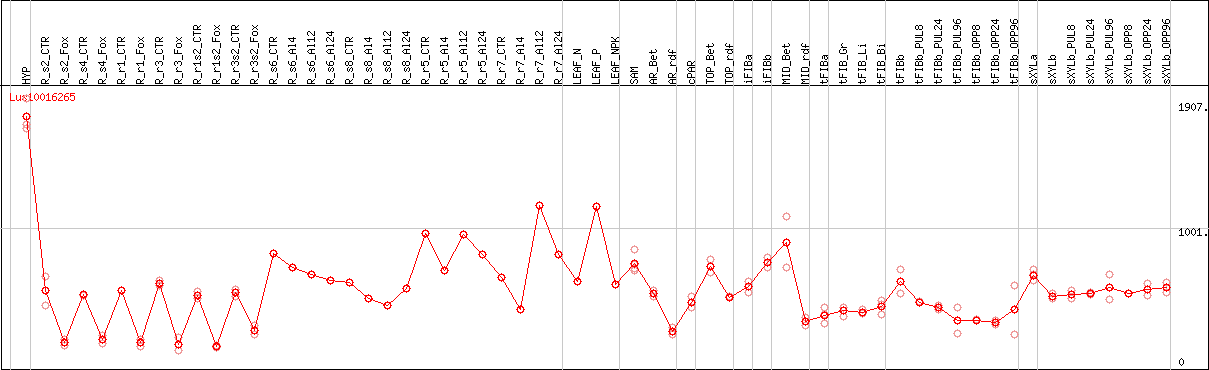
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016265 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.