Lus10016753 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Lus10016753 pacid=23149061 polypeptide=Lus10016753 locus=Lus10016753.g ID=Lus10016753.BGIv1.0 annot-version=v1.0
ATGGAAAACGATCGTATGCTATACCAAAGCAGCGTCTTCTCCCGGCTAAAACCTTGCTGTCTAGAGCTCCTTCAGCTTCTCCAAAACCCTAACAAAGTTT
CTTCCTCTGCATTACCTTCACTGCTCCAACTCCTCCGCAGCTGTTCATCTAGTGATTTGCAGTCTTTCTTCGACTACACCTCGTTCCCGTTGCTGCTCCT
ACTTGACGCTGCGGTCAGTTGCAGATCGTCCAATAAGGATGATCCTCAGAAGTATTCGGTGACCTTCAATCCGCAACAATCGTCTGATAGTGTTAGCGAC
AAAGTGGCGGAAGGTGTGCTGCAGTGCTTGGAGGAGCTTCTCCAGAAATGTCGTTTTGGATCGGTCGACCAGATGGTTGTTGTGATGAAGAAATTGACTA
ATGCCGCGTTGCTGTCTCCATCAGAGGCTTCAGAAGAATTTCGTGAGGGCGTAATCAAGTGTTTAAGGAAATTACTTATGGGCTTAAGTCCGTGTGCAGA
TGAAACTTGTTCATGCAGACAAACGCATGGTATGCCAGCATTGTTGGAGAATGGGGAGCTCCACTGGTCTCTGTCCAAGCCTTTGAGTTATGATACAGAA
GTGAAAGAGTGTGTGATTGCATTTCTCCAATCACATGCCGCTTCTGCCGCCGTCGGACACTTGCTGTCACTGCTACTCAAGGCAGCAGACAGTGAGGCTG
CCCGAGGACATCGAGGCAGTGCAAGGCTCCGTGTTGAAGCCTTTTATACCTTACGTGTGCTTGTAGCCAAGGTCGGTTCTGCAGATGCATTGGCTTTCTT
TTTACCTGGTGTTGTCAGCCAATTTTCAAAAGTTTTGCATGGCTCAAAAACTATGATGAGTGGAGCAGCTGGAAGCACAGATGCTACTGATCAAGCAATA
AGAGCGTTGGCAGAGTATCTAATGATAGTGCTTGATGGTGTTTCCAATGTAAGCTCCATTATAACCGAGAATCTCCCAGCTGATGCAAGTTCGGACAGTA
ATAGATCTGGCGAGTCCATATTGGATCAACTCCGTCACTTACCAGTCAGCAGTCAGGGGCATATGAAGAATATGGCTGATGAGTCAATAGTAGATCTGGA
AAACTCAGTAACTCGTGCGTCTGAATACGAAATGAGTAGGGGCATGAAAAGAAGCAGGTACACAGGATCATTGCATGTTGAGCGAACAAGAGATTGGTTA
AGGCAAACTTCAGCGCATGTGGTTAAATTGTTGAGTGCCACATTTCCACATATCTGTGTTCATCCTGTGAATAAGTTGAGATGGGGACTACTGGCTGCCA
TCAAAGGATTAATATTAAGGTGCAGTTCTACTTTGAAGGAAAGCAGATCTATGCTACTAGAGTGCCTTTTGGTCTTGGTTGTTGATGACTCTGAAGATGT
TGCTGGATCTGCCAAGGAGTTCCTTGAGTTCTTATTCTCATCGTGCGGAAAGCATTCAGTAAAAGCTGATATTGGTCCGATTTTTGCTAGGATAATTGAA
AAGCTTCCAAAGGTAGCACTTGGTGGTGACGAATCAGTGGCTCTCTTCCATGCACAACAATTATTAGTTGTCATTTACTATTCTGGTCCTCTATTTGTTT
TGGACCACCTCCAAAATCCGGTTACTGCAGCAAGACTCTTGGATGTGTTTTCCCTCTGTCTGAGTCCGAGTTCTGTATATGCTGGCGCTCTTGATAAGGA
CACATTGACAAGGACATCATCCATTAGTTTTCTTCCATCTATTATGGAGTTGAGATGTGGTTCTCTTGCCCCTAGTTACCAGAATCTCAAAGATCAGTCT
CTTGTAAATGTGTCCGATACAAGTTTTATTCATAGGAAACAAATTGAGTATCCTTTTGAAACTCTCGGGCACAGTTATGAACTTCCACGAATACCACCTT
GGTTTGTCCATGTGGGTTATAAGAAGTTATATGACACACTGGCTGGAGTACTCAGACTAGTAGGCTTATCTTTTTACTCTCTCTTGTTGCTAGACCACCA
TGAAGGGCATACGGTTGCTATTACAGATATACCACTAGGTCATATACGCAGATTAGTTTCTGAGGTTCACTCCAGAGATGCAAATGACGGCTGGCAGTAT
TTGTATAATGGAACTTGGGCTGGAAAGTTAATACGTCAGGCGAGCACTTCGGCCTGCATTCTAAATGAGATGATATTTGGTGTATCTGATAAAGCAGTTG
AAGGCATTACTATGTTGTTTCAAAATTCCAGGGAGAAGGGGGGAGAAGTTGTACTGTCTGATGTGAGAGGTTCTAGTGCTTGGCCGCCTTCACTTCAGAG
CAGAGAAAGTATGGCTTCTGTTTGGAAGGCCTCACAACAAACTGTCAAAAAGAGTCACTTAATTGATTGTATTGGGAGAGTCTTACATGAGTATCTATCA
TCTGAATTGTGGAATTTACCCATAGATCACAGGGATCTAGTGGGACAATGTGACGAGGAAGTTGAAGTTAACTTGCACTTATTCCAAGATGCAGCTGCCC
TACAACAGGTTGCTATTGATGGAATTGGCATCTTTGCGACCTGCCTTGGAAATGATTTCTCCTCCAGTGGATTTCTTCACTCTACTCTTTATCTGCTGCT
TGAGAGCCTCATATCCGCTGATATCCAAGTCAGAACTGCTTCTGATACTGTCCTGCATGTTCTTTCTGCTACCTGTGGCCATCCAACAGTTGGGCAGTTG
GTCTTGGCCAATGCCGATTATGTTGTAGATTCAATATGTCGGCAATTACGTCACTTGGATCTTAACCCACAAGTCCCGAACGTGCTTGCTTCTATGCTTT
CTTATATTGGAGTGACTAATAAAGTACTGCCCCTTTTGGAGGAAACGATGGGCTCTGTTTCAAGAGAACTAGAGATTCTTAGCAGACATCAGCACCCAGA
GTTGACCACTCCTTTCCTAAAGGCCATAGCACAAATTTGCAAGGCATCACAATCTGAGGCTTCATCACTGCCATGTAATGCAGCATTGTATCTGAAGGAT
GTGAAGTCCAAAGTCTCAAGGTTGGAGAAGAAAGTGAGAGCTCAATCCAATAAAGGTCCAACATCATCTTGTGATGACAATGTTGGTACTTGGCATGGAG
AATCAGAACAATGGGAGGGTATAATCTTCAAGTTGAATGAATCTAAGAGGTACAGACGTACAGTTGGATCTATTGCTGCTTCCTGCTTATTATCCGCGAC
TCCTCTACTTGGTTCAATGGAGCAAGCGGCATGCCTGGTGGCATTGGATATAGTTCAGGACGGCATTGCATCCCTGGCAAAAGTGGAAGAAGCTTATCAA
TATGAGAAAGAAACCAAGGAAGTAGTCGAAGAAGTAGTGCAAGCATATTCAACTTATCAGCTTCAGGATACTATTGATGCGGCCGATGAAGGGATTGACG
AGAACAGATTGCTTCCAGCAATGAATAAGATATGGCCATTCTTGATTTCTTGCGTTCGTAACAAAAATCCTGTGGCAGTTAGGAAATGCGCAACTGTGAT
AGCCAATGTGGTGCAGACATGTGGAGGGGACTTCTTCACACGGCGGTTCCACACTGATGGTCCGTGCTTGTGGAAGCTTCTTTCCTCATCTCCGTTTCAG
AGGAAGCCATTATTGAAAGATGAAAGAAGGACACTACGACTTCCTTATAGGAGCAATTTAGCCGCAAAGGAGGAGGAATCAATGGCTGAAACGTCCAACT
TGAAGGTCCAGGCTGCGGTGCTGAAAATGGTTGCTGATTTGGCACGAAGCAAGAGGAGTTGTAGGGCATTGGAACTGGTACTGAAGAAGGTGAGCGGTCT
AGTTGTTGGGATAGCTTGCAGCGGCGTCATTGGGCTTCATGATGCCTCGGCGAGTGCTCTTGATGCACTTGCGACTATGGACCCTGATCTGATATGGCTT
TTAGTAGCCGATGTTTACTACTCATTGAAGGAGGAGAATGCCACATACCCTCCCAGCAGCTGCTCACGTTTGTTGGAAATATCTAGGCTTTTACCACCGC
CTGTTTCACCCAAGGAGTACCTTTATGTGCAGTATGGAGGGCAAAGCTATGGTTTTGACATCAACGTAGCTTCAGTGGAGAAAGTATTCCGTTCATTGGC
TTTTCCAGACCAACACAGGTTGTGA
|
|||||||||||||||
|
AA sequence
|
>Lus10016753 pacid=23149061 polypeptide=Lus10016753 locus=Lus10016753.g ID=Lus10016753.BGIv1.0 annot-version=v1.0
MENDRMLYQSSVFSRLKPCCLELLQLLQNPNKVSSSALPSLLQLLRSCSSSDLQSFFDYTSFPLLLLLDAAVSCRSSNKDDPQKYSVTFNPQQSSDSVSD
KVAEGVLQCLEELLQKCRFGSVDQMVVVMKKLTNAALLSPSEASEEFREGVIKCLRKLLMGLSPCADETCSCRQTHGMPALLENGELHWSLSKPLSYDTE
VKECVIAFLQSHAASAAVGHLLSLLLKAADSEAARGHRGSARLRVEAFYTLRVLVAKVGSADALAFFLPGVVSQFSKVLHGSKTMMSGAAGSTDATDQAI
RALAEYLMIVLDGVSNVSSIITENLPADASSDSNRSGESILDQLRHLPVSSQGHMKNMADESIVDLENSVTRASEYEMSRGMKRSRYTGSLHVERTRDWL
RQTSAHVVKLLSATFPHICVHPVNKLRWGLLAAIKGLILRCSSTLKESRSMLLECLLVLVVDDSEDVAGSAKEFLEFLFSSCGKHSVKADIGPIFARIIE
KLPKVALGGDESVALFHAQQLLVVIYYSGPLFVLDHLQNPVTAARLLDVFSLCLSPSSVYAGALDKDTLTRTSSISFLPSIMELRCGSLAPSYQNLKDQS
LVNVSDTSFIHRKQIEYPFETLGHSYELPRIPPWFVHVGYKKLYDTLAGVLRLVGLSFYSLLLLDHHEGHTVAITDIPLGHIRRLVSEVHSRDANDGWQY
LYNGTWAGKLIRQASTSACILNEMIFGVSDKAVEGITMLFQNSREKGGEVVLSDVRGSSAWPPSLQSRESMASVWKASQQTVKKSHLIDCIGRVLHEYLS
SELWNLPIDHRDLVGQCDEEVEVNLHLFQDAAALQQVAIDGIGIFATCLGNDFSSSGFLHSTLYLLLESLISADIQVRTASDTVLHVLSATCGHPTVGQL
VLANADYVVDSICRQLRHLDLNPQVPNVLASMLSYIGVTNKVLPLLEETMGSVSRELEILSRHQHPELTTPFLKAIAQICKASQSEASSLPCNAALYLKD
VKSKVSRLEKKVRAQSNKGPTSSCDDNVGTWHGESEQWEGIIFKLNESKRYRRTVGSIAASCLLSATPLLGSMEQAACLVALDIVQDGIASLAKVEEAYQ
YEKETKEVVEEVVQAYSTYQLQDTIDAADEGIDENRLLPAMNKIWPFLISCVRNKNPVAVRKCATVIANVVQTCGGDFFTRRFHTDGPCLWKLLSSSPFQ
RKPLLKDERRTLRLPYRSNLAAKEEESMAETSNLKVQAAVLKMVADLARSKRSCRALELVLKKVSGLVVGIACSGVIGLHDASASALDALATMDPDLIWL
LVADVYYSLKEENATYPPSSCSRLLEISRLLPPPVSPKEYLYVQYGGQSYGFDINVASVEKVFRSLAFPDQHRL
|
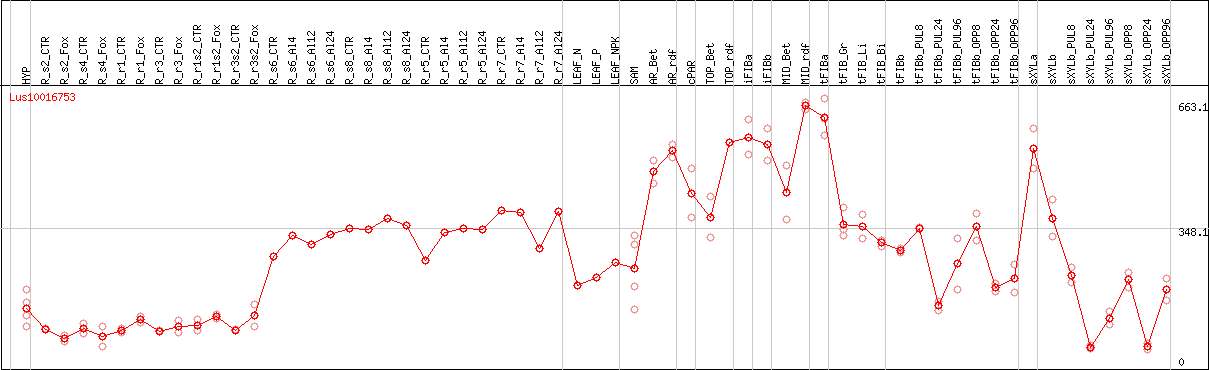
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016753 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
