Lus10016589 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016589 pacid=23143927 polypeptide=Lus10016589 locus=Lus10016589.g ID=Lus10016589.BGIv1.0 annot-version=v1.0
ATGAGATCCACCACCGAAAGCGGCGGCGACAGCCGCTTCCTTGGAACCATATCGGCCTCCTCAATCCGAAATCTGCTCCCTAGATCCATCTCTTCCAAGA
AGAAAACCAAGTCCAAACTCTTCCGCAGCGAAAACGACCACCCCAACGTTAGAATTAGTGACCCACCACTATCTCCTTCACTCCCAAAACCCTGGAAATC
ACCTAAATCCCTCGATTCCAACTTTCAGCGTGAAATTTATCCCGCTAGAGAACGTCTCTCTCTCAAGGAGGACTTGGTAACGGAATCAGTGGAGAAGAAG
GATTTCCCTGAAGCTTCAGATCCATCTGTGAAGGTTGTGGTGAGAATTAGGGTTCCAAATGACCAGACAAGAGAGTGGAATCGAACAGTAAGGAAGGTTT
CTTCAGATTCTATGTCGATGGGTGAAAGAAGTTTCACATTTGACTCAGTTTTGGACTCCAATTCAACTCAGGAAGACGTGTTTCAGGAAGTTGGAATTCC
GTTGGTCCAGAGTGCCTTGGCTGGATACAATGCTTCTATTTTGTCTTACGGGCAGACTGGAAGTGGGAAGACCTACACGATGTGGGGTCCTCCAAGCGCC
ATGGTTGAAGATCCTTCTCCCAGCAGTCACCAGGGGCTTGTTCCACGGGTTTTCCAAATGTTGTTTGACAAGATCCAGAAAGAGCAAGAGAACTCAGAAG
GAAAACATATCAATTATCAGTGCAGATGCTCTTTCCTTGAGATTTACAATGACCAAATAGGCGACTTGCTTGATCCAGTTCAAAGAAACCTTGAGATAAT
GGATGATCCCAAGAATGGATTGTACGTTGAGAATTTGACCGAGGAGTATGTGACAAGCTATGAGGATTTATCTCAAATATTAATCAAGGGACTTTCAAAC
AGGAAAGTTGGAGCAACAAGCATAAACTCTAAGAGCTCTAGGTCTCATATCGTGTTTACATTCATCATTGAGTCCTGGTGCAAGGGGACTTCATCAACAT
GCTTTAGCAGTTCAAAAATCAGCAGAATTAGCTTCGTTGATCTTGCTGGAATGGACAGAAATAAACCTGATGATGCTGGTCCGCAGCTTCTCAGGGAAAG
CAAACATGTGAAAAAGTCCCTCTCTCAGCTTGGGCATGTAGTGAGCACTCTTGCCAAAGAAGCTCAACCAGGAAAGTGCACAGATGTTCCTTACGAAGAG
TTCTGTTTGACACATTTGCTTCGCGAGTCACTTGGTGGAAATGCAAAACTTACAGTTCTTTGTACCATCTCTCCTGACAATAGGAATGATAGTGAGACAT
TGAGAACACTTAGATTTGGACAGCGAATCAAGTCAATTCAGAATAAGCCAGTCATAAATGAAATATCAGAGAATGATGTAAATGATTTGAGTGATCAGAT
ACGCCAACTGAAGGAGGAACTGATAAAGGCCAAATATGATTCATGTAAGTCAGCGAGAAGCAAAGGGGGTAGCTTTAGTGGGCGGAATGTGCGTGACAGC
TTGAATAGTTTGAGAGTTAGCCTAAACCGCTCCTTAATGTTACCCAAAATTGATATTGATTCTGACGACGAGGTTAACAATGTTGGCGTGGATGATGTAC
TGGAACTGCATAATCATCTTAACGAGTTCCGGACCTCAGGGGAGGGGGACTTAACAAGTTTATCCGAGAGCAAGGACTCAATGCATTTCACATCAGTAGA
CCAAAGTTTTGAGACTGCCGTGATGAGCGAAGACGAAGTCAATGGTCCGGCTGAACATGAAGAAATGGATTCGTGGAGAAACAAAGTAAATGTTGCATCT
GAAGACGAATTTCCAATTAACCTTAACGCCACCAGGGCAGAAGACGCTGCTGCCATCAGGAATAGCATTTCTATCAGCTTGTGCCGCCAGTCTCAAGTTC
TCCAAGAGCCAGTGCTGTCAGAATCTCCGAAAATTGGAAATACTAGGAAGAGCATTACAATATCACCCTCAGCATTTTCCTCTAGTCGGAGCAATGTTTC
TGAGAGTTCTAACTTCAACTCAATCGCTGCAAGTAAGACACTAAGGAAAAGTGAGCAAATAAGATCATCTCTGCGTTCAAGCAAGATATTAGCAGGTCCC
ACTGATTCTCTGGCAGCCAGCCTACAGAAGGGATTGCATATAATTGAACATCACCAACGAGTCTCCCCATCTAACAGATCCTCAGCGTCTTTCTCTTTTG
AACACTTTGCATTGACATCAACTGCTTCGGTTCAAAAGTTAGCAGAAGGTTCCCCATCTACAGAAGTGGCATCTGCCTCTTTACTGTGTGCATCCTGTCA
GGAGAAAATCATTGTGACAGGTGGGGAAGTCGGGAGTCATAATGAACTTGTCAATGGAACGACTAAGGGTTCAAGCAGTGATATAGTGGAATGTAGCACA
AGAGAGAAGGAACTTGAAACTTTATGCAAGGAACAGGAAGCCAAAATTGAGCAGCTAAATCAAATGGTTGAGCAATACAAATATGTGGCAGAACAACAGT
CCTCTGGACATGCACTAGTACTGGAGGCACCCAAACATGAGGAATCAGTGGATGACGAAGACAAGACAAATGACGAATGTGAAGTGAAGGAGGTGGTTTT
GGATAATGTGGACACAGGTTTTGACACAAGGGAGAAAGAAGAACTCCTATCCGAAATTGAAAGTCTGAAGGCCAAGTTGAGGGTGTATAGTGATGCACCA
TCGTCAACGAGGTCTACTGAGGAGCTCAGGTCCTCCTTACTGGCAAGATCCGTTCTACTGAAGAGCATAGACATTCGGAACAACTCTGAAGAACTGGAGA
AAGAAAGGGAGAGGTGGACAGAAATAGAGAGTGGCTGGATATCTCTAACCGACGAGCTGAGGATTGAAATTGAGTCCAGCCGGCGGCATGCAGAGAAAGT
GGAAATGGAACTAAGACTGGAGAAGGCAGTTACGGAGGAGCTGGACGATGCTCTCCACAGGGCTATGGTTGGACATGCCAGGATGGTGGAACACTATGCT
GATCTACAGGAAAAGCACAATGACTTGGTTGAAAAGCACAAAGCAATCTTAGAAGGAATAGCAGAGGTGAAGAGGGCAGCGGCAAAAGCAGGAAAGAAAG
GCGGTAATCGATTTGCGAAGTCGCTTGCTGCAGAACTTTCAAACGTGAGGGTGGAAAGAGAGAAGGAGAGGCAGTTGTTGAGGAAGGAGAACAAGAGCCT
GAAACTGCAGCTAAGGGATACTGCTGAAGCTGTACATGCGGCAGGAGAGCTCCTTGTGAGGCTCAAGGAAGCAGAGCAAGCTGCTTCACTTGCAGAGGAG
AAATTGAGCAATGTAGATGAAGAGAATGAGAAGTTGAGGAAGCAGATGGAGAAGCAGAAGCGGAAACACAAGATGGAAATGGTGACAATGAAACAGTATC
TGGCAGACAGCAGGTTGCCTCAATCAGCACTGCGGAAGCCGGCAATGTACGATGAGGCTGAGGATTCAGAATTGCCACAGAGTACAATCCCATATGATGA
TCAAGCTTGGAGAGCTGAATTTGGAGCCATATATAATAATCATCACTACTGCAACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016589 pacid=23143927 polypeptide=Lus10016589 locus=Lus10016589.g ID=Lus10016589.BGIv1.0 annot-version=v1.0
MRSTTESGGDSRFLGTISASSIRNLLPRSISSKKKTKSKLFRSENDHPNVRISDPPLSPSLPKPWKSPKSLDSNFQREIYPARERLSLKEDLVTESVEKK
DFPEASDPSVKVVVRIRVPNDQTREWNRTVRKVSSDSMSMGERSFTFDSVLDSNSTQEDVFQEVGIPLVQSALAGYNASILSYGQTGSGKTYTMWGPPSA
MVEDPSPSSHQGLVPRVFQMLFDKIQKEQENSEGKHINYQCRCSFLEIYNDQIGDLLDPVQRNLEIMDDPKNGLYVENLTEEYVTSYEDLSQILIKGLSN
RKVGATSINSKSSRSHIVFTFIIESWCKGTSSTCFSSSKISRISFVDLAGMDRNKPDDAGPQLLRESKHVKKSLSQLGHVVSTLAKEAQPGKCTDVPYEE
FCLTHLLRESLGGNAKLTVLCTISPDNRNDSETLRTLRFGQRIKSIQNKPVINEISENDVNDLSDQIRQLKEELIKAKYDSCKSARSKGGSFSGRNVRDS
LNSLRVSLNRSLMLPKIDIDSDDEVNNVGVDDVLELHNHLNEFRTSGEGDLTSLSESKDSMHFTSVDQSFETAVMSEDEVNGPAEHEEMDSWRNKVNVAS
EDEFPINLNATRAEDAAAIRNSISISLCRQSQVLQEPVLSESPKIGNTRKSITISPSAFSSSRSNVSESSNFNSIAASKTLRKSEQIRSSLRSSKILAGP
TDSLAASLQKGLHIIEHHQRVSPSNRSSASFSFEHFALTSTASVQKLAEGSPSTEVASASLLCASCQEKIIVTGGEVGSHNELVNGTTKGSSSDIVECST
REKELETLCKEQEAKIEQLNQMVEQYKYVAEQQSSGHALVLEAPKHEESVDDEDKTNDECEVKEVVLDNVDTGFDTREKEELLSEIESLKAKLRVYSDAP
SSTRSTEELRSSLLARSVLLKSIDIRNNSEELEKERERWTEIESGWISLTDELRIEIESSRRHAEKVEMELRLEKAVTEELDDALHRAMVGHARMVEHYA
DLQEKHNDLVEKHKAILEGIAEVKRAAAKAGKKGGNRFAKSLAAELSNVRVEREKERQLLRKENKSLKLQLRDTAEAVHAAGELLVRLKEAEQAASLAEE
KLSNVDEENEKLRKQMEKQKRKHKMEMVTMKQYLADSRLPQSALRKPAMYDEAEDSELPQSTIPYDDQAWRAEFGAIYNNHHYCN
|
DESeq2's median of ratios [FLAX]
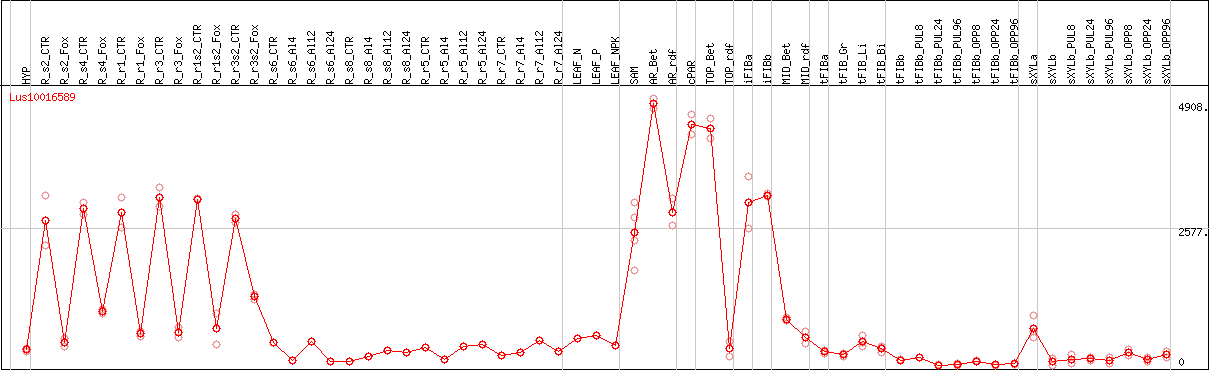
Coexpressed genes
Lus10016589 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
