Lus10016788 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10016788 pacid=23149138 polypeptide=Lus10016788 locus=Lus10016788.g ID=Lus10016788.BGIv1.0 annot-version=v1.0
ATGGAACTTCTTCCTCAGTGTGGTGTTCATGATGGCGGTGAAGCTGATTCTAGTCAACATAACATTGGAGCAGGTAAAAGCCACGAAGAAGGAAACCAAG
TAGAAAATGCCAGTGTTATAGTCAGTGAATCAATGCAAAACCTTGGAGGGTCTGAAGTCGTAGGACAAACTAGAGCTTCAGGAAGTGAGAGAATCTCTTA
TGAGTATCATGATTTTGATGAAGATGACTCGAGTCTACAGCAGGGCACAAGCGAATCTTGTGCAGCTCCTATGAAATCTAATGTTGTTCTACATACTATG
GAAATTGAACCAACCAACTCTAGGGAGGGAGATCATTCTTTCTCTGAACCTTTAGCTCTATGGGTAAAGTGGAGAGGGAAGTGGCAGGCAGGAATCAGAT
GTGCAAGAGCTGACTGGCCGCTGTCAACTTTGAGAGCGAAACCCACTCATGATCGGAAGCAGTACTTCGTCGTATTTTTCCCGAATACCAGAAATTATTC
CTGGGCAGATATGTTACTTGTCCGATCGATAAGTGAAACTCCTGAGCCGATTGCATATAAGTCGCACAAAGGTGGACTTAAATTGGTTAAAGATTTGAGC
GTTGCACGACGGTTCATAATGCAAAAACTAGCCGTTGGGATGCTGAATATCGTTGATCAATTTCATAATGAGGCTCTGGTAGAATCTGCTCGCTCCGTGC
TGGTGTGGAAGGAGTTTGCTATGGAAGCATCACGCTGTAGTGGTTATGGTGATCTCGGAAGAATGCTTCTCAAGCTTCAAAATATGATTTTGCAGAAGTA
CATAGAGGCTGATTGGTTGTCCAGTTCTTTTCAATCATGGGTGCAACAGTGTCAGGTTGCATGTAGTGCTGAATCAATCGAGTTGTTGAGAGAGAAATCT
ATGATTCTGCAGCAACATGCATTGCTCTTGCTGATGTCTTCTGTTTCTCCGCCTCCCTTCCTGTCACAACAGGAACTATATGATTCCATCTTGTGGAATA
AAGTCAATTCTCTTGGGGATGCATCAGTGCAATCAACATTAGGCTCTGAGTGGAAAACCTGGAAGCATGAAGTTATGAAATGGTTTTCAACATCTCACCC
CTTATCTGCTGGTGGAGAGACAGAGCGTAGACGTTCTGGTAGCCCTTCAGAAGTAGGACATCAAGTAAATCTGAAAAGGCCCAAGCTTGAAGTTCGCCGT
GCAGAACCACATGCTGGGCAAGTGGAAATGAGCTATCCACGGCAAACTATGACAGTGGATATTGATTCCGAGTTCTTCAGCAAGCGAGAAACTACGAGCA
GTGCACCTTTGTCAACCGGACTTTCAAGAGTGGAAGATATCCAAAAAGGAGCTATGCCAATAGATTCTCCTAGTAGTGTGCCTGGTAGATGGGATGGAAT
TGTTGTAGAAGCTTCAAGTTCATTGCCTAGCTCTAGCAAGGGGGCAGAAAAAACTCCTCCCAGTGGCCTCATGGCTATGAATGCTACAGATCCTGGTAAT
GGTAGAAACAGGCAGTGCACTGCTTTCATTCAGTCTAAGGGTAGGCAATGTGTGAGGTGGGCAAATGAAGGTGATGTCTACTGCTGTGTGCATCTAGCCT
CCCGTTTCACAGGTGGCTCTACCAAGGGAGAGGTAAGTTCTCCAGTCCATAACCCAATGTGTGACGGAACCACAGTACTTGGCACTCGATGCAAACACCG
TTCCCTGCCAGGTTCTTCATTCTGCAAGAAACACGGACCAAAGACAGATACCAATTTACTACAGAATTCACTCAAAAGAAAGCACGGTGTATTTGTTTCC
ACTCCAGAAACATCACAGTGTCGGGATATAGTGCTATTTGAACAAGCTGAAAGTCCCCTTCAAGTGGAGCCTGTCTCAGTCGTGGCATGTGATAATGGTG
ATTCATACCATGCAATAAATGGTTTATCGGGTATGTTTGATGAGTCCAGTATGAATTATCCTGGTAAAGAGGTGCAGCATTGCGTAGGTTCAAACGCGTT
GGATAACAATGCTCCTTGTCATGAAACCCCGAAGCGCTATACATTATACTGTGATAGCCACATCCCAAGCTGGCTTAAGCGTGCAAGGAATGGTAGGAGT
AGAATAATATCAAAAGAGGTATTTCTGGATCTCTTGAAAGGTTGTTGCTCTGTGGAGCAAAAGTTGCATTTACACAAAGCATGTGAGCTATTTTACAAAC
TCTTCAAGAGCATTTTGTCTCTAAGAAACCCAGTTCCGTTGGAGGTTCAACTTCAGTGGGCCCTGACTGAAGCTTCTAAAGATTTCGGGGTGGGGGAGTT
CTTGCTGAAGTTAGTTTGCACTGAAAAAGAGAGACTGGGAAGCATTTGGGGCCTTGGTGGTGATGAAGATCCACATGTTTTACCCTCTCGTGTTGGTAAA
TCAGCAGGTTTACCCACACCTATTGATGGTGGTCATGAAGATGGACAATATATTAGGTGCAAGATTTGCTTTGATAAGTTTACTGATGACCAAGAACTTT
GCAGCCACTGGATGGACAATCATAAAAAGGAGGCACAATGCCTCTTCAGAGAGCATGCTTGTGCTATTTGTCTTGATTCTTTTACAGACAAGAAAGGTTT
AGAAGCACATGTGCAGGACAGACATCATGTGCAGTTTGTTGAACAATGCATGCTTCTTCTTTGCATCCCATGTAAAAGTCACTTTGGCAATGCTGAACAG
TTATGGTCACATGTCCTCTCAGTTCACCCACTTGATTTTAGAGTGGAAAAATCTGATCAACAGCTTATTGTTTCTGCACCTGATCCTAAGCAGAAACTCA
AGCAAACAGCTGAGCTGGATAATGGATCTTTTGACAAACCTACCACTAGTAATCCAAATGGTGTACGGAAATTTATTTGCAGGTTCTGTGGTTTGAAGTT
CGACCTCCTGCCTGATCTTGGTCGCCACCATCAGGCTGCTCATATGGGACCAAATATATTGAACTGTCGACCCCAAAAGAAAAGATTTGGCTACTATGCT
TATAGATCAAAATCTGGTAGACTTACCCGTCCAAGATTTAAGAAAGGTATGGCTACAGCTGCATACAGGATCAGGAATAGGGCTAATGCTAGTTTGAAGA
AACGTATCCAGGCTACTAAGTCACTGAGTTCCATGGACTTAAATGTTCAGCCTCGCTTAAGTGATCCAACGGCTCTTGGTCGGTTGGCCGAATCTCAATG
CTCAGCAGTTGCTAAGATTCTGTTTTCTGAGATTCAGAAGACCAACCCTCGCCCAAATAACCCTGAAATTTTAGCTACTGCACGCTCTGTGTGCTGCAGG
GTGAGCCTAAAAGCGTCCCTTGAGGGCAAATATGGACTATTGCCAGAACGTATTTATCTCAAAGCAGCCAAACTGTGCAGTGAGCATAATATTCAAGTGG
AGTGGCATCATGAAGGATTTGTTTGTCCCAGAGGATGCAAGAGTTATAAGGACCTCGAGTTGCTATCACCCTTGATCCCGCTTCCTAATGTTCCCACAGG
AAAACGGTCAGATAATTCTTTGAATCATGTCCACGATGATTGGCTGGTGGATGAATGCCACTATATTATTGATTTGGATGATGTTAGAGAGGAAAGTAAG
CGAAAGGTAGTTGTCCTGTGTCAGGATATCAGCTTTGGTAAGGAGTCGATACCAATTGCTTGTGTGGCAGACGAGCAGTTAATCAATTCCCTTGATGGCT
CGGCAGATAACTCAGATGTTCAAAACACTGTTAGGCCTTGGGAATCTTTCACCTACCTAACTAACACATTATTTAATCATTCCAGCGATCCTGATTTTGA
GAGTTTGCAGGTGGGATGCTCCTGTCCATATTCAGTGTGCTGTTCTGAAACATGTGATCATGTTGACCTCTTTGATGATGATTATGAAGATGCAAAAGAC
ATATATGGAGCGCCAATGCATGGAAGATTTCCATATGACGATAAAGGAAGGATAATCCTGGAGGAGGGATACCTTGTTTATGAGTGTAACAATATGTGTA
GTTGCAATAATACATGCCCAAATAGAGTACTGCAGAATGGAGTACGTGTGAGGTTGGAAGTTTTCAAAACAGAAAATAAGGGCTGGGCAGTCAGGGCAGG
TGAGCATATCCTACGTGGCACATTTATATGTGAATACATAGGAGAAGTGTTAGGCGAAGATGAGGCGAATAAGAGACGAATAAGCTATGGCGAAGGTGGT
TGCAGCTATATGTATAGTATTGATGATCATACGAATGACATGAGCAGAATGATACAAGGGGAAGCCAACTACGTCATTGATGCCACAGCTTATGGAAATG
TGTCGCGGTTCATAAATCACAGCTGCTTACCAAATCTTGAAACCCATCAAGTTATTATCAATAGCATGGATTCTCTGCGCTCTCATATTGGCCTTTATGC
AAGTCGAGATATAGCTTATGGTGAAGAACTGACGCACAACTACCGGTACAATATTGTGTCTGGACAAGGTCATCCATGCCATTGTGGTACCTCCAGGTGT
TGCGGCCACCTGCACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10016788 pacid=23149138 polypeptide=Lus10016788 locus=Lus10016788.g ID=Lus10016788.BGIv1.0 annot-version=v1.0
MELLPQCGVHDGGEADSSQHNIGAGKSHEEGNQVENASVIVSESMQNLGGSEVVGQTRASGSERISYEYHDFDEDDSSLQQGTSESCAAPMKSNVVLHTM
EIEPTNSREGDHSFSEPLALWVKWRGKWQAGIRCARADWPLSTLRAKPTHDRKQYFVVFFPNTRNYSWADMLLVRSISETPEPIAYKSHKGGLKLVKDLS
VARRFIMQKLAVGMLNIVDQFHNEALVESARSVLVWKEFAMEASRCSGYGDLGRMLLKLQNMILQKYIEADWLSSSFQSWVQQCQVACSAESIELLREKS
MILQQHALLLLMSSVSPPPFLSQQELYDSILWNKVNSLGDASVQSTLGSEWKTWKHEVMKWFSTSHPLSAGGETERRRSGSPSEVGHQVNLKRPKLEVRR
AEPHAGQVEMSYPRQTMTVDIDSEFFSKRETTSSAPLSTGLSRVEDIQKGAMPIDSPSSVPGRWDGIVVEASSSLPSSSKGAEKTPPSGLMAMNATDPGN
GRNRQCTAFIQSKGRQCVRWANEGDVYCCVHLASRFTGGSTKGEVSSPVHNPMCDGTTVLGTRCKHRSLPGSSFCKKHGPKTDTNLLQNSLKRKHGVFVS
TPETSQCRDIVLFEQAESPLQVEPVSVVACDNGDSYHAINGLSGMFDESSMNYPGKEVQHCVGSNALDNNAPCHETPKRYTLYCDSHIPSWLKRARNGRS
RIISKEVFLDLLKGCCSVEQKLHLHKACELFYKLFKSILSLRNPVPLEVQLQWALTEASKDFGVGEFLLKLVCTEKERLGSIWGLGGDEDPHVLPSRVGK
SAGLPTPIDGGHEDGQYIRCKICFDKFTDDQELCSHWMDNHKKEAQCLFREHACAICLDSFTDKKGLEAHVQDRHHVQFVEQCMLLLCIPCKSHFGNAEQ
LWSHVLSVHPLDFRVEKSDQQLIVSAPDPKQKLKQTAELDNGSFDKPTTSNPNGVRKFICRFCGLKFDLLPDLGRHHQAAHMGPNILNCRPQKKRFGYYA
YRSKSGRLTRPRFKKGMATAAYRIRNRANASLKKRIQATKSLSSMDLNVQPRLSDPTALGRLAESQCSAVAKILFSEIQKTNPRPNNPEILATARSVCCR
VSLKASLEGKYGLLPERIYLKAAKLCSEHNIQVEWHHEGFVCPRGCKSYKDLELLSPLIPLPNVPTGKRSDNSLNHVHDDWLVDECHYIIDLDDVREESK
RKVVVLCQDISFGKESIPIACVADEQLINSLDGSADNSDVQNTVRPWESFTYLTNTLFNHSSDPDFESLQVGCSCPYSVCCSETCDHVDLFDDDYEDAKD
IYGAPMHGRFPYDDKGRIILEEGYLVYECNNMCSCNNTCPNRVLQNGVRVRLEVFKTENKGWAVRAGEHILRGTFICEYIGEVLGEDEANKRRISYGEGG
CSYMYSIDDHTNDMSRMIQGEANYVIDATAYGNVSRFINHSCLPNLETHQVIINSMDSLRSHIGLYASRDIAYGEELTHNYRYNIVSGQGHPCHCGTSRC
CGHLH
|
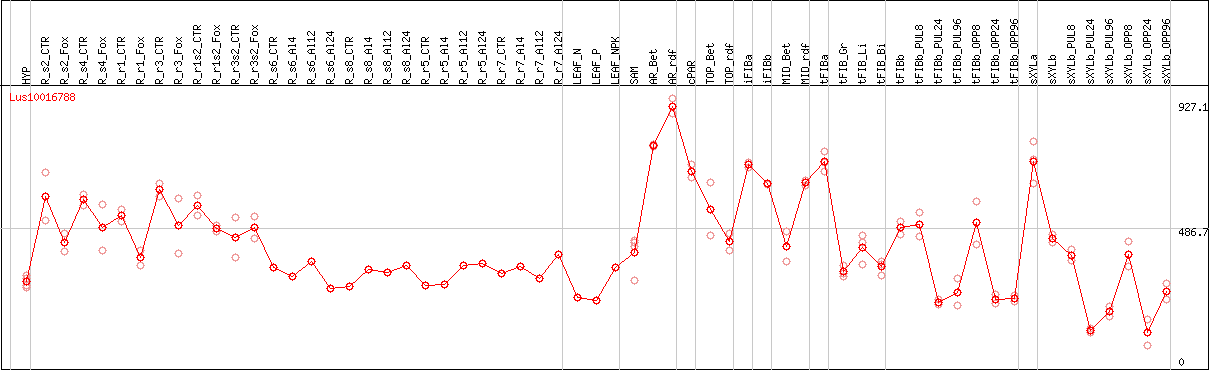
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016788 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
