Lus10016664 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016664 pacid=23143934 polypeptide=Lus10016664 locus=Lus10016664.g ID=Lus10016664.BGIv1.0 annot-version=v1.0
ATGGGGAAAAAGTTAAAGAAAAAGGGCCGTGCGCCTCAGACTCAGAAGGAGAAGAGGTTAGTTAGCCACTCTCCTAGCAAGGTACCTCATCAACCAAATA
TCCGTTTGAGTGAGGACAAAGTTAGTGCTGATGTTGTCATGGTACCTGAACCGCGAAGAATATGTGCTCATCTGGAGAAGGGGTTTGATTTAACCCAGTT
TACTGAGAAAATTAATTCTTCAGATTCTGCGGAGTGTGAGGATTGTCAGGAGGCTGTTGCTGACAGAAGGGGAGGAGCCAAAGGAAAGGGTAAACATGGG
AAGAAGAAAGGAGCTGATTCCAAGTCTGATTCTAAGGCCATCTGGATATGCTTGAAATGTGGCCATTATGCTTGTGGAGGGGTTGGTTTGCCAACTAACT
CTCAGTGCCATGCTATGCGGCATGCTAAACAAAATGGTCACCCCTTGGCGATCCAGTTGGTGAACACGCATCTTAGGTGGTGCTTCAAGTGCAACAGCTT
CATTCCAGCTGATAAATCAGAAGATAATGATGAAAATAGGGATGCAATGTCTGATGTTGTAAGAGTTATCGAGGCTCATGCATCTCATACGCCATCAGTG
CATGTTAAAGATGCTTGGTTTGGAAGTGGCAGTGTCTTGAGTGAAAAAAAACCACCAGCCACTGTCTCCAGTAGTTTACAAGGCAATGGCGGATACATGG
TGAGGGGCTTGGTTAACCTGGGGAACACTTGCTTTTTCAATTCCGTAATGCAGAATCTACTAGCTCTGAATAGTTTGAGGAGTTACTTTTTCAGTGAAGA
TTCTTCTTATGGTCCTCTTTCTATTTCTTTGAGGAAGCTTTATGATGAAACAAGACAAGAGACAGGTATGAGAAATGTGATTAATCCGCGATCGTTTTTT
GCGGGTCTCTGTTCTAAGGCTCCTCAGTTCAGGGGATATCAACAGCATGATAGTCATGAATTATTGCGTTGCTTGCTTGATGCATGGTCCACCGAAGAGT
TGGCCAGAAGAAAACAGATCAGCGCATTTGAGCAAAGTGCTGTTTCTGTACCGAATCTTCCTACATTTGTGGATGCTGCGTTTGGAGGACGAATCTCTAG
TACTGTATGTTGCATAGAATGTGCACATTCCTCAACTGTCTATGAGCCATTCTTGGACCTTTCTCTTCCAGTTCCAACAAAGAAGCCTCCAACTAAAAAG
CTCCAGTCTTTATCTCGAGGAAAGAAGACCAAGCTACCACCAAAGCGAGCTGAAAAAGTTAGGCAAAAAACCAGCAAGGATACAGATCGAATGCATGCTG
AGAGTGCCACAAGCTCCTCAACGAGCAATGAATTGGGCCACGAAACGCATTTAGCGACCCCTCTTAATGATAATGCAGCATCTTCCTTGGGTCCTACAGT
GCCTGACTCTGCTGGCCGAACTCCCGAGCCTGAAGAAAAGGAGTCGGTACCTAGAAACTTATTGGACACTCATTGTGTGCAAAAGGAAGACCTTGGGGGT
GCGCTGGAGCAAACATCATCTTTAGATGAATTTGCATGGATGGATTATCTAGAACCTGAAACTGAAACTATTTCAAACGAGAACGATCTGGCTCTACAAG
TTGATGACTTATCAATTCAATATTCACGGGAGCAAGATTTGGGCTCACCGGAGAAAGAGGAGACTCTTCCTTTTGGTCCAGTTCAACAAATCCCATATCC
AGAGCCAGACTCATCTTCGATAAATCCTTGGGAAGAGGAAGTCCCGTTGCAGGTCCAAAGCTCTGAAGTTCTGCTGCTTCCTTACTATGAAGAGAGCGTC
ACTTGTGGAGAAATTACTGCTGGAGAAGCTGAGGCCTCATCTTCAGTTGTGGGATGTGGACAGGATGAAGCAGCCTTTGACGGCTTTGGTGATATGTTCA
ACGAGCCTGAGGTTTTTATAGGCCCTGTTGCAGGGCCTGCTTTAAGCAATGGTCTTACGGCTGGCAATAGCAGCGACTCTGACCCTGGTGAAGTTGATGA
CTCTGATTCTCCAGTATCCGTGGAAAGTTGTCTGGCCCACTTTATAAAGCCAGAAATTCTCTCCAATGATAATGCTTGGGAATGTGAGAACTGTTCCAAA
ACCGTACACCTCCAGGAGCGGAAGGCAAAGAAGAAGCTGGGAAGAATTGTAGTAGGAGTTTTAAATGGAGGCGAGAGTGAAAGCCAAAATTTAGGCACCT
CGTGCTCGTCAGAAAATGCCACAAATGGGGAAGCCATCACAGATGCTCTTTACCATTCCAAAGATGAAAACATAGTTTCTAATGGTGGAGAAAATGATTG
CACAAGTGAAATTGTCATGATATCTGAAAACGGTCCAGTGCCACAATGTTGTCCTATTCAGGAAGAATCAGAAAGTCAGGATGGCTCACATACCCGTGGA
GTTGATGATTCTGTTTCTACCTGCACTGGAAATACTGCTGCTGTGAATCAAGAGACTCAGTCACAGTCATCTACAAATGTTGAATCAGAAGAAAGTGAAG
ACGAAGAGAAAACAATCAAAAAAGTGATTGTGAAGAGGGACGCAACTAAAAGGGTCCTTATTGACAAAGCACCGCCAATACTGACAATACATTTGAAGAG
GTTCAGCCAGGATTTTCGTGGACGATTGAGTAAATTGAATGGCCACGTTAACTTTAGAGATCAACTTGATCTTAGACCATACATGGATCCGAGGTGTGAA
GATAAAGAGAGGAGTGTGTACCGATTGACAGGGGTCGTTGAACATCTGGGGACAATGCGAGGAGGCCATTATGTTGCATACATAAGAGGAGGAGGAGGAG
GAGGAGGGAAGAGCGGTGGTGGTTGCGTATGGTATCACGTGAGCGATTCGTATGTGAGGCAGACTTCGTTGGACGAAGTTATGCGCTGTGAAGCGTACAT
CTTATTCTACGAGAGGGTTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016664 pacid=23143934 polypeptide=Lus10016664 locus=Lus10016664.g ID=Lus10016664.BGIv1.0 annot-version=v1.0
MGKKLKKKGRAPQTQKEKRLVSHSPSKVPHQPNIRLSEDKVSADVVMVPEPRRICAHLEKGFDLTQFTEKINSSDSAECEDCQEAVADRRGGAKGKGKHG
KKKGADSKSDSKAIWICLKCGHYACGGVGLPTNSQCHAMRHAKQNGHPLAIQLVNTHLRWCFKCNSFIPADKSEDNDENRDAMSDVVRVIEAHASHTPSV
HVKDAWFGSGSVLSEKKPPATVSSSLQGNGGYMVRGLVNLGNTCFFNSVMQNLLALNSLRSYFFSEDSSYGPLSISLRKLYDETRQETGMRNVINPRSFF
AGLCSKAPQFRGYQQHDSHELLRCLLDAWSTEELARRKQISAFEQSAVSVPNLPTFVDAAFGGRISSTVCCIECAHSSTVYEPFLDLSLPVPTKKPPTKK
LQSLSRGKKTKLPPKRAEKVRQKTSKDTDRMHAESATSSSTSNELGHETHLATPLNDNAASSLGPTVPDSAGRTPEPEEKESVPRNLLDTHCVQKEDLGG
ALEQTSSLDEFAWMDYLEPETETISNENDLALQVDDLSIQYSREQDLGSPEKEETLPFGPVQQIPYPEPDSSSINPWEEEVPLQVQSSEVLLLPYYEESV
TCGEITAGEAEASSSVVGCGQDEAAFDGFGDMFNEPEVFIGPVAGPALSNGLTAGNSSDSDPGEVDDSDSPVSVESCLAHFIKPEILSNDNAWECENCSK
TVHLQERKAKKKLGRIVVGVLNGGESESQNLGTSCSSENATNGEAITDALYHSKDENIVSNGGENDCTSEIVMISENGPVPQCCPIQEESESQDGSHTRG
VDDSVSTCTGNTAAVNQETQSQSSTNVESEESEDEEKTIKKVIVKRDATKRVLIDKAPPILTIHLKRFSQDFRGRLSKLNGHVNFRDQLDLRPYMDPRCE
DKERSVYRLTGVVEHLGTMRGGHYVAYIRGGGGGGGKSGGGCVWYHVSDSYVRQTSLDEVMRCEAYILFYERV
|
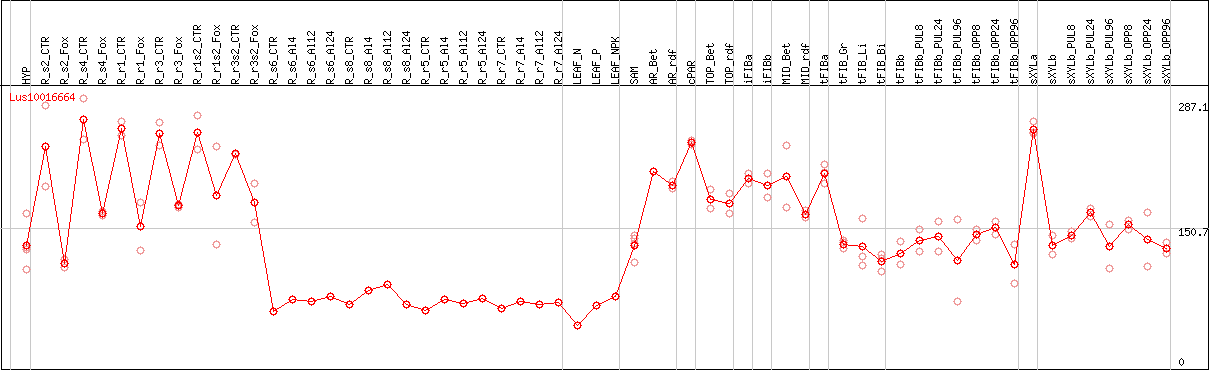
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016664 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
