External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G64350 189 / 2e-62
FKP12, ATFKBP12, FKBP12
ARABIDOPSIS THALIANA FK506-BINDING PROTEIN 12, FK506-binding protein 12 (.1)
AT3G55520 72 / 2e-15
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT5G48570 70 / 5e-14
ROF2, ATFKBP65
FKBP-type peptidyl-prolyl cis-trans isomerase family protein (.1)
AT3G25230 67 / 6e-13
ROF1, ATFKBP62
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
AT4G25340 67 / 7e-13
ATFKBP53
FK506 BINDING PROTEIN 53 (.1.2)
AT5G45680 64 / 1e-12
ATFKBP13
FK506 BINDING PROTEIN 13, FK506-binding protein 13 (.1)
AT5G05420 61 / 6e-12
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT5G48580 55 / 1e-09
FKBP15-2
FK506- and rapamycin-binding protein 15 kD-2 (.1)
AT3G25220 54 / 4e-09
FKBP15-1
FK506-binding protein 15 kD-1 (.1)
ATMG00310 53 / 5e-09
ATMG00310.1, ORF154
RNA-directed DNA polymerase (reverse transcriptase)-related family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003760
224 / 3e-76
AT5G64350 185 / 2e-62
ARABIDOPSIS THALIANA FK506-BINDING PROTEIN 12, FK506-binding protein 12 (.1)
Lus10003171
80 / 2e-17
AT3G25230 871 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10004714
75 / 1e-16
AT3G55520 265 / 3e-91
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Lus10040280
74 / 2e-16
AT3G55520 263 / 1e-90
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Lus10002338
77 / 3e-16
AT3G25230 874 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10038216
71 / 3e-14
AT3G25230 870 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10031700
71 / 4e-14
AT4G25340 324 / 3e-105
FK506 BINDING PROTEIN 53 (.1.2)
Lus10025888
71 / 4e-14
AT3G25230 870 / 0.0
FK506 BINDING PROTEIN 62, rotamase FKBP 1 (.1.2)
Lus10031121
69 / 1e-13
AT4G25340 335 / 3e-110
FK506 BINDING PROTEIN 53 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0487
FKBP
PF00254
FKBP_C
FKBP-type peptidyl-prolyl cis-trans isomerase
Representative CDS sequence
>Lus10016586 pacid=23143945 polypeptide=Lus10016586 locus=Lus10016586.g ID=Lus10016586.BGIv1.0 annot-version=v1.0
ATGGGAGTAGAGAAGGAGATCGTCAGCCCGGGAAACGGTCCCAAACCCTCTCGCGGACAGAAAGTCACCGTTCATTGCACCGGCTTTGGGAAAAACGGTG
ATCTATCTGAAAAGTTCTGGAGCACCAAGGACCCTGGGCAGCAACCTTTCTCTTTCAATATTGGCAAGGGAAGCGTCATTAAAGGATGGGATGAAGGTGT
CATGGGAATGCAAGTGGGAGAAGTCGCTCGTTTACGGTGCTCCCCAGACTACGCTTATGGAGCTAATGGATTTGGAGCATGGGGAATCCTGCCCAATTCA
ACTCTGGTTTTTGAAATTGAAGTCCTGAGCTTAGATTTATGGGGACGAGTGCTGAAAGCCAGATATTTCCCTAAATCCACCTTCATGGAAGAAAAGCTGG
GAGGAAGGCCGTACTGGATCTGGTCCAGCCTGATTAAAGCCAGGGACCTGCTAAGGATGGATGCCTTCATGGCGGTGGGGAATGGAACCTCATTTCGCAT
AGCATGCGACCCGTGGATCCCATCCTCTAATGTGATGACCCTGCATATCCCCTTTGGCGGTGGTCAATGGGCCTCAAGACAAATGGCCATGGGCCTGTGA
AA sequence
>Lus10016586 pacid=23143945 polypeptide=Lus10016586 locus=Lus10016586.g ID=Lus10016586.BGIv1.0 annot-version=v1.0
MGVEKEIVSPGNGPKPSRGQKVTVHCTGFGKNGDLSEKFWSTKDPGQQPFSFNIGKGSVIKGWDEGVMGMQVGEVARLRCSPDYAYGANGFGAWGILPNS
TLVFEIEVLSLDLWGRVLKARYFPKSTFMEEKLGGRPYWIWSSLIKARDLLRMDAFMAVGNGTSFRIACDPWIPSSNVMTLHIPFGGGQWASRQMAMGL
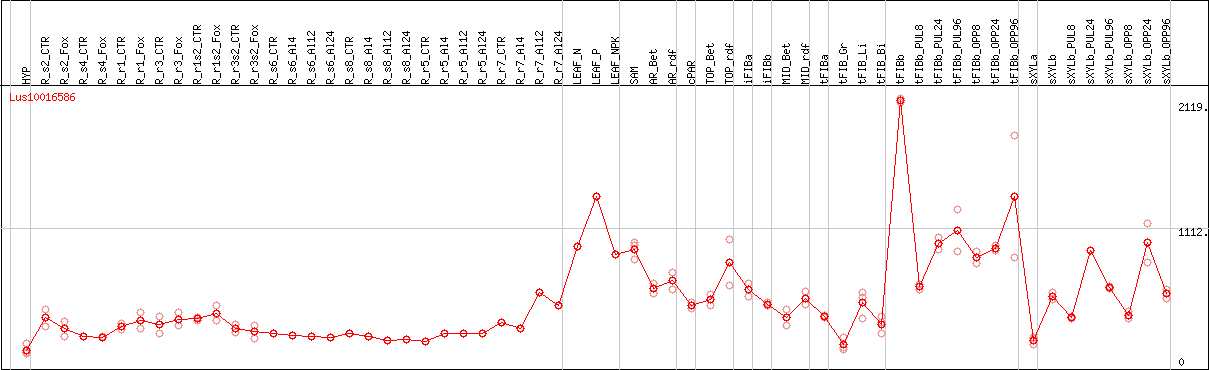
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016586 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.