External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G01630 313 / 2e-108
Sec14p-like phosphatidylinositol transfer family protein (.1)
AT1G14820 135 / 9e-39
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
AT1G75170 103 / 3e-26
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
AT4G36640 101 / 2e-25
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT4G08690 98 / 4e-24
Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT1G22180 89 / 8e-21
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
AT4G09160 88 / 8e-20
SEC14 cytosolic factor family protein / phosphoglyceride transfer family protein (.1)
AT4G39180 78 / 3e-16
ATSEC14, SEC14
ARABIDOPSIS THALIANA SECRETION 14, Sec14p-like phosphatidylinositol transfer family protein (.1.2)
AT4G34580 77 / 4e-16
SRH1, COW1
SHORT ROOT HAIR 1, CAN OF WORMS1, Sec14p-like phosphatidylinositol transfer family protein (.1)
AT5G63060 76 / 4e-16
Sec14p-like phosphatidylinositol transfer family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022428
463 / 6e-168
AT1G01630 323 / 1e-112
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10004684
342 / 1e-119
AT1G01630 326 / 2e-113
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10040252
338 / 3e-118
AT1G01630 319 / 8e-111
Sec14p-like phosphatidylinositol transfer family protein (.1)
Lus10001584
151 / 4e-45
AT1G14820 320 / 2e-111
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10003699
147 / 2e-43
AT1G14820 316 / 1e-109
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10034577
128 / 2e-36
AT1G14820 257 / 3e-87
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10026020
100 / 3e-24
AT1G75170 397 / 4e-139
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10010804
98 / 3e-24
AT1G75170 393 / 7e-139
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Lus10016312
97 / 5e-24
AT1G75170 387 / 2e-136
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G063966
315 / 9e-109
AT1G01630 340 / 2e-118
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.001G068000
301 / 9e-104
AT1G01630 361 / 3e-127
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.003G162100
299 / 7e-103
AT1G01630 356 / 2e-125
Sec14p-like phosphatidylinositol transfer family protein (.1)
Potri.008G135600
154 / 2e-46
AT1G14820 323 / 2e-112
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.010G105400
143 / 5e-42
AT1G14820 326 / 1e-113
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.015G023000
141 / 4e-41
AT1G14820 276 / 5e-94
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.007G025800
108 / 5e-28
AT1G75170 408 / 2e-144
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.005G123200
108 / 7e-28
AT1G75170 414 / 9e-147
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.007G025900
100 / 5e-25
AT1G75170 369 / 3e-129
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
Potri.002G261000
99 / 2e-24
AT1G75170 427 / 3e-152
Sec14p-like phosphatidylinositol transfer family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0214
UBA
PF03765
CRAL_TRIO_N
CRAL/TRIO, N-terminal domain
CL0512
CRAL_TRIO
PF00650
CRAL_TRIO
CRAL/TRIO domain
Representative CDS sequence
>Lus10016733 pacid=23149033 polypeptide=Lus10016733 locus=Lus10016733.g ID=Lus10016733.BGIv1.0 annot-version=v1.0
ATGTTCCGTAATGGCGATAAACCATCAGATCCAGATGACAAGCTTGAAGTTGAGCGGACCAAGTTGAGTCTCTTGAGGGACTTGGCCCAGACCAAGGATC
CTTCTTCCAAGGAAGTAGATGATGGGACACTGAGGAGGTTTCTGAGGGCAAGAGATATGGATGTAGAGACTGCATGTGCAATGTTCCTCAAATACCTCAA
CTGGAAGCACAGTTTCCTCCCCAATGGCACCTCCTCTGTTTCCACCTCCGACATCTCCATCGACATTTCCCACAACAAGGTCTTCCTCCAAGGTACTGAT
CGGACTGGCCGACCCATCTCCGTTGCTTTCGGAGGTCGCCATTTCCAGAACGGCGTTGACGAATTCAAGCGTTATGTAGTTTATGTCATCGAGAAACTCT
GCGCAAGTATGGCGGAAGGACAAGAGAAATTTGTGGTGATCGGAGATCTGGAAGGTTGGGGTTATGCAAACAGCGACATTAGGGGTTATCTGGCTGCTCT
GCACATTTTGCAGGATTACTACCCGGAAAGACTTGGTAAGCTGTTCATTGTGAATGTGCCATACATTTTCATGGCAGTGTGGAAAGTTATTTACCCCTTC
ATCGACAGCAATACCAAAAGAAAGATTGTATTTGTGGATAACAAGAAGCTGAAATCCACACTTGTAGAGGACATTGATGAAAGCCAGCTGCCAGAGATAT
ACGGAGGCAAACTGCCTTTAGTTCCTATCCACCTCACCTGA
AA sequence
>Lus10016733 pacid=23149033 polypeptide=Lus10016733 locus=Lus10016733.g ID=Lus10016733.BGIv1.0 annot-version=v1.0
MFRNGDKPSDPDDKLEVERTKLSLLRDLAQTKDPSSKEVDDGTLRRFLRARDMDVETACAMFLKYLNWKHSFLPNGTSSVSTSDISIDISHNKVFLQGTD
RTGRPISVAFGGRHFQNGVDEFKRYVVYVIEKLCASMAEGQEKFVVIGDLEGWGYANSDIRGYLAALHILQDYYPERLGKLFIVNVPYIFMAVWKVIYPF
IDSNTKRKIVFVDNKKLKSTLVEDIDESQLPEIYGGKLPLVPIHLT
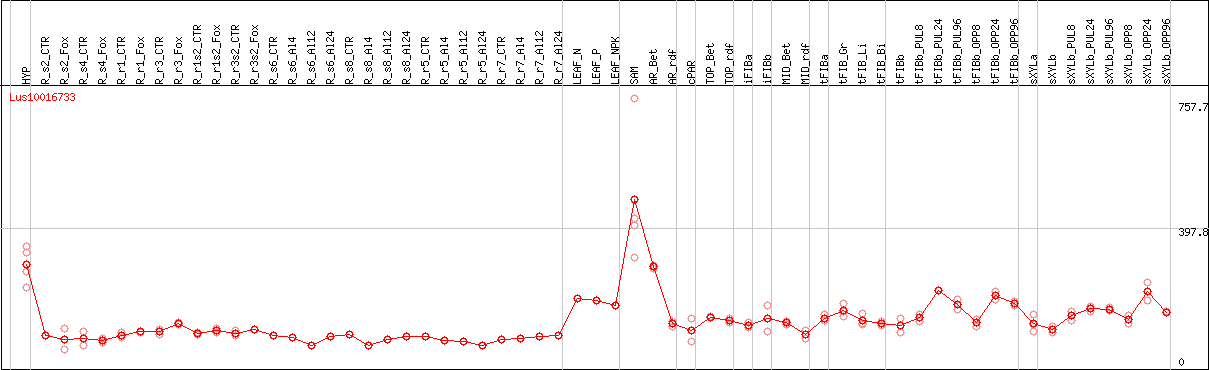
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016733 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.