Lus10017180 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10017180 pacid=23165832 polypeptide=Lus10017180 locus=Lus10017180.g ID=Lus10017180.BGIv1.0 annot-version=v1.0
ATGAAGGCGAATCGCAGAGCCAAGAAGCCGGACGCCTCCTCGAAGCTGGTTCTTGAAGAGATAATCGGGCTGACTACAAAGAACGCCAATGGAATCGCGT
CGACGGAATCCACTTCAGCGTGTGCTTATATCGCTGGATGTGCGGTGGTCCTCCACGACGTTGATTCCGGCCGACAGTCGCACCTCATCGCTTCTCATAG
AATGCCTAAGCCATTGAGCTGCGTCGCAATGTCACGAGATGGACGATTGGTTGCCGCGGGAGAGTCAGGAAATCAACCTGCAGTACTAGTATGGGATCGA
TCGACTCTGTCATTTTTATCTGATCTGAATGGCCATGTTTATGGAGTGGAGTGCATTGCTTTCTCGCCAGATGGGGAGCATCTTGTGTCTGTTGGAGGAT
ACATATATCTTTGGGATTGGAAGAACGGCGTGTTGGTGGCCAAGCTTAAAGCAAGTTCATCTTGCTCGGCTATCACATCAGTCAGCTTCACATCGAATGG
AAAATTTATGGTTACTGCTGGGAAGAAGCACTTGAAGTTTTGGACAGTTGGTTCGACATCGAAGAATACTCGATTAAACGCAGCTATAGGGGCACTGATA
ATTCAAGGGAAGCCTGTTAATCTCGGCCCACATAAAGGAAGCTCGTTTGTGTGTGTCGCTAGTTCTGCAAACGTTGAACCAGTAGGGACTATGTTCCCTA
TTTATGCTTTAACTGATGGAGGTCTCTTGTGTGTCATCGATAAGGGTTTGTCAGTAGCAAAGTCTGCGGACTTGAAGGTGGATAAAGGTCTTGCACTATC
TGCCTCCCGTAAGCTTATTGCTTGTGGTTGTTCTAATGGAATTGTGAAGCTCTTTAGCAATGAGACTCTCAACTATACTGGAACTCTGCTGCATTCCAAG
GCCAAAATTTGTGATGAGTCAATTGATAATCTTCGAAACACTGAAGACAGTGGAGAGAACCCGGTTCCTGCCTCTCCTGATTCTGTTGCTTGTCAGTTCT
CTGCCTCAGAGAAGTTGGTGGTCGTTTATGCAGATCATACCCTCTGTATATGGGACGTTCTTGATGTGAACCAGGCCTCCAGGTGTTGCATGCTGGTTTC
ACACTCAGCCTGTATTTGGGATGTAAAGAACCTGTGCTGTGAATACATGCACGATCCTTCTCTTGTGTGTGCTGCTAGAGGATGTGCTGGAGGAGTTTCT
TTTGCTACCTGCTCAGCAGATGGCACTATTAGGTTGTGGGATCTTGCACCACAACAACAATTTGTTGGAGATGATTCTGATCACCATGGTTTGAATGCTG
AACCATTAAGCCCTTCTTGCTTAGCAAGCACCGGAATTTTTGAGCGAGATACTCTGGATGTTGGTTTGAGCAGTCATGGCTTTCGTTGCATGGCAATTAG
TTCAGATGCACAGTATCTAGCTGCTGGTGATAATGAGGGAAACCTTCACATCTACAATCTGCGCACTTTTGATTATACGTGTTTCCAGGATGTACACCAC
TCAGAGATCCTATCATTGAGCTTCAGTGCATCACAAGAGGAACATGATGCCCCAGAAGTCGATCATGGCAATTACTTGCTTGTGTCAGGGGGAAGGGATC
GAGTTATCCATCTTTATGATGTCAAAAGGAACTTTAATCTCATTGCAAGTGTTGACGAGCACTCAGCTGCAGTGACCTCTGTGAAGCTTGATAGAGTGGG
TTGTAAGGCTATCAGTTGCAGCGCTGATAGATCACTGGTGTTCCGGGATGTTTTCGTGTCAGATAGTGGCTGTCAGATATTTCGGCGTCACCATCAAATG
GCATCTCTTGGCACTGTATATGATATGGCTGTTGATAAGAAAATGGAGTTTGTTGTCACAGTTGGACAGGATAAGAAGATAAACACATTTGATGTTGCGT
CCGGAAAGATGGTGAGATCTTTCAAGCAAGACAAGGATTTTGGAGACCCTATTAAGGTTATTATGGATCCAAGCTGCAGCTATCTACTTTGCTCTTACTC
CAACAAGAGTATCTGCATGTATGACTCGATGAGCGGGGATCTGGTCATGCAGGCCATGGGACATGGTGAAGTTGTAACTGGTTTCGTCTTTTTACCCGAC
TGCAAGCATATTGTATCTGTCTCTGGTGATGGTTGCATTTTCGTGTGGAAACTACCTCCTCGTATTACCTGCAGAATACTGCAGACAATAAAAGAAAACT
TTGTTCCATTGTCTCCAGAAGAGTTGGGGCTACCGGCAACGTTTACTACACGAGTTGTGATTCCTGAAGAAGAAAGCCTGCAATTCAGAGTCAATCCTGA
AGATGCACAAGAAAGGACCCCAACCTTCAGATTTAGCATTTCGCGACTTCCGAAGTGGGCGCAATCTAAACTTGCAGACTCTAGTAGTCCTCGGATGAAT
CTTGATTGCTCGCCAAATAAGGACCAACAACATGTTGAATTGAAGATCGTTTCCCCTCTGACTCATAATGTTGAAGAATCTGAGCATATGTTCATTGAAG
TCCAAACACCTTCCAGACTAGATGCATGTGGCAAAGAGTCCTCCTTTAATAGCTCAGATTCGGATTCTGCTCGAACAAGTGAACGTGAAAGAAAGTCAAT
GACACGGAAGAATCTCAGTCGTTTTGCATTGGATAAACGTTGGCTCAGTGTTTATACCGTGTGTCACATCCTGAATTCCCCCGAAGTGCAGCGCTCAATT
GATTTAAAGATGCCAGTCCCATCAAGTTCTAGTAATTGTGATATTTTGTCTGAAGATAACTGTCGGTTTTCTGGTGAAATAACAAATGCAAGTGATCAGC
CGAATGGTAGCCAGATATCTGAGCAGTGTTCGTTCAATTTGGAGACCCAAACTCAAGAGGGCATAGCAACTGAAGAGAGTGAACTTTTCAAGCAGCACTT
TGGTAGTTTATCAACAGCGAGCAAGATAGAAGGCTGTAAATCATCAGTAAGAAGAAGTTACTCTGCAAGATATGTTGTGCGGCGGGAGTATCCTGGCGTT
CCCCAGAAACTCTTCACCACACCCACCCGTGATTCTGGTCGTAAACCATCAGATGCAAAGGTTACTGGAGAATCATCCGTTCCTATCGACATGGAGAAAC
CAGCAATACAGTATCTGGGAGAAGGTCAACATATAATTTCTTCCATGCAGGTCCTATCACACACCTCTCAAAAAGATTTAACAAAATGCATCACAGAGGA
AGAGCCTTACGAAGGAGAACAAGACCGAAAGGAACATACTTTGGAGGAAAATGACCTGCAGTCTACTGAGGAAATAGTATCACTGTGTAGAGAAGCATTA
TTGAAACTGAATGCTGCAGCTGACAACGCATCTCTTATGTTCTCACAATTAGAAAGCGTGGTCTCGAGGCAAGAAACTTCAAATAGACCCGGAGCTAAGT
TTTATGAAAGGGCAAGCGACCTGCTCTCTTCGATCACAGAGAAAATTAATGCAGTTGCAAAACTGGCGCAGCGTGGCAATGGTGGAAGCACAAGTAAAAT
GGAAGTTCATTGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10017180 pacid=23165832 polypeptide=Lus10017180 locus=Lus10017180.g ID=Lus10017180.BGIv1.0 annot-version=v1.0
MKANRRAKKPDASSKLVLEEIIGLTTKNANGIASTESTSACAYIAGCAVVLHDVDSGRQSHLIASHRMPKPLSCVAMSRDGRLVAAGESGNQPAVLVWDR
STLSFLSDLNGHVYGVECIAFSPDGEHLVSVGGYIYLWDWKNGVLVAKLKASSSCSAITSVSFTSNGKFMVTAGKKHLKFWTVGSTSKNTRLNAAIGALI
IQGKPVNLGPHKGSSFVCVASSANVEPVGTMFPIYALTDGGLLCVIDKGLSVAKSADLKVDKGLALSASRKLIACGCSNGIVKLFSNETLNYTGTLLHSK
AKICDESIDNLRNTEDSGENPVPASPDSVACQFSASEKLVVVYADHTLCIWDVLDVNQASRCCMLVSHSACIWDVKNLCCEYMHDPSLVCAARGCAGGVS
FATCSADGTIRLWDLAPQQQFVGDDSDHHGLNAEPLSPSCLASTGIFERDTLDVGLSSHGFRCMAISSDAQYLAAGDNEGNLHIYNLRTFDYTCFQDVHH
SEILSLSFSASQEEHDAPEVDHGNYLLVSGGRDRVIHLYDVKRNFNLIASVDEHSAAVTSVKLDRVGCKAISCSADRSLVFRDVFVSDSGCQIFRRHHQM
ASLGTVYDMAVDKKMEFVVTVGQDKKINTFDVASGKMVRSFKQDKDFGDPIKVIMDPSCSYLLCSYSNKSICMYDSMSGDLVMQAMGHGEVVTGFVFLPD
CKHIVSVSGDGCIFVWKLPPRITCRILQTIKENFVPLSPEELGLPATFTTRVVIPEEESLQFRVNPEDAQERTPTFRFSISRLPKWAQSKLADSSSPRMN
LDCSPNKDQQHVELKIVSPLTHNVEESEHMFIEVQTPSRLDACGKESSFNSSDSDSARTSERERKSMTRKNLSRFALDKRWLSVYTVCHILNSPEVQRSI
DLKMPVPSSSSNCDILSEDNCRFSGEITNASDQPNGSQISEQCSFNLETQTQEGIATEESELFKQHFGSLSTASKIEGCKSSVRRSYSARYVVRREYPGV
PQKLFTTPTRDSGRKPSDAKVTGESSVPIDMEKPAIQYLGEGQHIISSMQVLSHTSQKDLTKCITEEEPYEGEQDRKEHTLEENDLQSTEEIVSLCREAL
LKLNAAADNASLMFSQLESVVSRQETSNRPGAKFYERASDLLSSITEKINAVAKLAQRGNGGSTSKMEVHC
|
DESeq2's median of ratios [FLAX]
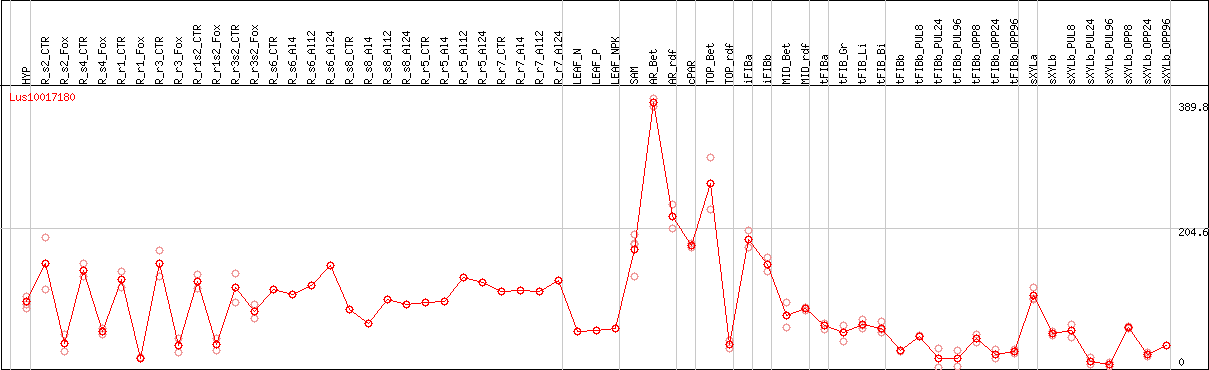
Coexpressed genes
Lus10017180 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
