External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G04820 87 / 2e-20
OFP
ATOFP13, OFP13
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
AT2G36050 81 / 4e-18
OFP
ATOFP15, OFP15
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 15, ovate family protein 15 (.1)
AT3G52540 74 / 4e-15
OFP
ATOFP18, OFP18
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 18, ovate family protein 18 (.1)
AT1G79960 62 / 3e-11
OFP
ATOFP14, OFP14
ovate family protein 14 (.1)
AT2G18500 59 / 6e-10
OFP
ATOFP7, OFP7
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 7, ovate family protein 7 (.1)
AT5G19650 56 / 3e-09
OFP
ATOFP8, OFP8
ovate family protein 8 (.1)
AT4G14860 55 / 4e-09
OFP
ATOFP11
ovate family protein 11 (.1)
AT5G22240 54 / 8e-09
OFP
ATOFP10, OFP10
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 10, Ovate family protein (.1)
AT1G05420 53 / 3e-08
OFP
ATOFP12, OFP12
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
AT2G30400 53 / 6e-08
OFP
ATOFP2, OFP2
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 2, ovate family protein 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021303
280 / 7e-94
AT5G04820 103 / 8e-26
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10043433
78 / 7e-17
AT5G04820 146 / 2e-42
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10034150
77 / 1e-16
AT5G04820 134 / 8e-38
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10010554
62 / 2e-11
AT1G05420 104 / 2e-27
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10006120
61 / 2e-10
AT1G05420 103 / 6e-26
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10006912
56 / 2e-09
AT1G05420 112 / 2e-30
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
Lus10035891
57 / 3e-09
AT1G79960 166 / 2e-48
ovate family protein 14 (.1)
Lus10043406
55 / 4e-09
AT5G04820 64 / 1e-12
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10014677
55 / 6e-09
AT1G05420 109 / 2e-29
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 12, ovate family protein 12 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04844
Ovate
Transcriptional repressor, ovate
Representative CDS sequence
>Lus10016979 pacid=23173019 polypeptide=Lus10016979 locus=Lus10016979.g ID=Lus10016979.BGIv1.0 annot-version=v1.0
ATGTTGGGGAAGAAGAAATTAAGATTAAATCCCATCCCTTCCTTTCTCTTCTCCAAATCCAACTCCTCAGATCAATGCTCTTCTTCTCCTTCCTCTTCCT
CATCCTCTGTTAAATCATCCTCTGTTTCCTCCTCCTCCTGGCCTTGGACTCCAAACTACTACTACTGCCACCAACAGCCTCGCACTCTCTCCTTCCGCAC
CATCACCTCCGCCGACATGTTGGATACTAAATACCCACCAGAATCTGGACCTGGGTCTGGACCCAGCAAGCCCGATTCGTCGTTTTTCCCCAACACCCCT
GCCGTCGCCAGCGACGGCGGTGAAGACCCGGCAGTCGAGAGCGTGATTCGCGGCGCGAAATCGGAGCGGTTGTTCTTCTCTCCTGGAGAGACTAGCTCAT
CGTTACTGCAAGATTCAGAAAAGGGGAAGATCGGCGGCGACGGGAGTGGAGGAGAGGCGGGGGATGCTCTGTTTTCGATTAAGGAGGGCGGAGAAGAAGC
CGGAGCGGTGGCGGTGGGGGTGGATTCTAAGGATCCGTTCGAGGATTTCAAGAACTCGATGGTGGAGATGGTGGAGGCTCACGGGTTGAAGGATTGGGAG
TCTTTAGAGAATCTATTGAGCTGCTACCTCAGGGTTAATAAGAAGAGCAACCACGGATACATCGTCGGGGCTTTCGTTGATTTGCTCGTCAGTTTGCCGC
CGTTTGGTCCTAAATCGACTCCGGGCGCGGGGGTGGCGGGGCGGGGGGAGGAGGGAGGGGTGTGA
AA sequence
>Lus10016979 pacid=23173019 polypeptide=Lus10016979 locus=Lus10016979.g ID=Lus10016979.BGIv1.0 annot-version=v1.0
MLGKKKLRLNPIPSFLFSKSNSSDQCSSSPSSSSSSVKSSSVSSSSWPWTPNYYYCHQQPRTLSFRTITSADMLDTKYPPESGPGSGPSKPDSSFFPNTP
AVASDGGEDPAVESVIRGAKSERLFFSPGETSSSLLQDSEKGKIGGDGSGGEAGDALFSIKEGGEEAGAVAVGVDSKDPFEDFKNSMVEMVEAHGLKDWE
SLENLLSCYLRVNKKSNHGYIVGAFVDLLVSLPPFGPKSTPGAGVAGRGEEGGV
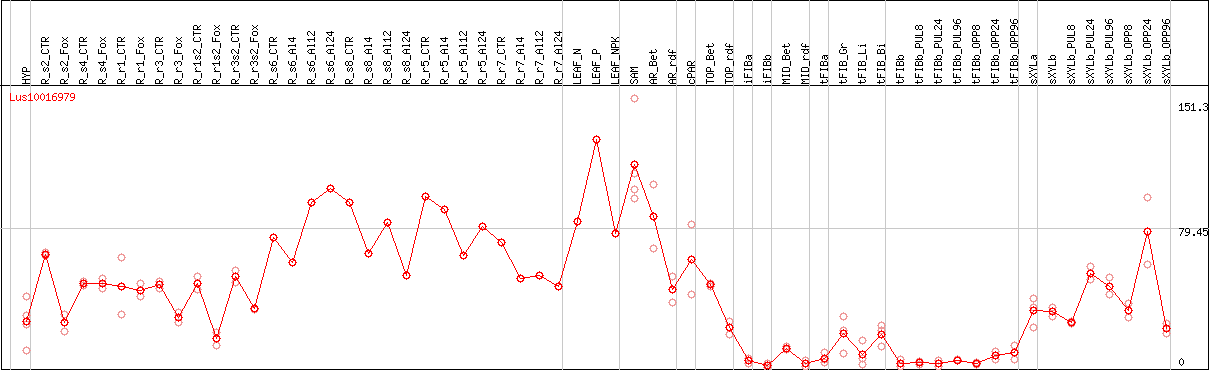
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10016979 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.