External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39090 314 / 4e-108
EMB3005, RD19A, RD19
RESPONSIVE TO DEHYDRATION 19A, RESPONSIVE TO DEHYDRATION 19, Papain family cysteine protease (.1)
AT2G21430 305 / 1e-104
Papain family cysteine protease (.1)
AT4G16190 281 / 6e-95
Papain family cysteine protease (.1)
AT3G54940 250 / 8e-83
Papain family cysteine protease (.2)
AT3G19400 107 / 1e-27
Cysteine proteinases superfamily protein (.1.2)
AT5G60360 107 / 1e-27
SAG2, AALP
SENESCENCE ASSOCIATED GENE2, aleurain-like protease (.1.2.3)
AT3G19390 107 / 2e-27
Granulin repeat cysteine protease family protein (.1)
AT3G45310 105 / 4e-27
Cysteine proteinases superfamily protein (.1.2)
AT1G06260 104 / 1e-26
Cysteine proteinases superfamily protein (.1)
AT1G20850 102 / 5e-26
XCP2
xylem cysteine peptidase 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017487
370 / 2e-130
AT4G39090 550 / 0.0
RESPONSIVE TO DEHYDRATION 19A, RESPONSIVE TO DEHYDRATION 19, Papain family cysteine protease (.1)
Lus10028797
370 / 7e-130
AT4G39090 559 / 0.0
RESPONSIVE TO DEHYDRATION 19A, RESPONSIVE TO DEHYDRATION 19, Papain family cysteine protease (.1)
Lus10002103
288 / 4e-99
AT4G39090 431 / 5e-153
RESPONSIVE TO DEHYDRATION 19A, RESPONSIVE TO DEHYDRATION 19, Papain family cysteine protease (.1)
Lus10024303
289 / 6e-98
AT4G39090 520 / 0.0
RESPONSIVE TO DEHYDRATION 19A, RESPONSIVE TO DEHYDRATION 19, Papain family cysteine protease (.1)
Lus10038684
234 / 2e-76
AT3G54940 499 / 1e-177
Papain family cysteine protease (.2)
Lus10037951
206 / 5e-64
AT3G54940 451 / 6e-157
Papain family cysteine protease (.2)
Lus10020734
105 / 4e-27
AT5G45890 390 / 4e-136
senescence-associated gene 12 (.1)
Lus10034895
105 / 6e-27
AT3G45310 524 / 0.0
Cysteine proteinases superfamily protein (.1.2)
Lus10020722
103 / 2e-26
AT5G45890 392 / 1e-136
senescence-associated gene 12 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0125
Peptidase_CA
PF00112
Peptidase_C1
Papain family cysteine protease
Representative CDS sequence
>Lus10017489 pacid=23163130 polypeptide=Lus10017489 locus=Lus10017489.g ID=Lus10017489.BGIv1.0 annot-version=v1.0
ATGAGGTTTGATCTTCTTATAACCTTCCCTCTTTACATAACTTCAGCAAGAGCTATTATTAACGAATTGGTCGATTACCAGTGTGACCCAGAAGAGAAAG
GAGCATGCGACGCTGGATGTGGAGGAGGCCTAATGAACAGTGCCTTCGAGTACACTCTAAAAGCCGGAGGGCTCATGCGAGAGGAAGACTACCCTTACAC
TGGCACTGACCGCAGCACCTGCAAGTTCGACAAGAGCAAGATCGCAGCCAAAGTAGCAAACTTTAGTGTCATCTCCATTGATGAAGACCAGATTGCTGCT
AACCTCGTCCAGAGTGGCCCTCTCGCTGTGGCGATCAATGCGGTGTACATGCAGACCTACATCGGCGGGGTGTCGTGCCCTTACATCTGCTCGAAGAGGC
TAGACCACGGTGTGCTATTGGTGGGGTATGGGGAGGCAGGGTACGCACCGATCAGGATGAAGGAGAAGCCGTACTGGATCATAAAGAACTCGTGGGGGGA
GACCTGGGGGGAGAGTGGCTACTACAAGATCTGCAGAGGAAAGAACGTCTGCGGAGTTGATTCCATGGTTTCCTCTGTTGCTGCTGTTCAGACCACTGCT
CATTAG
AA sequence
>Lus10017489 pacid=23163130 polypeptide=Lus10017489 locus=Lus10017489.g ID=Lus10017489.BGIv1.0 annot-version=v1.0
MRFDLLITFPLYITSARAIINELVDYQCDPEEKGACDAGCGGGLMNSAFEYTLKAGGLMREEDYPYTGTDRSTCKFDKSKIAAKVANFSVISIDEDQIAA
NLVQSGPLAVAINAVYMQTYIGGVSCPYICSKRLDHGVLLVGYGEAGYAPIRMKEKPYWIIKNSWGETWGESGYYKICRGKNVCGVDSMVSSVAAVQTTA
H
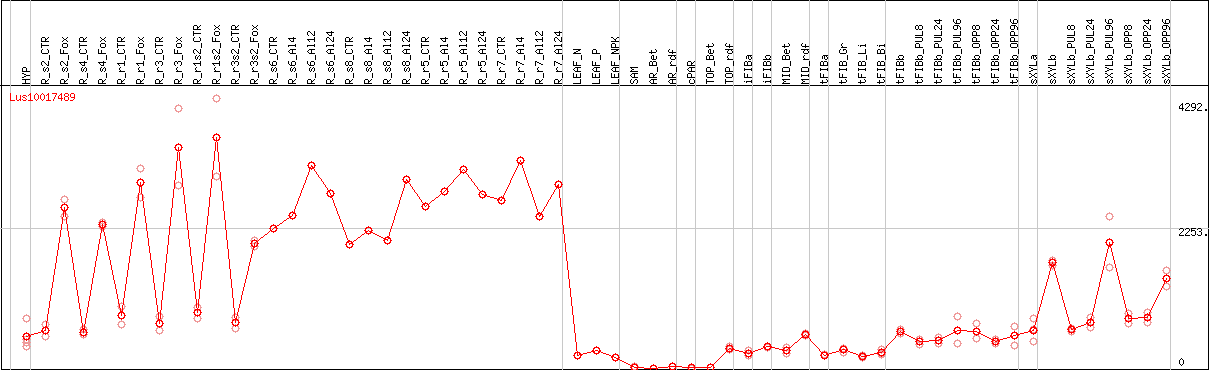
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017489 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.