External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G23770 212 / 1e-63
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
AT2G33580 199 / 3e-58
Protein kinase superfamily protein (.1)
AT3G21630 168 / 3e-47
LYSMRLK1, CERK1
LYSM DOMAIN RECEPTOR-LIKE KINASE 1, chitin elicitor receptor kinase 1 (.1)
AT1G51940 167 / 9e-47
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
AT1G79620 164 / 3e-45
Leucine-rich repeat protein kinase family protein (.1)
AT2G28970 162 / 2e-44
Leucine-rich repeat protein kinase family protein (.1)
AT2G25220 157 / 2e-44
Protein kinase superfamily protein (.1.2)
AT4G29990 162 / 3e-44
Leucine-rich repeat transmembrane protein kinase protein (.1)
AT2G17220 156 / 3e-44
Kin3
kinase 3, Protein kinase superfamily protein (.1.2)
AT1G67720 160 / 1e-43
Leucine-rich repeat protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022488
613 / 0
AT2G23770 391 / 2e-128
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10018788
249 / 5e-77
AT2G23770 312 / 3e-97
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10016299
248 / 6e-77
AT2G23770 368 / 3e-119
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10032735
248 / 2e-76
AT2G23770 305 / 1e-94
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10000577
231 / 2e-70
AT2G23770 490 / 1e-166
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10019661
222 / 4e-67
AT2G23770 483 / 8e-164
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10041171
200 / 3e-59
AT2G33580 224 / 1e-64
Protein kinase superfamily protein (.1)
Lus10008586
199 / 2e-58
AT2G33580 594 / 0.0
Protein kinase superfamily protein (.1)
Lus10021889
197 / 2e-58
AT2G23770 256 / 2e-77
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G128300
352 / 3e-117
AT2G23770 394 / 2e-129
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.014G040000
291 / 5e-94
AT2G23770 345 / 1e-110
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.005G128200
248 / 6e-77
AT2G23770 514 / 4e-176
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.007G032100
246 / 4e-76
AT2G23770 513 / 6e-175
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.005G259600
218 / 1e-65
AT2G33580 637 / 0.0
Protein kinase superfamily protein (.1)
Potri.010G078700
204 / 2e-60
AT2G23770 288 / 5e-89
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.008G160600
194 / 9e-57
AT2G33580 217 / 8e-62
Protein kinase superfamily protein (.1)
Potri.008G187500
183 / 7e-53
AT1G51940 362 / 7e-117
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.014G156400
181 / 5e-52
AT3G21630 678 / 0.0
LYSM DOMAIN RECEPTOR-LIKE KINASE 1, chitin elicitor receptor kinase 1 (.1)
Potri.011G010000
180 / 1e-51
AT3G21630 673 / 0.0
LYSM DOMAIN RECEPTOR-LIKE KINASE 1, chitin elicitor receptor kinase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF07714
PK_Tyr_Ser-Thr
Protein tyrosine and serine/threonine kinase
Representative CDS sequence
>Lus10016794 pacid=23149059 polypeptide=Lus10016794 locus=Lus10016794.g ID=Lus10016794.BGIv1.0 annot-version=v1.0
ATGTCTTTGATCCTAACAGGAGCATTTTTGGTCTACCATAAGAAAAAAGGCCTTGAAACTCTAAAGGGGAAAGACAGGAAGAAGAAATTGACCTTGCCAG
AGGATTTCCTTGATAGCATGTCCACAGTTGATCAAGGGTTGAGAATCTATGACTTGGAGGAGTTAAAGCTAGCAACTGCCGATTTCAGCCATGACAATCA
GTTGAGTGACCGTGTGTACAAGGGTGTTACAAATGGACAAGAAGTAGCTATCAAGAAGATGAGCAGAAATGTTTCAGGTGAAGTGAATTTGCTGAGAAAG
ATCAACCATTTCAATCTGATAAGCCTTCTTGCTGCTGCATATGAGCACCAGGGTGAATACTACCTGATCTACAAGTACATGGAGAATGGTTCTTTGAAGG
ATTGGTTACAGAAAGGTAGTTTACATCTAGTCCTAAGTTGGAGCCATAGGCTTCAGATTGCTTTGGATATTGCTAATGGACTTCACTATCTTCATAAGTT
TACTGAACCTCCATATGTTCACAAAGATATCAGCAGCAGCAATATTCTGTTAGATGCTGATTTGAGGGCTAAGATTGCCAACTTCAGTCTTGCAAGGTCT
GCAGAGGCAGACTTGTATGCAAGTTCTTCAATCAGGCTTGCAAAGGGAACAAAAGGTTATCTGGCACCTGAGTACTTAGAGTATGGCTTGGTGATTCCTG
AGATTGATGTGTATGCATTTGGGGTCGTTTTAACTGAGCTCATCACTCGAAAAGAAGCCGTATTTTGGCAGAATGAAAGAGAAGATTCAGAGTCCGAGCT
TTGCAACTTCATAGATCCTAGCCTGCATCCAAAGCAAAAAATGAGCCTTGTGCTGAGAATGGTTAGGTTGGTTCTAATGTGCTTGGCTAAAGAACCTGAA
GATAGACCAAGCATGGGTGAAATAGTTTCTTCACTTTCAAAGATACAACTGGAAGTGTATAGATCCGACTTTATGCATTGTAGCAGTTCATAG
AA sequence
>Lus10016794 pacid=23149059 polypeptide=Lus10016794 locus=Lus10016794.g ID=Lus10016794.BGIv1.0 annot-version=v1.0
MSLILTGAFLVYHKKKGLETLKGKDRKKKLTLPEDFLDSMSTVDQGLRIYDLEELKLATADFSHDNQLSDRVYKGVTNGQEVAIKKMSRNVSGEVNLLRK
INHFNLISLLAAAYEHQGEYYLIYKYMENGSLKDWLQKGSLHLVLSWSHRLQIALDIANGLHYLHKFTEPPYVHKDISSSNILLDADLRAKIANFSLARS
AEADLYASSSIRLAKGTKGYLAPEYLEYGLVIPEIDVYAFGVVLTELITRKEAVFWQNEREDSESELCNFIDPSLHPKQKMSLVLRMVRLVLMCLAKEPE
DRPSMGEIVSSLSKIQLEVYRSDFMHCSSS
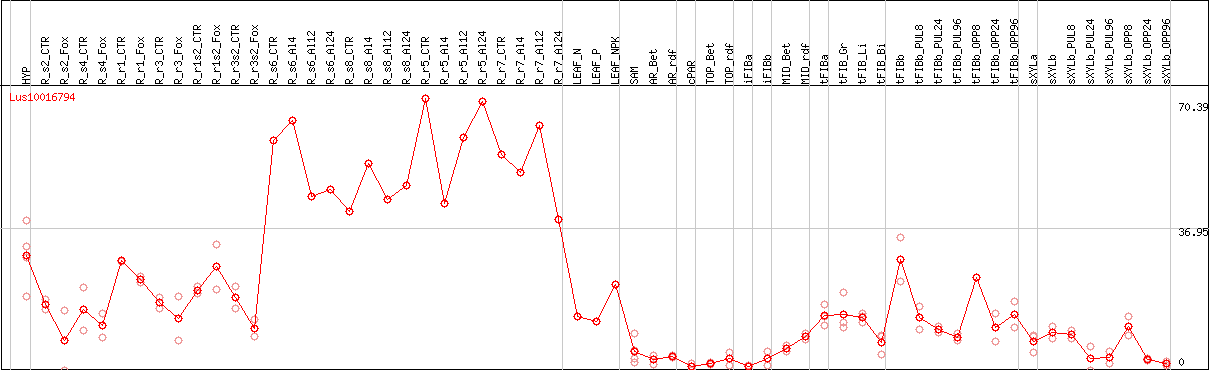
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016794 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.