Lus10016397 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016397 pacid=23161874 polypeptide=Lus10016397 locus=Lus10016397.g ID=Lus10016397.BGIv1.0 annot-version=v1.0
ATGGTGGCGAGCAAGAAGGCACCACTGATGACGAGCAACAGCGCCAACGTCGGCGCTGATCCGTCGCCTGACAACTTCCCCCAGAAGACGATCGAAGAGA
TGTACCAGAAGAAGACGCAACTGGAGCACATCCTGCTCCGGCCGGACACCTACATCGGCTCGATCGAGAAGCACACGCAGATGCTCTGGGTTTACGAAGA
TGGCAAGATGGTGAATCGCTCCGTCACGTATGTTCCGGGGCTGTACAAGATCTTCGACGAGATCTTGGTGAATGCTGCCGATAACAAGCAGAGGGATCCG
AAGATGAATTCGTTGAAGGTGTCTATTGACGTGGAGAACAACCTCGTCAGCGTTTACAACAACGGGGATGGAGTTCCTGTCGAAGTTCACAAGGAGGAAG
CCGTGTACGTTCCTGAGTTGATTTTTGGCCATTTGTTGACTAGTAGTAACTATGATGACAATGAGAAGAAGACGACTGGTGGTAGGAATGGTTATGGAGC
TAAGCTGACCAATATATTCTCTACCGAGTTTGTGATTGAGACTGCTGACGGCAAGAGGCTGAAGAAGTACAAGCAGGTATTCACTAACAACATGGGCACG
AAATCTGATCCTGAGATAAGCAATTGCAAGGCATCGGAAAGTTGGACAAAGGTTTCCTTCAAGCCTGATTTAGATAAGTTCAACATGACTCATCTTGAAG
AAGACGTGGTCGCCCTGATGAAGAAAAGGGTGCTTGACATGGCTGGCTGTTTGGGAAAAGGAGTGAAGGTTGAGTTGAACGGTATCCATATCCCTGTAAA
GTCATTCCAGGACTATGTGAATCTCTACCTGGATTCCACTTCTGAAAATGGCGTGCGTCGTATGAGCTTTTACGAAAAATGCGGGGACAGGTGGGAATAC
TGTGTCAGCATAACTGAAGGGCAATTCCAGCAGGTCAGCTTTGTCAATTCTATTGCCACAATCAAGGGTGGTACTCATGTAGACTATGTGACAAACCAGA
TCACAAGTTATGTGACTGAAGTTGTTAATAAGAAGCATAAAAATGCTCACCTGAAAGCCCATAATGTTAAGAACTATCTGTGGGTTTTTGTCAATACTCT
GATTGACAATCCTGCATTCGATTCTCAAACGAAGGAAACTCTAACACTACGCCAGAGCAGTTTCGGGTCTAGTTGTCAAGTGTCTGAGGACATTTTGAAG
AAAGTGGTGAAATCTGAGATTGTTGAAAATCTTCTCAACTGGGCACAATTCAAGCAGCAAAAAGACCTCAAGAAAACTGATGGGTCAAAAACATCCCGGA
TCAATGTTCCAAAGTTGGAAGATGCTAATGAGGCCGGAGGCAAGAATTCTGCGAAGTGCACCCTCATTTTGACTGAAGGAGATTCAGCCAAGGGCTTGGC
TGTTTCTGGGCTCTCTGTAGTTGGTCGTGACTACTATGGCGTGTTCCCACTAAGAGGTAAGTTGCTCAATGTCAGGGAAGCCACCGTTAAGCAGCTGAAT
GAGAACAAGGAAATTCAGTACTTGAAGAAGATATTAGGTTTAAGACAAGATAAAACGTACGAAGATGTTAAGTCATTGAGATATGGCCATATGATGATCA
TGACAGATCAGGATCACGACGGTTCTCATATTAAGGGATTACTGATCAATTTCATCCACTCTTTCTGGCCATCATTACTCAGAGTTCCTTCCTTCGTTGT
TGAGTTCATCACTCCCATTGTGAAGGCAACTAGTAAGAGAGGCCAAGTTCTGTCCTTCTATACGATGCCTGAGTACAAGACGTGGTCTGACAATCTTGGA
ACTGCAAAAAGCAACTGGTCCATCAAGTATTATAAGGGGTTGGGAACGAGTACAGACAAAGAAGGAAAGGAGTACTTCAGCGATCTTGATAAGCACAAGA
AAGATTTCATGTGGGTGGATGACAATGATGGAAATGCTATAGAGCTTGCTTTCAGCAAGAAGAAGATAGAAGCCAGGAAAGACTGGATTCGACAACATGA
GCCTGGAACTCATCTAGATCAGGCTGACAAGCTCATCAAATACAGCGATTTCATCAATAAGGAGCTTATATTGTTTTCAGTTGCCGATCTTCAAAGGTCT
ATCCCTTCCATGGTTGATGGACTGAAGCCAGGTCAAAGAAAAATCCTTTTTGCTGCTTTCAAGAAGAACCTTGTGAAAGAATTGAAGCTGTCTCAGTTCA
TTGGTTATGTATCTGAGCAAAGTGCTTACCATCACGGGGAGCAAAGTCTCAACAGTACAATTGTTGGGATGGCACAGAACTTTGTTGGTAGCAACAATAT
TAACCTCTTCGTACCAGCTGGAAATTTTGGCACTCGAGCCAGTGGCGGTAAAAATGCAGCAAGTGCAAGATACATTCACACTCGCCTTTCTCTTATAACG
AGGTTCCTTTACCCCAAAGACGATGACTGCATTCTGGACTACCTTGAAGATGATGGACAGTCCATTGAACCAACATGGTACATGCCAATAATACCTATGG
CTCTGGTCAATGGTAGTGAAGGAATTGGCACAGGATGGAGTACCTTTGTTCCTAACTACAACCCTAGAGATATTGTTGCTAACATAAGGCGGTTGTTGAA
CGAGGAAATGATGGAGCCCATGGAACCGTGGTACAAAGGCTTCAGAGGAACAGTCGTAAGAGGTGTATCAAAGGAATCTGGAGTCACCTACTATTGTAAT
GGAGTCATTGACGAAGTTGACGAGACAACTATCAAAGTAACAGAGCTGCCCATCCGTACTTGGACCGACAATTTCCGTGAATATCTTAAATCAGTGACGG
AAGGGCTCAAGGATGAGAATGGAAACGCTCCTAAGGATCCTTTGATCAAGGGTTATCATGAACAAGGTGATGACAATGAGTTTGAGTTCACCATTACCTT
GTCAGAGGAGAAGGTGCAACTTGCTAGGACCGAAGGTTTGCTGAAGAAGTTTAAGCTCACTAGCTCTATCAGCACAACTAACATGCATCTCTTCGATCCA
TCTGGGAAGATCAAGAAATATGACAATCCAGAACACGTGATTGAGGAATTCTTCCACTTGAGGCTTGAGTATTACGACAAGAGGAAGAAAACAAAATTAG
CTGCTCTTGAGCTAGAGCTACTAAGGTTGGAGAACAAAGTGAGGTTTATCCTGGCAGTGGTGAACGGCGAAATCATAGTGAACAACAGAAAGAGAGCTGA
TCTGATCCAAGAGCTTCACCAGAAGAGATTTACTCCGATGCCAATCAAGGCCAAGTCTTCTGTGACTTCTGATACAACTGAAGAATCTGAGGAAAATGAA
GACTCCGATCCTGTAAACCCTTCTGGAGTCCGAGCTAGCGATTACGACTACTTGTTGTCGATGGCCATTTCAACATTGACCTTCGAGAGAGTGCAGAAGC
TGCTTGAAGATAGGGACCGGTTCAACGGTGAGGTCGATAATCTGAGGAAAATGTCAGTCAAAGACTTGTGGATGCATGATCTTGACGCTTTCGAGAAGCA
GCTCGATGAGCTTGAAGCACAAGAGGAATTGGATAGGAAGCTGAAGCCGAAAAGTAAACAGACACGTGTTGTCAATGCTCCTAAGCCTAGGAACAGAGCG
AAACCTGTTCAGAAACCGAAAAGCGTAGAAGAAGATATGGAATCAAAGCTGAAACTTTCCTCTACTAGTTCAGCAATGGAAACTAATGAACAGCCAGAAC
CTGTTCCAGCTGCAAAACCAAAGGCAAAGGCCGCAGCTACGAAGAAAGCTCCTGCGAAAAAGGGAAAACAAGCAGCTTTGGTTGACAGTGACGACGATGA
AGTCGAGTCTCTGAAAGACCGCCTGAAAGCTTACAAGCTCGACGATTCTTCCCCGGAGAACTCTGCAGAAAAAAAGCAAGCCCCAAAGAAAAGAGGAGCG
AAGAAAGCGGTGATAGACCTCGACGACGAAGAGTCCGACGAAGAAGTGGCTGCGGCACCAGCAAAGAAGGGTGGTAGAAAGCCAGCAGCACCGAAAGCGG
TTCCAAAGGCGCCGGCAGCAAAGAAAAGAGGTGCAGCGGTGGCGAAGCAGAAGCTGATACCGGAGATGCTGAAGCCGGCAGAGGGAGAAGGGATATCGCC
GGAGAAGAAGGTGAGAAAGATGAGGGAGTCTCCTTTCAACAAGAAGAGTAGTTCCGTGCTGGGAGTATCTACTTCGTCGTCGAGTAATGAGATCTCGCCG
GGTTCGGAGATGACTGAAGAAAATATGGCTATGCCGGCGAGGGCCCGGCCGCAGAGGAACCGGAAGCCGACTATGTATGTGTTGAGCGATTCTGATGAGG
AGGACGACAAGGAGGATTCAGACGTCGGCGGCGAGTCAGACTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016397 pacid=23161874 polypeptide=Lus10016397 locus=Lus10016397.g ID=Lus10016397.BGIv1.0 annot-version=v1.0
MVASKKAPLMTSNSANVGADPSPDNFPQKTIEEMYQKKTQLEHILLRPDTYIGSIEKHTQMLWVYEDGKMVNRSVTYVPGLYKIFDEILVNAADNKQRDP
KMNSLKVSIDVENNLVSVYNNGDGVPVEVHKEEAVYVPELIFGHLLTSSNYDDNEKKTTGGRNGYGAKLTNIFSTEFVIETADGKRLKKYKQVFTNNMGT
KSDPEISNCKASESWTKVSFKPDLDKFNMTHLEEDVVALMKKRVLDMAGCLGKGVKVELNGIHIPVKSFQDYVNLYLDSTSENGVRRMSFYEKCGDRWEY
CVSITEGQFQQVSFVNSIATIKGGTHVDYVTNQITSYVTEVVNKKHKNAHLKAHNVKNYLWVFVNTLIDNPAFDSQTKETLTLRQSSFGSSCQVSEDILK
KVVKSEIVENLLNWAQFKQQKDLKKTDGSKTSRINVPKLEDANEAGGKNSAKCTLILTEGDSAKGLAVSGLSVVGRDYYGVFPLRGKLLNVREATVKQLN
ENKEIQYLKKILGLRQDKTYEDVKSLRYGHMMIMTDQDHDGSHIKGLLINFIHSFWPSLLRVPSFVVEFITPIVKATSKRGQVLSFYTMPEYKTWSDNLG
TAKSNWSIKYYKGLGTSTDKEGKEYFSDLDKHKKDFMWVDDNDGNAIELAFSKKKIEARKDWIRQHEPGTHLDQADKLIKYSDFINKELILFSVADLQRS
IPSMVDGLKPGQRKILFAAFKKNLVKELKLSQFIGYVSEQSAYHHGEQSLNSTIVGMAQNFVGSNNINLFVPAGNFGTRASGGKNAASARYIHTRLSLIT
RFLYPKDDDCILDYLEDDGQSIEPTWYMPIIPMALVNGSEGIGTGWSTFVPNYNPRDIVANIRRLLNEEMMEPMEPWYKGFRGTVVRGVSKESGVTYYCN
GVIDEVDETTIKVTELPIRTWTDNFREYLKSVTEGLKDENGNAPKDPLIKGYHEQGDDNEFEFTITLSEEKVQLARTEGLLKKFKLTSSISTTNMHLFDP
SGKIKKYDNPEHVIEEFFHLRLEYYDKRKKTKLAALELELLRLENKVRFILAVVNGEIIVNNRKRADLIQELHQKRFTPMPIKAKSSVTSDTTEESEENE
DSDPVNPSGVRASDYDYLLSMAISTLTFERVQKLLEDRDRFNGEVDNLRKMSVKDLWMHDLDAFEKQLDELEAQEELDRKLKPKSKQTRVVNAPKPRNRA
KPVQKPKSVEEDMESKLKLSSTSSAMETNEQPEPVPAAKPKAKAAATKKAPAKKGKQAALVDSDDDEVESLKDRLKAYKLDDSSPENSAEKKQAPKKRGA
KKAVIDLDDEESDEEVAAAPAKKGGRKPAAPKAVPKAPAAKKRGAAVAKQKLIPEMLKPAEGEGISPEKKVRKMRESPFNKKSSSVLGVSTSSSSNEISP
GSEMTEENMAMPARARPQRNRKPTMYVLSDSDEEDDKEDSDVGGESD
|
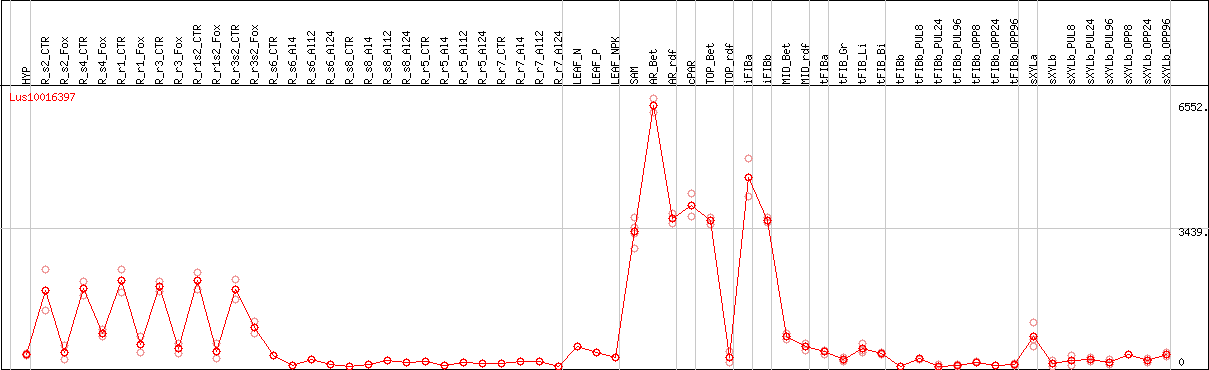
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016397 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
