Lus10017162 [FLAX]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10017162 pacid=23167488 polypeptide=Lus10017162 locus=Lus10017162.g ID=Lus10017162.BGIv1.0 annot-version=v1.0
ATGTTGTCGAGAGTCGAACATGGGAAGAAGACTAGGGATGATGAATCAGTAGAAAATAATTTGTTATTGAATGACTTGCAAACTATAAGCAAAGCTCTGA
ACCTTGACAAACAAAACCCATCCACGACTTTACTTCCTACTGCCAGTAGGGAGGAGTCAGGAGAGGAGAAATTGTTACTGAATGAGTTGCAAACTATAAG
CAAAGCTCTGTACCTTGATAAACCAAAACCTTCGAGGACTTTACTTCCCACCGCCAGCAGCCGAGCAAAAGCAACTGGAAGACCACCAACCTTAGTCGAA
CCCAAAACAAAACCCAAGCCTGGGGGGCTTGTTAATGACGAGTCCTCACGCAAGGACAAGAAGTCGTCTATATGGAACTGGAAAGCATCATTCAGAGCCC
TCTCCAGTGTTAAGAACCGCAAGTTCAATTGTTGTTTCTCTCTTCAAGTTCATTCCATCGAAGGCCTTCCCTCGACTTTGGACGGTTGCAGCTTGTGTGT
AAATTGGAAGTGGAGGGATGGGGAGTTGGTGACACGTCCTGTGAAATCTTTTGAAGGGACTGCAGAATTTGAAGAGAAGTTAGTGCATACTTGTCTTGTT
TATGGCAGTCGGAGTGGTCCTCATCATTCTGCAAAGTATGAGGCCAAACATTTCTCGCTCTTTGCATCTATCTTTCGTATGCCTGAGGTTGATTTGGGGA
AGCACAGGGTTGACCTTACTAGGTTGCTTCCTCTCACATTGGAGGAATTGGATGATGAGAAGAGTTCTGGAAAGTGGACTACAAGTTATAAGTTGTCGGG
AGAAGCAAAGGGTGCAGTGATGAACTTGAGTTTCAGTTATACAGTTGTTGGAGAGTCTCCGACTGGAAATGGTCACAATGTTCCCGAGGAAGTAAGTATC
AAAAGGAATAATCTGAAAACAGCTAAGACAATGCCTAGACTTGGCCAATCTGAAGGAAAGGTTGCATTACATCGTCATGAAAGTCTTACCAATACTTCTA
ACAAGCAGTTTCCTGCAAAGCCTCAGCCGGTGAAAAAGGAAAAAGATCTTCATGAAGTTATGCCTGACAAAAATCTAGAACTAGATACGACAAGAAACGT
GATGAATGGGAGATTTGATGTCGATAAGTTGGATGCTTCAACAAGTGAAAATGAGCATGATGATAAGCTGCAGTTTCCAACCATATCTGAATCTGGGGAA
GCCAATGTTGAAAATGAAAATGGGAATGTTGAATTTTCTATTGCTGAAGAAGGCAACTATTTGCTAAATGAGAGCAACGAAGATGGGAATGTTGAATTTT
CTATTGCTGAAGAAGCAAAGGTTTTGCCAAGTGAGAGCAGTAGAACGGAGACCCTTACCTTCCAGGATGTTGGCGTTTCTATTCCAGAGACTGATTTTGC
TTCCAAGTTTCAGGCAATTCCTCAAGAGTCTATTAATTGTCAGGCTGAGGAAAAAGGAGATGCTGGTCAAAAAGATGAACTTTTTACCTGTGACTCTGAC
TCTGATGATAATCTATGTTCAAAAGAATATGTGATGAAAGAATTAGAATCTATGTTAAACAGTGTTTCAAATCTTGAAACTGAAGCACTTGATTCTTCTG
ATGACCTGAAAATTGACAATCTAGTAGACAGGGACAAAAGCATAGTTAACCCGGACGATGCTGATTCTATTGCAACTGAATTTTTAGACATGTTGGGGGT
TCAGCATAGTCCATTTGGCTTGAGTTCTGAGAGTGAACCTGAGTCCCCAAGGGAGCAGTTACTAAGGCAATTTGAGAAGGATGCCTTGGCTGGTGGCTTT
TCTTTGTTTGACTACAATGCAGATGATTGGGATGATCCGGCGAGTACAAATAATGAGATGAATGAGTCAGGATATGACCAATCGGTATGTGGTTTTGATC
TGTCAGCATTCACGGATGATGCAGAAGAAGACCATCATAAAGGAACTCACTCTCAGGGTAAAAAGATTGGAGCTAAGATGTTGGAGGACTTTGAGACAGA
GGTTTTAATGCATGAATGGGGCCTGAATGACAAGACATTTGAGTGTTCTCCATCCAAGACTTGTGATGGCGTTGGGGTTCAATGTGATCCATCTGTAGAA
GAGCCATATAAATTACCACCTCTATCAGAAGGTTTAGGTCCTTTTCTGCAGACAAAGAATGGTGGTTTTGTACGCTCTATGAATCCGTCACTTTTTGGGA
GTGCGAAAAGTGGTGGGAGCTTGATTATACAAGTCTCCAGTCCTGTTGTGGTACCTGCAGAAATGGGCGCTAATGTTACTGATGTACTTGAGCAACTCGC
GTCAGTTGGAATTGAGAAGCTCTCTATGCAGGCAAATAAGCTAATGCCTTTGGAGGATATCACTGGGAAAACAATGCAACAGGTTGCTTGGGAAGTTTCT
GGTGCTTTAGAGGGACATGATGGGGGGAGTTCTTTCCCGCAGAAACATGAGTTAGGGCATGAGGGATCAGATATGCCAGAGAGGACGAAGAAAAAGCGTT
CTAGCTCAACAGGAACTGAGATTGGCTCGGAATATGTGTCTCTCGATGACCTTGCACCATTAGCTATGGATAAGATTGAAGCACTTTCATTGGAGGGTTT
GAGAATACAATCCGGGCTGTCTGGCGAGGAAGCAACCGTAGGGGAAATTTCAAGTCTCCATGGCAAAGGAGTTAATGTTGGGAGTTCCCTTGGGTTGGAA
GGAACTGCTGGAATGCAGTTGTTGGACATTAAAGACAACGATGGTGACGTTGACGGGTTAATGGGTCTTTCGTTGACTCTTGACGAATGGATGCGGTTGG
ACTCTGGTGATATTGGTGATGATAACGATCAGATAAGTGAACGGACTTCAAAGATACTTGCTGCTCATCATGCAAGCTCCTTGGAAATGCTGCAAGGGGG
TTCGAAGGGGGAGAAAAGGCGTGGAAAGGGATCCGGTAAAAAATGTGGTTTGCTGGGTAATAACTTTACAGTAGCATTAATGGTTCAACTACGTGATCCT
CTAAGAAACTACGAGCCTGTTGGGACGCCAATGCTTGCTCTGATTCAAGTTGAGCGAGTTTTCATCCCACCGAAGCCCAGAATATACTGCAAAGTAGCAG
AGTTGAGGGAGAGCGGCAACAACGAGGATGAAGAAGAGTCCGAGTTGACATCGAACGAGGCAAAGGACAGGAGAGTAGTAGAAGAAGAAGATAATAAAAA
GCTCTCTATGCAGGCAAATAAGCTAATGCCTTTGGAGGATATCACTGGGAAAACAATGCAACAGGTTGCTTGGGAAGTTTCTGGTGCTTTAGAGGGACAT
GATGGGGGGAGTTCTTTCCCGCAGAAACATGAGTTAGGGCATGAGGGATCAGATATGCCAGAGAGGACGAAGAAAAAGCGTTCTAGCTCAACAGGAACTG
AGATTGGCTCGGAATATGTGTCTCTCGATGACCTTGCACCATTAGCTATGGATAAGATTGAAGCACTTTCATTGGAGGGTTTGAGAATACAATCCGGGCT
GTCTGGCGAGGAAGCAACCGTAGGGGAAATTTCAAGTCTCCATGGCAAAGGAGTTAATGTTGGGAGTTCCCTTGGGTTGGAAGGAACTGCTGGAATGCAG
TTGTTGGACATTAAAGACAACGATGGTGACGTTGACGGGTTAATGGGTCTTTCGTTGACTCTTGACGAATGGATGCGGTTGGACTCTGGTGATATTGGTG
ATGATAACGATCAGATAAGTGAACGGACTTCAAAGATACTTGCTGCTCATCATGCAAGCTCCTTGGAAATGCTGCAAGGGGGTTCGAAGGGGGAGAAAAG
GCGTGGAAAGGGATCCGGTAAAAAATGTGGTTTGCTGGGTAATAACTTTACAGTAGCATTAATGGTTCAACTACGTGATCCTCTAAGAAACTACGAGCCT
GTTGGGACGCCAATGCTTGCTCTGATTCAAGTTGAGCGAGTTTTCATCCCACCGAAGCCCAGAATATACTGCAAAGTAGCAGAGTTGAGGGAGAGCGGCA
ACAACGAGGATGAAGAAGAGTCCGAGTTGACATCGAACGAGGCAAAGGACAGGAGAGTAGTAGAAGAAGAAGATAATAAAGCAGAAGAGATGGAAGGAAT
AGCGCGGTTCCAAATTGCAGAAGTACATGTTGCTGGTATAAAGACCGAACCTGAGAGGAAGGGTTGGGGTACAGCAACACAGCAACAAGCTGGTTCTCGT
TGGCTGGTAGCTAATGGAATGGGAAAGGGGAATAAAATGAAGTCAAAAGCTGGAAGCTCTGGTGATAGTGTGTGGAGTTTATCATCATCTCGTGTTCAGC
AACTGGGAGGAGGAAAGCAGATTAAGAGGAACCCAAATTTAGTTTTGCCTAGTAAGAAACTGAATTGGTTGAAGTAA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Lus10017162 pacid=23167488 polypeptide=Lus10017162 locus=Lus10017162.g ID=Lus10017162.BGIv1.0 annot-version=v1.0
MLSRVEHGKKTRDDESVENNLLLNDLQTISKALNLDKQNPSTTLLPTASREESGEEKLLLNELQTISKALYLDKPKPSRTLLPTASSRAKATGRPPTLVE
PKTKPKPGGLVNDESSRKDKKSSIWNWKASFRALSSVKNRKFNCCFSLQVHSIEGLPSTLDGCSLCVNWKWRDGELVTRPVKSFEGTAEFEEKLVHTCLV
YGSRSGPHHSAKYEAKHFSLFASIFRMPEVDLGKHRVDLTRLLPLTLEELDDEKSSGKWTTSYKLSGEAKGAVMNLSFSYTVVGESPTGNGHNVPEEVSI
KRNNLKTAKTMPRLGQSEGKVALHRHESLTNTSNKQFPAKPQPVKKEKDLHEVMPDKNLELDTTRNVMNGRFDVDKLDASTSENEHDDKLQFPTISESGE
ANVENENGNVEFSIAEEGNYLLNESNEDGNVEFSIAEEAKVLPSESSRTETLTFQDVGVSIPETDFASKFQAIPQESINCQAEEKGDAGQKDELFTCDSD
SDDNLCSKEYVMKELESMLNSVSNLETEALDSSDDLKIDNLVDRDKSIVNPDDADSIATEFLDMLGVQHSPFGLSSESEPESPREQLLRQFEKDALAGGF
SLFDYNADDWDDPASTNNEMNESGYDQSVCGFDLSAFTDDAEEDHHKGTHSQGKKIGAKMLEDFETEVLMHEWGLNDKTFECSPSKTCDGVGVQCDPSVE
EPYKLPPLSEGLGPFLQTKNGGFVRSMNPSLFGSAKSGGSLIIQVSSPVVVPAEMGANVTDVLEQLASVGIEKLSMQANKLMPLEDITGKTMQQVAWEVS
GALEGHDGGSSFPQKHELGHEGSDMPERTKKKRSSSTGTEIGSEYVSLDDLAPLAMDKIEALSLEGLRIQSGLSGEEATVGEISSLHGKGVNVGSSLGLE
GTAGMQLLDIKDNDGDVDGLMGLSLTLDEWMRLDSGDIGDDNDQISERTSKILAAHHASSLEMLQGGSKGEKRRGKGSGKKCGLLGNNFTVALMVQLRDP
LRNYEPVGTPMLALIQVERVFIPPKPRIYCKVAELRESGNNEDEEESELTSNEAKDRRVVEEEDNKKLSMQANKLMPLEDITGKTMQQVAWEVSGALEGH
DGGSSFPQKHELGHEGSDMPERTKKKRSSSTGTEIGSEYVSLDDLAPLAMDKIEALSLEGLRIQSGLSGEEATVGEISSLHGKGVNVGSSLGLEGTAGMQ
LLDIKDNDGDVDGLMGLSLTLDEWMRLDSGDIGDDNDQISERTSKILAAHHASSLEMLQGGSKGEKRRGKGSGKKCGLLGNNFTVALMVQLRDPLRNYEP
VGTPMLALIQVERVFIPPKPRIYCKVAELRESGNNEDEEESELTSNEAKDRRVVEEEDNKAEEMEGIARFQIAEVHVAGIKTEPERKGWGTATQQQAGSR
WLVANGMGKGNKMKSKAGSSGDSVWSLSSSRVQQLGGGKQIKRNPNLVLPSKKLNWLK
|
DESeq2's median of ratios [FLAX]
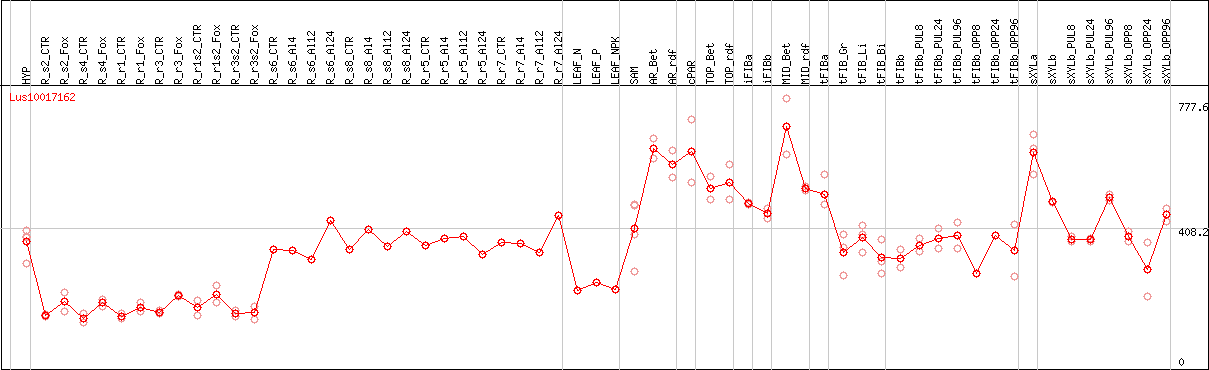
Coexpressed genes
Lus10017162 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
