External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G60920 300 / 4e-101
COB
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
AT3G02210 288 / 2e-96
COBL1
COBRA-like protein 1 precursor (.1)
AT3G29810 281 / 8e-94
COBL2
COBRA-like protein 2 precursor (.1)
AT5G15630 231 / 2e-74
IRX6, COBL4
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
AT1G09790 214 / 1e-67
COBL6
COBRA-like protein 6 precursor (.1)
AT5G49270 59 / 3e-10
DER9, MRH4, SHV2, COBL9
SHAVEN 2, MUTANT ROOT HAIR 4, DEFORMED ROOT HAIRS 9, COBRA-LIKE 9, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
AT4G16120 57 / 3e-09
ATSEB1, COBL7
ARABIDOPSIS THALIANA SEC61 BETA 1, COBRA-like protein-7 precursor (.1)
AT3G20580 53 / 3e-08
COBL10
COBRA-like protein 10 precursor (.1)
AT3G16860 49 / 6e-07
COBL8
COBRA-like protein 8 precursor (.1)
AT4G27110 48 / 2e-06
COBL11
COBRA-like protein 11 precursor (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034671
382 / 3e-131
AT5G60920 732 / 0.0
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Lus10007986
327 / 1e-111
AT5G60920 755 / 0.0
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Lus10021144
325 / 6e-111
AT5G60920 749 / 0.0
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Lus10012598
303 / 2e-102
AT3G02210 697 / 0.0
COBRA-like protein 1 precursor (.1)
Lus10034379
287 / 3e-96
AT3G02210 685 / 0.0
COBRA-like protein 1 precursor (.1)
Lus10017863
236 / 5e-76
AT5G15630 717 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Lus10034670
235 / 1e-75
AT5G15630 715 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Lus10034666
234 / 1e-75
AT5G15630 627 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Lus10017866
231 / 2e-74
AT5G15630 625 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G060000
320 / 3e-109
AT5G60920 759 / 0.0
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.004G117100
295 / 4e-99
AT3G02210 724 / 0.0
COBRA-like protein 1 precursor (.1)
Potri.015G060100
237 / 1e-76
AT5G15630 739 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.015G060200
230 / 4e-74
AT5G15630 669 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.004G117200
230 / 4e-74
AT5G15630 679 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.017G098600
225 / 2e-72
AT5G60920 601 / 0.0
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.017G098700
224 / 8e-72
AT5G60920 614 / 0.0
COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.004G219200
212 / 3e-67
AT1G09790 511 / 2e-180
COBRA-like protein 6 precursor (.1)
Potri.012G062200
207 / 3e-65
AT5G15630 586 / 0.0
IRREGULAR XYLEM 6, COBRA-LIKE4, COBRA-like extracellular glycosyl-phosphatidyl inositol-anchored protein family (.1)
Potri.004G167100
169 / 1e-50
AT3G29810 407 / 2e-139
COBRA-like protein 2 precursor (.1)
PFAM info
Representative CDS sequence
>Lus10017861 pacid=23148846 polypeptide=Lus10017861 locus=Lus10017861.g ID=Lus10017861.BGIv1.0 annot-version=v1.0
ATGTCTCAGCGAGTGTTGGGGTTAGAGGAGGGAAACCTAGCCGACTTACGCGGTCCTGACACTCCGCATTTAGCCTCGGTGGTTTCAGGTCAAGGAAAAC
AAGATCTCGCACCTTTGGTGCAGTGTACCAAGCATATGTGTCCGATCCGGATCCATTGGCATGTGAAGCTTAACTACAAGGAGTACTGGAGAGTGAAAGT
GACCATCACAAATTTCGATTACAGAATGAATTACTACACACAGTGGAATCTAGTGGTGCAACACCCCAACTTTGCCAACCTTACCCAGCTTTTCAGCTTC
AACTATAAGTCCTTAACTCCTTATGATGGTTTCAATGATACATCTATGTTATGGGGAGTCAAGTTCTACAACGATTTTCTTTCGCAAGCTGGGCCCTTGG
GGAATGTCCAGTCCGAGCTTCTATTCCGGAAAGATTCATCAACTTTCACATTCGAGAAAGGATGGGCTTTCCCCAGGAGGATATACTTTAATGGTGACAA
CTGTGTGATGCCTCCGCCAGATGCATATCCATGGTCGCCTAATGATGGTCACAGATCATCAACCATGTCTCTACTTCGACCTCTAGGCGTAGCATTGTTG
TCTATCATGATCCTCGTAGCTTATGCGTGA
AA sequence
>Lus10017861 pacid=23148846 polypeptide=Lus10017861 locus=Lus10017861.g ID=Lus10017861.BGIv1.0 annot-version=v1.0
MSQRVLGLEEGNLADLRGPDTPHLASVVSGQGKQDLAPLVQCTKHMCPIRIHWHVKLNYKEYWRVKVTITNFDYRMNYYTQWNLVVQHPNFANLTQLFSF
NYKSLTPYDGFNDTSMLWGVKFYNDFLSQAGPLGNVQSELLFRKDSSTFTFEKGWAFPRRIYFNGDNCVMPPPDAYPWSPNDGHRSSTMSLLRPLGVALL
SIMILVAYA
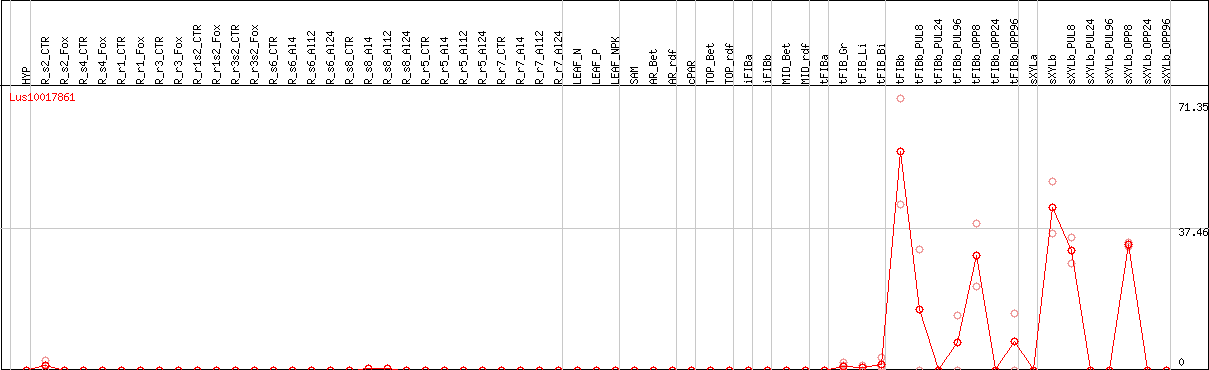
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017861 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.