Lus10016620 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016620 pacid=23143976 polypeptide=Lus10016620 locus=Lus10016620.g ID=Lus10016620.BGIv1.0 annot-version=v1.0
ATGGCAAGTGTAGAGAAGCTGCTCGTACAGATCTTCGAGAAGAATAAGTTGATAATTGAGCAGGTCAAGAAGCAATCTGACCTCTTCGATCAGCATCTCG
CTTCAAAATGTATCCTCGACGGAATCTCCCCTCCTCCATGGCTTTTCTGCACCTCTTTCTCGTCTGTTTTATCGGATTCAAACGTCCTGAAGAAGGAGGA
GCTCATATCAGAGCTTTTGCTTACGCGTTCTCAACCAGGAAGCATTTACCCTAATAGTCATGACTCCCTTCATGAAAAGCCTTCTGATCATGTTTTGAAC
GAGGTTCCACCAAATCACTTGGAAGGAGATGGTCATGATGTCAACAACGCCCCTAATACAGTTGACAGGTCTGCTCTGTCTCAATTGCCTACCACTGACA
GTCCCTGTGCTCTTAATCAAGTCCTTGAGCAGGATTCTAGTGTCACATCTCCGCAGGAGTGTAGAGATACCCAAGGTCCTGATAATTTTGCTGATCAAAG
TCAATCATTGGTGAGGATTGAAAGATCAAAATCGAGGCAAAGAGCATTGGAGACTCGTAGTTATTCAATACTTGCTAAACGACGAGCAAATGATGAGCCT
GAACCTGGTGTCTGTACTAGTCAATATGAAGCAATTTCTACAAAGAGCAAACTGAAAGTGACTGAGTCTTTGAAGAAAGGGCACAACAACGATAATTGTA
ATTTTGGAACTGGGACGAGGACACCGTGCCTGTCATCCAGTCCTATGAATGAGTTATTCAGTGTTAGCAAGTGTTCAGATATTGCCAAGGTAGATGGTCT
TACAGATACTAATGGCAAGTTGATGGACCAGTCTTATTATGCTAATCCGATGTCAGCAACAGCCAATCCTACTGCGACTGTGGATGCAGTTTGTGCCGCA
GAAGAAGCATATGGTGGTGATTGCAGCATAAATGAAGTTGGTAGTGCTATTCATTCTGATAGAGTTACAAACTCTAGACATTCAAGCATTCGAACTGGAC
TTGTAAAGGATACATCAGAGGTGGTAACATGCATTCATAATGTAGCTGAAGACTGTAGATCTGAGCAGTCCACTAAAAAATCCTCAAAAGATGCTTATAA
TGAGAACTCGTTACATGGATGGATCAAACCTTCCTCTGTAATTTCCAATCACAGTCATGCACATCCAAATTCGCAGGCGACGGATGTCAAGAGCATAGAA
ATGGGGAGCGGTGGTGATCTTGTGAGAGTAACAAGCCACGATAATGAAGGCATGCAGCTAGATTTTCCTCTCAGTATTGGGAATCAAGATGATTGTGAGC
AAAACCAGGTTCCCCCCCATGCCAATATTGTAATGGACTTGATAAAACCTGCAACTGAAGAACCTGAATCTCAAATCACTAAGATGGGCAATGACAGCAT
GGGATTACATAAACCAACACAGTCCATAAGCTCCAATCTACAAAATGGCAATGTTTATGGTGGTTCGATAGAAATAGATGTTTCATCTAACAATGCTGAA
GATGCAGATATTGCAGTTGCAGATTCTAATGGATTATCGCAAATACTTCAATCGTCAAGTTTGGTTGAGCAAGAAGTCTGCTTGCTTCCAGCTTTGGGTA
TTCATCCTAGTAGCTTGCCTTGTACTGAAAGCACCAGTCTTGCTGACATGGAAACATGTGGTCCAGTTCCAGCTGAAACAGATTTAAGGTCTCCTGGACC
AGTTGCACCTGATTCTGTAACTTCTAGGTCTAATTCAGCTGCCAATACTGGATTGGCCTCAAGGTCAATAAGTGAAGATGCCATGGTGATCGAGCCCAAT
CAACTTCATTTTGATGACGTGGAGGCTAACCGGGTTTATGATATTGACGGCCAAGACAAGGATGGTTCATTATCACTGAAACAATCTCCGACAGTACGAT
CCAGTACCTGTTTATTGACTGAAGTAGCTTCTGTGAGCGACCAAGAAAATCACAAATTATTGCCTGATGCACTTGAGGGCATTGATAGCCAGAAGCACTC
TAGGAAGGATTCGTTGCGAGATCTTGTCAGCGACAGTATCAAGTCTGTTGATCCAAGTAATATCAACTTTGTACAGGATCAGAATTTTTTTGATGCTTCT
CAAAATCCATATGTTTATAGTTCTGTGATGGATTCATGGCCTCAATACAAACGCATAAAGATTGCTGATAGGCCAAGTTATGCATGTCCCGCTCCTTCAA
GCTTATCTACATTGCCATTTCAGCCCTTCCTAAAACAAAATCTGCAGGAGGCTATGGGCTCAAATGACTATACACCTGTTGGTGAAAAGAAGCGTGAATA
TGCGGATAACCATAAAGTACAGCCATGTGAAGTTGTACCATCCACGCTGCCAGTTAGAGAGTTAGGATGGGCTGATGACCCTAGGTCAGCTCTACTTAAG
TTTACATGGTTCCGTTTGATTACAGCCCTTAACCTAGCTGGGGTAGATTCCAACGATGAGAATACCTTGGGAGGGAGGGACAGAAGTGAAAGTTCCATTG
TCACTTTTACGCATGAACAGCTAGAAGCACCTTCAGTCTCAAGTCTGACAAAGCTGGCTACTGGCTCATCTGAAATATCTTTGTCTGGGCATACCAAATT
TTCCGAGATGCGAACTTCTGATGAAAGCCAAGTTTCATCACAGGTTTCCGATAGTTTTTATTTGGGAAGCACCTTTTTAACTCGCTCCGACAACATTATC
CAGATTGAAAATTCTGATCTTGGAGAGAATAGCAGACCGTCCGATTACTCTGTCGTTTCTCCATTTTCTGTGGATCTGATTGCACCTAATCAAAATTCAG
TTGAGTTGGAGAGATTTTTTGTGGAAACGGATGACTCAGTACCTTGTGCTACCAGAAACGGGACAAGCTTTGGCACACTGGACTGTCCGAGCATTGCCCT
TGATCATGCTAGTGTTTTGGAGCAGTTTCGCAAATCTGCTTGTTTGCAAACTCCACTGTCCAGTTTTCTGGATACTTGTAAATTACAAAACACTTTAGGT
CTAGAACAAACCATCCCGGATGTCTTTTTGGGGAGCTTGGACTTGAAGGGGCACCTTGTTGAGGACGGTGACACCACTAAGCAGTTTGAAGTCAGTGGCC
ATTGCTTGAATGAAGAACTGCACTGTGCATATCGTGAGATTCCTCATCCTTACTGCTTACCATCTCCTAATGCTCATTTTCATCAGGAAAGTAGCAAACC
TTACATGTCCCCAGTAGGGAGATTCTGGGATGGAATCCCACCAAAATCTCGCAGTTCAGAGAAACGAGCAAGTTCAATTCCAGATCTTCCCAGCATTCAA
GAGGAGACTGAGAATATGGATGCCATAACTGATTCAATGCATGATGACATCATTCCAGAAAAGGCAATTAAGGATGCTGAGAGAAAACCACTTACCGAGC
TGACTGAACATGCAAATTTTCCAATATCTGTATGTGAAGATGAGAGGTTAGAAGATAGAGATAGTCTTGCATCTGTGAGCACAGAATTCAGTTTGCCTAG
CACTTGCAAGATGACAAATTTGAAACCTGGGAATCGGATGAGCACCAGATCGAGATTTGCATATGCAAGTAATGGGAAGGTGAATCCGAGGGTCCGACTT
GGATCAGATGGCGTTAGAAGGAATGATGGATCATTACAGAATAGAATCAGTAAACCAAAGTTGTCTAGGAAAAACGAGTCAAGAAAACATAGCTCTGGCG
TGTTGGGGAAGGAGCCTAAATGTAACAATATCGTGTCCAATATTACTTCCTTTGTTCCTCTAGTGCAACAAAAGCAAACGGCTGCTGCTATTACAGGGAA
GAGGGATATCAAAGTGAAGGCCCTGGAGGCTGCCGAGGCTGCAAAGCAACTTGCTGAAAAGAAAGAAAATGATCGAAGGTTGAAAAAGGAAGCCTTGAAA
CTTGAGCGGGCAAGGATCGAAGAGCAGAACTTAAGGCAGCTTGAGTTAGAGAAAGAAAGGAAAGAAAAAGAACGGAAGAAAAAGGAGGCTGAGATGGCGG
CCAAAAAAAGACAGAGAGAAGAAGAACAGAAGAGGCAGAGAGAAAGGAAAAAGAAGTGTGTTGAAGAAACAAGAAGGCAGAACATGGAACACAGGGAAAA
GAGGAACGGGAACAAAGAAGAAATGAAGCCTGGAGCAACGGATGTGAATGCATGTAAAACAAAGGAATCTAAGGAAAACTCAGCAGAACATCTAAGGATA
GGGAAGGCAAAGCCAGGTAGTCAAGTACCTAGAGAGACCGAGACGGAGCATACAGACACTCATCTCCCTGCAATTGGCACAAAGAGTCCAAGTATTGCAA
CTGAAGCTTGTGATGCGATGAGCAATTCTGGTGATAAATCAAAGGTGATAAATGAAATATGCGAGAGAAAAGAGAAAGACAGCTTGGTTTCCAACATGAT
ACCTGAAGAATCATACGCAATATCTCCATACAAAGGCTCAGATGATGAAGATGAGTACGATGATGATAATAAGCAGAATACGAAGTTTGTTCCTTCCTGG
GCCAGAAAGCGTAGCATATCATTGGCTATCTTGTCCCAGCAGACGATAGATCCGGAATCAATCTTCCCACCTGAAAGCTTTTGCAGTAGAGACAAAGTAC
AAGTCCTCATGGAATTCAGCGCCACTAGCAAGGAATTTGGCATTCCCCGTGCCAAGTCGGATTATTGTATATTCAATGCTTCCATCCGCCTCATTCCTTC
TCCTCTTCCTCATCATCACATACGGTGCCGTCGGGTCACATCCTCCGCCGCCGCTGGAGCCGCCAACTACACCTTCGACTTTGACTTCCCATCCCTCGCC
GTCCGCAACCTCACCCTCCTCGGCGACTCCTACTTCCGCGACGGCGTGGTCGGCCTCACCCGCGAAGTCACCGTCCCTACATCAAGCTCCGGCTCCGTCA
TCTACAACAACCCCATTCCCTTCTCCGACTCGGATTCTCCTCCCTCCGCCGCTTCTTTCTCCTCAAGATTCACCTTCTCCGTCGCCAACGTCAACCCCAG
CTCTTTCGGCGACGGCCTCGCTTTCTTCCTCTCCCGCGACAACCAGACTATGGGCAGACCCGGCGGTTACCTTGGCCTCGTCAACGCCTCCCTGTTGACC
AAGAACAAGTTCGTCGCCGTGGAATTCGACACTCGGGTGGACCCCCACTTCGGCGACCCCAACGAGAACCACGTCGGCTTAGACATCGACAGCCTGATTT
CCGTCAAGACGGCGGATCCATTAACCCAGGGGATCGATTTGAAGAGTGGGGATCCAATCACCGCCTGGGTTGACTACGACGTCGGCGACCTTAACAATCC
CACCGCTGCAGCAGCAATCTTGCAAGTGTTCTTGAGCTACTCCAGTACTTCCAAGCCCCAAGATCCTCTGTTATCCGTCGACGTAGACCTCTCAGACTAC
CTCAAAGGAGACATGTACGTCGGATTCTCGGGCTCCACTGAAGGATCCACGGAGCTCCACTCGATTCATAATTGGAGCTTCAACATGACGAGACAAGGAT
TATCGCCATCGTCACTAAGGCCAAGATCCACGATTCCTCACAACGTCTCCGAAAGCTCGATCATCATAACCTCTCCGGATATTCCGTCTTCTTCCAACCC
TAAAAACAAGAACCACAAGAAGCTCGGATTAGGGCTCGGAATCGCCGGGCCATCTTTCTTCTGCGCGTTTCTGGTGGTATTCGGGTACATTTCATTCAAG
AAATTGCAGAGGATGAGAAGTAGTACCGGGAAAACAGAGCTTGTGACGGGGCGTCCGAAGGAATTCAATTACAGAGAACTGAGCTCTGCGACGAGAGGGT
TCCATGCCAGCAGAATAATCGGGAAGGGAGCGTTTGGGAACGTCTACAAGGCGTTCTTCGTCTCGACGGGAACACTCGCCGCCGTGAAGAGATCGAAACA
CTCACACGAAGGGAAAACAGAGTTCCTAGCTGAGTTGTCAATCATCGCCGGCCTCCGCCATAAGAACCTCGTCCAGCTGCTCGGCTGGTGCGTCGAGAAA
GGAGAGCTACTGTTGGTGTACGAGTTCATGCCTCACAGCAGCCTGGACAAGGTCCTCCACCACCAGGACTCCTCCTCCTCCGGCGGGCTGTCGTGGGCGC
AGCGGCGGAACGTAGCGGTGGGGCTAGCGTCGGCGCTGACGTATCTACACGAAGAGTGCGAGAAGCAGGTGATACACAGGGACATAAAATCGAGCAACGT
GATGCTGGACGTGCATTACAAGGCGAGGTTAGGGGATTTCGGATTGGCTAGGCTGATGGAGCATGATAAGAGCCCAGTGTCAACTCTGACAGCAGGGACG
ATGGGGTATTTGGCTCCGGAGTATCTGCACTACGGGAAGGCGACGGAGAAGACTGATGTGTTTAGCTACGGGGTAGTGTTACTGGAAGTGGGTTGCGGGA
GGAGGCCGATAGAGAGGGAGTCGGAGAGCCAGAAGATGGTGAATCTGGTGGATTGGGTTTGGGGTTTGTACGGTGAAGGGAAATTGTTGGAAGCTGCTGA
TGAGAGGTTGAATGGCGAGTTCGATGTTGAGGAGATGAGGAAGATGCTTATGATTGGACTGAGCTGTGCGAATCCTGATAGTGGGGAGAGGCCGAATATG
AGGAAGGTGCTTCAGATGCTGAATGGGGAAGTTGAGGTTAAGGCTCTTGCTAAGACTAAGCCTACTCTTACTTTCTCTTCTGGGTTCACTCTCTCGCTTC
AGGATATTACCTCTGACTCTGAATACTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016620 pacid=23143976 polypeptide=Lus10016620 locus=Lus10016620.g ID=Lus10016620.BGIv1.0 annot-version=v1.0
MASVEKLLVQIFEKNKLIIEQVKKQSDLFDQHLASKCILDGISPPPWLFCTSFSSVLSDSNVLKKEELISELLLTRSQPGSIYPNSHDSLHEKPSDHVLN
EVPPNHLEGDGHDVNNAPNTVDRSALSQLPTTDSPCALNQVLEQDSSVTSPQECRDTQGPDNFADQSQSLVRIERSKSRQRALETRSYSILAKRRANDEP
EPGVCTSQYEAISTKSKLKVTESLKKGHNNDNCNFGTGTRTPCLSSSPMNELFSVSKCSDIAKVDGLTDTNGKLMDQSYYANPMSATANPTATVDAVCAA
EEAYGGDCSINEVGSAIHSDRVTNSRHSSIRTGLVKDTSEVVTCIHNVAEDCRSEQSTKKSSKDAYNENSLHGWIKPSSVISNHSHAHPNSQATDVKSIE
MGSGGDLVRVTSHDNEGMQLDFPLSIGNQDDCEQNQVPPHANIVMDLIKPATEEPESQITKMGNDSMGLHKPTQSISSNLQNGNVYGGSIEIDVSSNNAE
DADIAVADSNGLSQILQSSSLVEQEVCLLPALGIHPSSLPCTESTSLADMETCGPVPAETDLRSPGPVAPDSVTSRSNSAANTGLASRSISEDAMVIEPN
QLHFDDVEANRVYDIDGQDKDGSLSLKQSPTVRSSTCLLTEVASVSDQENHKLLPDALEGIDSQKHSRKDSLRDLVSDSIKSVDPSNINFVQDQNFFDAS
QNPYVYSSVMDSWPQYKRIKIADRPSYACPAPSSLSTLPFQPFLKQNLQEAMGSNDYTPVGEKKREYADNHKVQPCEVVPSTLPVRELGWADDPRSALLK
FTWFRLITALNLAGVDSNDENTLGGRDRSESSIVTFTHEQLEAPSVSSLTKLATGSSEISLSGHTKFSEMRTSDESQVSSQVSDSFYLGSTFLTRSDNII
QIENSDLGENSRPSDYSVVSPFSVDLIAPNQNSVELERFFVETDDSVPCATRNGTSFGTLDCPSIALDHASVLEQFRKSACLQTPLSSFLDTCKLQNTLG
LEQTIPDVFLGSLDLKGHLVEDGDTTKQFEVSGHCLNEELHCAYREIPHPYCLPSPNAHFHQESSKPYMSPVGRFWDGIPPKSRSSEKRASSIPDLPSIQ
EETENMDAITDSMHDDIIPEKAIKDAERKPLTELTEHANFPISVCEDERLEDRDSLASVSTEFSLPSTCKMTNLKPGNRMSTRSRFAYASNGKVNPRVRL
GSDGVRRNDGSLQNRISKPKLSRKNESRKHSSGVLGKEPKCNNIVSNITSFVPLVQQKQTAAAITGKRDIKVKALEAAEAAKQLAEKKENDRRLKKEALK
LERARIEEQNLRQLELEKERKEKERKKKEAEMAAKKRQREEEQKRQRERKKKCVEETRRQNMEHREKRNGNKEEMKPGATDVNACKTKESKENSAEHLRI
GKAKPGSQVPRETETEHTDTHLPAIGTKSPSIATEACDAMSNSGDKSKVINEICERKEKDSLVSNMIPEESYAISPYKGSDDEDEYDDDNKQNTKFVPSW
ARKRSISLAILSQQTIDPESIFPPESFCSRDKVQVLMEFSATSKEFGIPRAKSDYCIFNASIRLIPSPLPHHHIRCRRVTSSAAAGAANYTFDFDFPSLA
VRNLTLLGDSYFRDGVVGLTREVTVPTSSSGSVIYNNPIPFSDSDSPPSAASFSSRFTFSVANVNPSSFGDGLAFFLSRDNQTMGRPGGYLGLVNASLLT
KNKFVAVEFDTRVDPHFGDPNENHVGLDIDSLISVKTADPLTQGIDLKSGDPITAWVDYDVGDLNNPTAAAAILQVFLSYSSTSKPQDPLLSVDVDLSDY
LKGDMYVGFSGSTEGSTELHSIHNWSFNMTRQGLSPSSLRPRSTIPHNVSESSIIITSPDIPSSSNPKNKNHKKLGLGLGIAGPSFFCAFLVVFGYISFK
KLQRMRSSTGKTELVTGRPKEFNYRELSSATRGFHASRIIGKGAFGNVYKAFFVSTGTLAAVKRSKHSHEGKTEFLAELSIIAGLRHKNLVQLLGWCVEK
GELLLVYEFMPHSSLDKVLHHQDSSSSGGLSWAQRRNVAVGLASALTYLHEECEKQVIHRDIKSSNVMLDVHYKARLGDFGLARLMEHDKSPVSTLTAGT
MGYLAPEYLHYGKATEKTDVFSYGVVLLEVGCGRRPIERESESQKMVNLVDWVWGLYGEGKLLEAADERLNGEFDVEEMRKMLMIGLSCANPDSGERPNM
RKVLQMLNGEVEVKALAKTKPTLTFSSGFTLSLQDITSDSEY
|
DESeq2's median of ratios [FLAX]
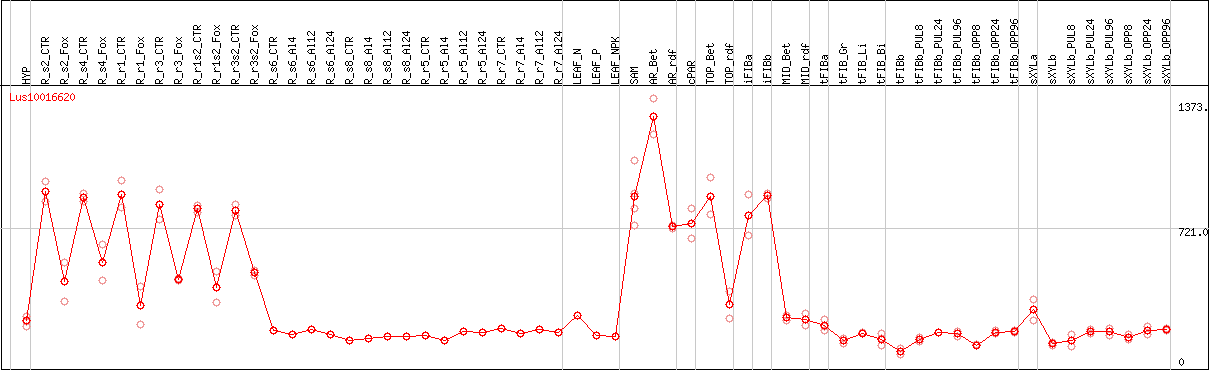
Coexpressed genes
Lus10016620 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
