External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G02400 305 / 4e-103
ATGA2OX4, ATGA2OX6, DTA1
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
AT1G47990 274 / 4e-91
ATGA2OX4
Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 4 (.1)
AT1G30040 251 / 7e-82
ATGA2OX2
GIBBERELLIN 2-OXIDASE 2, gibberellin 2-oxidase (.1.2)
AT2G34555 229 / 2e-73
ATGA2OX3
gibberellin 2-oxidase 3 (.1)
AT1G78440 213 / 6e-67
ATGA2OX1
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
AT1G78550 114 / 8e-29
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G21420 110 / 1e-27
LBO1
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G51810 110 / 2e-27
AT2353, GA20OX2, ATGA20OX2
gibberellin 20 oxidase 2 (.1)
AT1G80330 108 / 7e-27
ATGA3OX4
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 4, gibberellin 3-oxidase 4 (.1)
AT5G07200 106 / 7e-26
ATGA20OX3, YAP169
ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 3, gibberellin 20-oxidase 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037816
536 / 0
AT1G02400 298 / 4e-101
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
Lus10025016
308 / 2e-104
AT1G02400 367 / 4e-127
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
Lus10009998
306 / 2e-103
AT1G02400 362 / 3e-125
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
Lus10004511
303 / 4e-102
AT1G02400 347 / 3e-119
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
Lus10023311
195 / 3e-60
AT1G30040 381 / 6e-133
GIBBERELLIN 2-OXIDASE 2, gibberellin 2-oxidase (.1.2)
Lus10038498
159 / 1e-47
AT1G78440 258 / 2e-86
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
Lus10006350
125 / 6e-33
AT4G25420 486 / 1e-172
GA REQUIRING 5, ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10006722
122 / 1e-31
AT1G60980 489 / 5e-174
gibberellin 20-oxidase 4 (.1)
Lus10015016
119 / 8e-31
AT4G25420 494 / 3e-176
GA REQUIRING 5, ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G149700
373 / 1e-129
AT1G47990 318 / 4e-108
Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 4 (.1)
Potri.008G101600
372 / 2e-129
AT1G47990 350 / 2e-120
Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 4 (.1)
Potri.002G191900
347 / 2e-119
AT1G02400 394 / 7e-138
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
Potri.014G117300
335 / 1e-114
AT1G02400 367 / 2e-127
DOWNSTREAM TARGET OF AGL15 1, ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 6, Arabidopsis thaliana gibberellin 2-oxidase 4, gibberellin 2-oxidase 6 (.1)
Potri.001G378400
267 / 3e-88
AT1G78440 404 / 1e-141
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
Potri.011G095600
268 / 6e-88
AT1G78440 404 / 4e-141
Arabidopsis thaliana gibberellin 2-oxidase 1 (.1)
Potri.004G065000
252 / 2e-82
AT1G30040 448 / 6e-159
GIBBERELLIN 2-OXIDASE 2, gibberellin 2-oxidase (.1.2)
Potri.015G002800
118 / 8e-31
AT5G24530 521 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G355100
116 / 9e-30
AT1G17020 439 / 1e-154
senescence-related gene 1 (.1)
Potri.011G164200
115 / 1e-29
AT4G10500 305 / 4e-102
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10017094 pacid=23167390 polypeptide=Lus10017094 locus=Lus10017094.g ID=Lus10017094.BGIv1.0 annot-version=v1.0
ATGGTGGTGGCATCTCCTCCGCCTCTAGCCGGCACCGAGAAGGCGGCGGCACTCCCCCGCCTCCCAACGGTTGACATGTCGGGCCCACGCGATGAGGTGG
CCCGACAAATAGTGAAGGGGGGAGCAGAAGATGGCGAGGAGGGACTCGGGTTCTTCGGCCGACCCGCCCAGGAAAAGCAGCGGGCCGGACCCGCCGACCC
TTTCGGGTACGGGTGCAAGAACATCGGGTTCAACGGCGACGTCGGGGAAGTCGAGTACGTGATGCTGAATACCGATCCTCGCTCCATTGCGGAGAGGTCG
TTGACCATCAACTCCAACGACCCGACGACGTTCAGTTCCACTGTGAGCGGGTACATTGAAGCGGTAAAGGAAGTAGCATGCGAGATCCTAGATTTGATGG
CGGAAGGGATGTACATACCTGATACGTCAGTGTTTAGTGATCTCATCAGGGACGGCCATAGTGATTCTGTTCTCCGTCTGAATCACTACCCTCCATTTCT
CCTCCCTTCACCACCAACAGTCGATCACGCTGACGCCGAAGTTGCGACGATAGGATTTGGGGCGCATACTGACCCTCAGATCCTAACCATCTTAACATCC
AACGACGTTCCTGGCCTCCAGATCCACGTCGACCCGAACATGTGGGTCCCCGTATCGCCTGACCCCACCGCCTTCTTCGTTAATGTTGGTGACGTATTAC
AGGCGATGACAAATGGAAGAATGATAAGCGTAAAGCATAGGGCCCTCAACCTCAACTCGCCAAAGTCCAGGATGTCAATGGCGTATTTTGCAGCCCCACC
ACTAGAAACCACCATCTCTGTTCTGCCTCAAATGATTACTCCTTCCAGCCCTAATTGTCTGTTCAGACCTTTCACCTGGGCTGATTACAAGAAGGCAGCT
TACTCCCTTCGCCTTGGAGATTCACGCCTCGACCTTTTCACTGTTACTAACTCCTAA
AA sequence
>Lus10017094 pacid=23167390 polypeptide=Lus10017094 locus=Lus10017094.g ID=Lus10017094.BGIv1.0 annot-version=v1.0
MVVASPPPLAGTEKAAALPRLPTVDMSGPRDEVARQIVKGGAEDGEEGLGFFGRPAQEKQRAGPADPFGYGCKNIGFNGDVGEVEYVMLNTDPRSIAERS
LTINSNDPTTFSSTVSGYIEAVKEVACEILDLMAEGMYIPDTSVFSDLIRDGHSDSVLRLNHYPPFLLPSPPTVDHADAEVATIGFGAHTDPQILTILTS
NDVPGLQIHVDPNMWVPVSPDPTAFFVNVGDVLQAMTNGRMISVKHRALNLNSPKSRMSMAYFAAPPLETTISVLPQMITPSSPNCLFRPFTWADYKKAA
YSLRLGDSRLDLFTVTNS
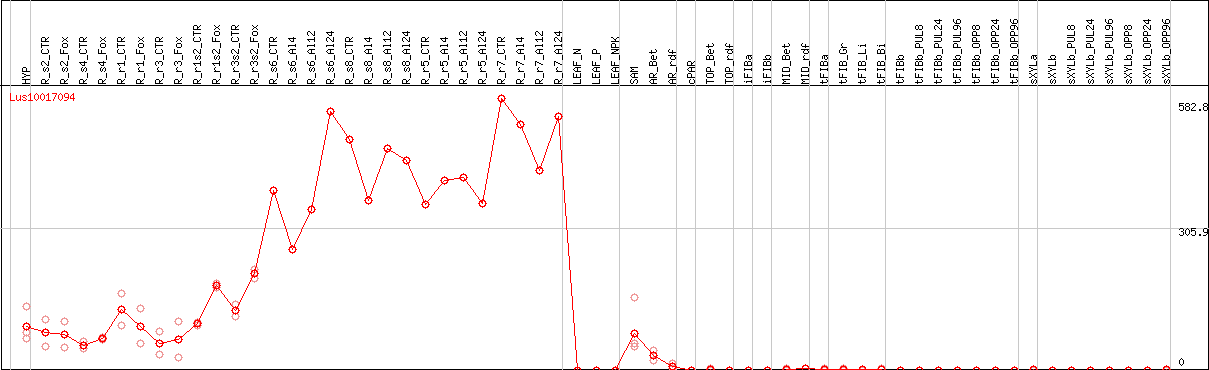
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017094 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.