Lus10016684 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016684 pacid=23149047 polypeptide=Lus10016684 locus=Lus10016684.g ID=Lus10016684.BGIv1.0 annot-version=v1.0
ATGGATGCGACGCAGGTACTACCTGTGGATGAAGTGGTGGCGGAAATCATGAAAATTCACAGATCTTTACCTGCAAGGCCCTCAATCGACGAGGTTGAGG
CAGCCAATGAATTGATCCAGACCGTTGACAAAGAAGAAGAGGCTAGGCTTGATGCCATCTACCGACAGCGCAAGGGCCCTGATGTACCTGAAGAACTCTT
CTCCATTCTCGTTGAGATGCACAACAACTCTGTCTCTTTCCATAGCAAGGACCAGAAGCGTGACGCCCTCAGATTGCTCGACCTAGAAAACCTCCACTCT
CATTTCGACCATCTTATCCAGAGAGCGTCCATGTGTTTACCTTCTTCCTCATCCTCTTCATCTACCCCTAACTATGCTACTGCTTCCGTTTCCGGCGGAA
CTACTTCTAAGCCTGCAGCCGGATCTTCGTCTTCCTACGCCATAAAATCCAGCTTTCATTTTTCGGAAAAGCAGCCTGTCGGAGCTGCGGATTTGGTTAC
CAGAGATGACAGCTACATGAAGAAGGCGAAGCCATTATTTTACTCCGGAGGAGCCGCTACTGCTGCTGGAATTGGTGTTCGAGAACCAACCATACTGGAT
TCGACTCTGAGATCCACGATTATCGCTTCAGGTCAAGATGGCGGTGATAAATTGAGCCTCATTAAGCTAGCTAGTCTAATAGAAGTCTCTGCTAAGAAAG
GCGCTACGGAGCTCAACCTCGCGAGTAAGCTGATGGATCAGATTGAGTGGCTACCTGATTCAATAGGCAAGTTATCAAGTTTGGTAACATTAGATTTGTC
TGACAATAGGATTATGGTCTTGCCACCCTCCATTGGCGGCCTCTCTTCCTTGACAAGATTGGATTTGCATTCAAACAGAATTGCTCAGCTACCCGAATCC
TTTGGCGATCTACTTAGCTTGGTCTTTCTGGATCTCCGTGCTAACCAGTTATCCACACTTCCTGCTACAATTGGAAGATTGGCTCGCCTCGAGCATCTCG
ATCTGAGCTCAAACCGGCTCAACTCCCTTCCGGATTCAATAGGATCACTAGCTAGCCTGAAAAACCTGAGCATTGAGACGAACGAGGTTGAAGAGCTTCC
ACATACTGTGGGACATTGTACTTCGCTGAAGGAGCTCCATGCAGATTACAACAAGTTGAAAGCCCTTCCGGAAGCTGTGGGTAAGATAGAGACATTAGAG
GTTCTCTCTGTGCGCTACAACAACATCAGACAGCTGCCAACAACCATGTCATCTCTAGTGAATCTGCGGGAGCTTGACGTGAGTTTCAATGAGCTGGAGG
CAGTGCCTGAGAGCTTGTGTTTCGCCACTTCTCTTGTGAAGATCAATGTTGGAAACAACTTTGCTGACCTTAGGTTCCTCCCGAGGTCCATTGGGAATCT
TAGCAACCTTGAGGAGTTGGATATAAGCAACAATCAGATTCGCATTCTTCCAGACTCATTCCAGCTGCTTTCCCGGTTGCGGGTCCTGCGAGTGGAAGAG
AATCCTCTTGAGATGCCGCCAAGGCATGTGGCTGACAAAGGCGCACAGGGAGGTCTTCAACATGTATCATGGTTCCAGTTTCTCCCTCATGAATCTGAGT
TGATCTCTGTATCTGATAAGAGTTTAAAAATTGATCAAAAAGATGCTGCTATGTGGTTGGTGCTTTCATCCCATTTGCAGTTGCAGAAGGGGGGATTTCT
GAGTACATGGACCAATTCTTTTGTTGGACCTTGGGATCCATCTCAAGGTCTTCACAATCCAGATGAGAAAATCAAGCTTTGGCTTTTTCTCCCCGGGCGT
CATTCTTCTGTTGTCGAGAGAGCTCAGTCAGCAGTTGCTAAATTAAGAGTTGTTGCATCTGGAATTTGGGTGGCACCTGGGGATTCTGAAGAGGTTTCTT
CTGCACTTTCTCAGTCTTTAAGGAATCGCATTGAAAAGGCACTTCTTGGAGTGTCTTACATGAGATATGGAGACGTATTTTCAAAGCATCGTCCATGTCA
AACTGAATCACTTTACAGGAGAGGACAGCCTACTATTGAATTCGTGTTTGCAGCCACTGAAGAAGCTATTTTTGTCCATGTTATAATATCTGCAAAGCAT
ATTCGCATGCTTTCAGCTGGTGATATCGAAAGTGTGTTGAAGCACTCATCTAAAGATTCTAGCTACAGACTTCCAGTCATCGTTTCTCCTCATGGAATAC
GTGGCTGGGTTACGGGATGTTGTCCAAATGATGTTATCAAGCAAGTCTATGCAAGTTCTGGCCACTCTAGGATGCAAGATGGTTTTACTGCTTCACTAAA
GGATGTCTCTCAAGGGCCTGCTTTCCAGCCGAGGGGGCAGCATTTATATGTTGAAGTTGCACTTGGTTGCCCTAGATATGAGAGTAGCAAGTCATTGCAA
TTGGGCTCCAGTTCAGGTAAGAATCTACAAAAAAACCATGCAGCAGAATCTCCAGCTGTGGCAAGAGGCGATCACAGTGGTTTACAAGATCACCCCTCTG
TGAATGAGAAGATATTCATCTATCCGGTTGAGGCTGTTGCTGTCCCTCTTTTACACACCTCATTTGCTCGGTCTTCTCTCAGACGATTTTGGTTACAAAA
TTGGGCAGGGCCATCAATGTCCGGTGCAGCTTTCTTTATGCATTGTGGAGCTAATATAGACAATGCAGATGGTTCTTGGCTCGACTCTGGCGGAGCACTT
GTGCACCATGGCTACAACAGTAGTAGCAATAGTAACAATAGTAGCATTAGCTTGATCAGCAACAGCTCCAGTGATAGTGATTATAATATGGCCATAGATG
GAGGGGACCTAGACGCAGATGCAGATTCGTTGTCATGCAGGCGATCTGGTTTGTCTTCAAATGATCATCTAGAAAATGATCTCCATAAACTGGGTTCTAA
GCGTACTCGGGCAGGAATGGAATCGTTTGGTCAGTTAGGTAGAGCTAAAAGTGCTTCTATTCAAGAAGCTTACAAGTCTGATTTGGGGTCCATTGAGGTT
AACACTTCAGCTATCACCGGGGTTGCAAATGAACTGAGTGGGTCTCCTTGGGATTGGGATGACGAGAGAGGTATGGGCATGGATATTCAAGCTCTACTTT
CCGAGTTTGGTGATTTCGGTGACTTCTTTGAGAATGATGCTTTGCCTTTTGGAGAGCCACCTGGAACTGCAGAGTCGGAGGCTCTAATGTTTTCAGGTCC
AGATGTTGGAGAAGTTGGCAGCAGTCCCGTTGGGTCAATGGATGTTGATCAAATGGTTTTACCTGTTGGTCTTCCATCTTTCGATAGCTTTAATCTGCCT
CCTTCAGTAACCATGGATGAGTGTGTGAGCAAAAATGAAGAAATGACTCACGGCTGTGTGAGTACAAGCTCAACAAATTCTGCTGCGCCCTCTTGCACGG
GTGAGTTTGATCATTTAATCAAGGCTGAAGCTCTTATGACATTTGCTTCAGAGTATGGAGCAGTTGAAACAGCTACTAGTGAGCTCTCTTCCTCAATCTT
CAAAAGGCCATATTGTCCCAAGTCTCGTAGTATAGAGAGCTCAAATTCAAATTCAAGCAATTACACATATGGTGCAACTCCTCCCCTTTCTCCTTGTTTT
GATGGGTTAGATGACAAGACTACTAGGCATGTTAACCTAAAAGCAGGTACTGGAAGAAGGGACACAAGAAAGTTTTACTCTCACGTTGAAGCTAGTAGAG
ACCAATACGAAGACTGTATTGGCAACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016684 pacid=23149047 polypeptide=Lus10016684 locus=Lus10016684.g ID=Lus10016684.BGIv1.0 annot-version=v1.0
MDATQVLPVDEVVAEIMKIHRSLPARPSIDEVEAANELIQTVDKEEEARLDAIYRQRKGPDVPEELFSILVEMHNNSVSFHSKDQKRDALRLLDLENLHS
HFDHLIQRASMCLPSSSSSSSTPNYATASVSGGTTSKPAAGSSSSYAIKSSFHFSEKQPVGAADLVTRDDSYMKKAKPLFYSGGAATAAGIGVREPTILD
STLRSTIIASGQDGGDKLSLIKLASLIEVSAKKGATELNLASKLMDQIEWLPDSIGKLSSLVTLDLSDNRIMVLPPSIGGLSSLTRLDLHSNRIAQLPES
FGDLLSLVFLDLRANQLSTLPATIGRLARLEHLDLSSNRLNSLPDSIGSLASLKNLSIETNEVEELPHTVGHCTSLKELHADYNKLKALPEAVGKIETLE
VLSVRYNNIRQLPTTMSSLVNLRELDVSFNELEAVPESLCFATSLVKINVGNNFADLRFLPRSIGNLSNLEELDISNNQIRILPDSFQLLSRLRVLRVEE
NPLEMPPRHVADKGAQGGLQHVSWFQFLPHESELISVSDKSLKIDQKDAAMWLVLSSHLQLQKGGFLSTWTNSFVGPWDPSQGLHNPDEKIKLWLFLPGR
HSSVVERAQSAVAKLRVVASGIWVAPGDSEEVSSALSQSLRNRIEKALLGVSYMRYGDVFSKHRPCQTESLYRRGQPTIEFVFAATEEAIFVHVIISAKH
IRMLSAGDIESVLKHSSKDSSYRLPVIVSPHGIRGWVTGCCPNDVIKQVYASSGHSRMQDGFTASLKDVSQGPAFQPRGQHLYVEVALGCPRYESSKSLQ
LGSSSGKNLQKNHAAESPAVARGDHSGLQDHPSVNEKIFIYPVEAVAVPLLHTSFARSSLRRFWLQNWAGPSMSGAAFFMHCGANIDNADGSWLDSGGAL
VHHGYNSSSNSNNSSISLISNSSSDSDYNMAIDGGDLDADADSLSCRRSGLSSNDHLENDLHKLGSKRTRAGMESFGQLGRAKSASIQEAYKSDLGSIEV
NTSAITGVANELSGSPWDWDDERGMGMDIQALLSEFGDFGDFFENDALPFGEPPGTAESEALMFSGPDVGEVGSSPVGSMDVDQMVLPVGLPSFDSFNLP
PSVTMDECVSKNEEMTHGCVSTSSTNSAAPSCTGEFDHLIKAEALMTFASEYGAVETATSELSSSIFKRPYCPKSRSIESSNSNSSNYTYGATPPLSPCF
DGLDDKTTRHVNLKAGTGRRDTRKFYSHVEASRDQYEDCIGN
|
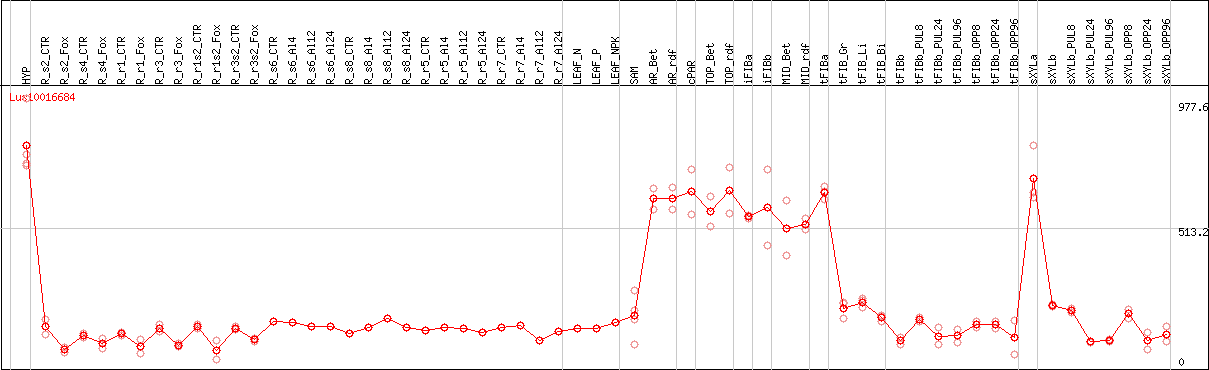
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016684 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
