External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G23690 186 / 4e-60
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT1G64160 182 / 2e-58
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT4G11190 162 / 1e-50
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT4G11210 161 / 3e-50
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT4G11180 140 / 4e-42
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT1G22900 77 / 2e-17
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT1G65870 70 / 9e-15
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT5G49040 69 / 2e-14
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT5G49030 71 / 3e-14
OVA2
ovule abortion 2, tRNA synthetase class I (I, L, M and V) family protein (.1), tRNA synthetase class I (I, L, M and V) family protein (.2), tRNA synthetase class I (I, L, M and V) family protein (.3)
AT5G42500 68 / 4e-14
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024715
251 / 2e-85
AT4G23690 186 / 2e-60
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10024714
248 / 3e-84
AT4G23690 186 / 3e-60
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10032331
246 / 1e-83
AT4G23690 187 / 5e-61
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10017539
176 / 6e-56
AT1G64160 238 / 9e-81
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10028749
174 / 3e-55
AT1G64160 232 / 2e-78
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10028750
94 / 6e-25
AT4G23690 52 / 3e-09
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10025064
82 / 1e-19
AT2G21100 197 / 9e-65
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10034479
81 / 3e-19
AT2G21100 189 / 9e-62
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10034478
79 / 6e-18
AT2G21100 191 / 1e-61
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G142702
303 / 4e-106
AT4G23690 191 / 2e-62
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.001G097001
303 / 4e-106
AT4G23690 191 / 2e-62
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.013G142602
262 / 4e-90
AT1G64160 181 / 3e-58
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.001G096800
262 / 4e-90
AT1G64160 181 / 3e-58
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.003G134600
259 / 8e-89
AT1G64160 182 / 5e-59
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.013G142501
258 / 2e-88
AT1G64160 179 / 1e-57
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.001G096680
258 / 2e-88
AT1G64160 178 / 2e-57
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.003G134400
252 / 3e-86
AT1G64160 184 / 1e-59
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.003G134800
241 / 1e-81
AT1G64160 188 / 4e-61
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.001G096440
182 / 4e-59
AT1G64160 119 / 1e-34
Disease resistance-responsive (dirigent-like protein) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0650
AOC_barrel
PF03018
Dirigent
Dirigent-like protein
Representative CDS sequence
>Lus10017538 pacid=23163131 polypeptide=Lus10017538 locus=Lus10017538.g ID=Lus10017538.BGIv1.0 annot-version=v1.0
ATGGACACCAAACCATCATCAACAACCCTGTTAGCTCTCTCATCATTCCTCCTCCTTTTCTTCCTCGCCAATTCTCAACCATCTCACGATCAACATCATA
ACTACCCTCGAAAGCCACGACCGCAACGTCGTGCCATACCAAAGCCATGCAAGGAGCTAGTCTTCTACTTCCACGACATCATCTACAACGGCAAGAACTC
GAGGAACGCCACCGCGGCCATCGTGGGGGCCCCTGCCTGGGGGAACAGGACCATCCTGGCCGGGCAGAACCACTTTGGTGACGTGGTCGTGTTCGACGAC
CCGATAACGTTGGACAACAACTTGCACGCCCCGGCGGTGGGGCGCGCACAAGGGATGTATGTTTACGACAAGAAGGAAGTGTTCACGGCGTGGTTGGGGT
TCTCGTTCGTGTTTAACTCGACGGAGCATGTGGGGACTCTGACGTTTGCCGGAGCTGATCCGCTGATGAACAAGACGAGGGATGTGTCGGTCGTCGGAGG
CACCGGTGATTTCTTCATGGCTAGAGGGATTGCTACGTTGATGACGGATGCGTTCGAAGGTGAAGTTTATTTTAGGCTGAGGGTTGATATTAAGCTTTAT
GAATGTTGGTGGTGA
AA sequence
>Lus10017538 pacid=23163131 polypeptide=Lus10017538 locus=Lus10017538.g ID=Lus10017538.BGIv1.0 annot-version=v1.0
MDTKPSSTTLLALSSFLLLFFLANSQPSHDQHHNYPRKPRPQRRAIPKPCKELVFYFHDIIYNGKNSRNATAAIVGAPAWGNRTILAGQNHFGDVVVFDD
PITLDNNLHAPAVGRAQGMYVYDKKEVFTAWLGFSFVFNSTEHVGTLTFAGADPLMNKTRDVSVVGGTGDFFMARGIATLMTDAFEGEVYFRLRVDIKLY
ECWW
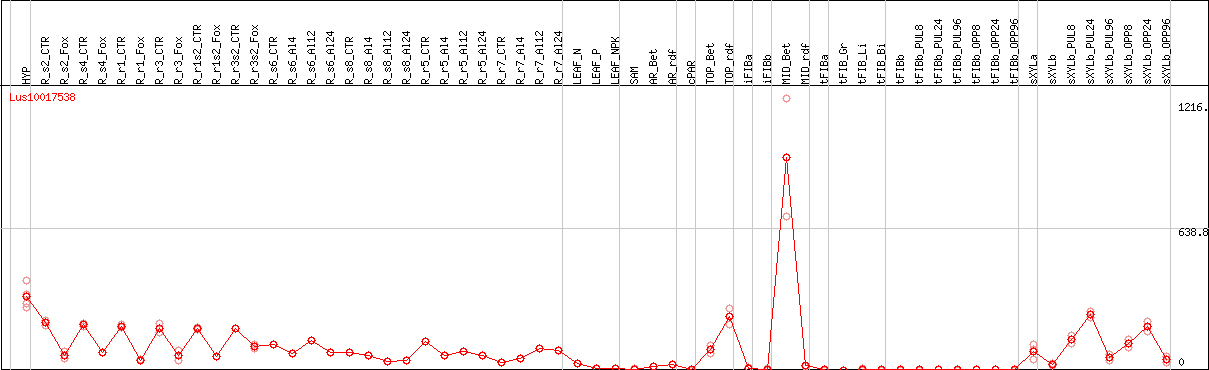
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017538 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.