External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G08100 152 / 9e-46
ASPGA1
asparaginase A1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
AT3G16150 108 / 7e-29
ASPGB1
asparaginase B1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
AT4G00590 42 / 5e-05
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034668
174 / 2e-54
AT5G08100 489 / 6e-176
asparaginase A1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
Lus10034655
172 / 9e-54
AT5G08100 507 / 0.0
asparaginase A1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
Lus10024795
111 / 5e-30
AT3G16150 536 / 0.0
asparaginase B1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Lus10040935
45 / 9e-06
AT4G00590 484 / 8e-171
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
Lus10009826
43 / 4e-05
AT4G00590 446 / 2e-156
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G063400
159 / 1e-48
AT5G08100 485 / 2e-174
asparaginase A1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
Potri.014G022900
112 / 1e-30
AT3G16150 556 / 0.0
asparaginase B1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Potri.002G122900
110 / 9e-30
AT3G16150 540 / 0.0
asparaginase B1, N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Potri.014G080600
45 / 9e-06
AT4G00590 486 / 1e-171
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF01112
Asparaginase_2
Asparaginase
Representative CDS sequence
>Lus10017879 pacid=23148861 polypeptide=Lus10017879 locus=Lus10017879.g ID=Lus10017879.BGIv1.0 annot-version=v1.0
ATGGGTTGGGCAATCGCTCTCCACGGCGGCGCCGGCGACATACCACGCTCTCTTCTGCCCGACCGCTGTCTCCCCCGCGAAGCCGCTCCCTCCAGCTCGG
TGTCTCCGCTCTTCAGTCCAACTCCCACCCTCTTGACGTCGTCGAGCTCGTCTTGGTTGTTCTCTCGTTTTGTCCTTAACTTTGGTGGTCTATGTACTGT
TGGTGAATTGGAGAACCACCCACAGTTTAATGCTGGTAAGGGATCTGTGTTAACCGCTGTTGGGACAGTGGAGATGGAAGCCATCATGGACGGCAAAACA
AAGAAATGTGGAGCAGTTTCTGGACTTACCACTGTTGTCAACGCTATCTCACTTGCACGCCTCGTCATGGAAAAGACTCCTCACATCTATCCAGGATTCG
AAGGAGCTGAAGCTTTTGCAAGGGAACAAGTCAGTTTATTCTTAGCTCCCTTTCTTCTGTGGAGCTCTGAAGTCTGA
AA sequence
>Lus10017879 pacid=23148861 polypeptide=Lus10017879 locus=Lus10017879.g ID=Lus10017879.BGIv1.0 annot-version=v1.0
MGWAIALHGGAGDIPRSLLPDRCLPREAAPSSSVSPLFSPTPTLLTSSSSSWLFSRFVLNFGGLCTVGELENHPQFNAGKGSVLTAVGTVEMEAIMDGKT
KKCGAVSGLTTVVNAISLARLVMEKTPHIYPGFEGAEAFAREQVSLFLAPFLLWSSEV
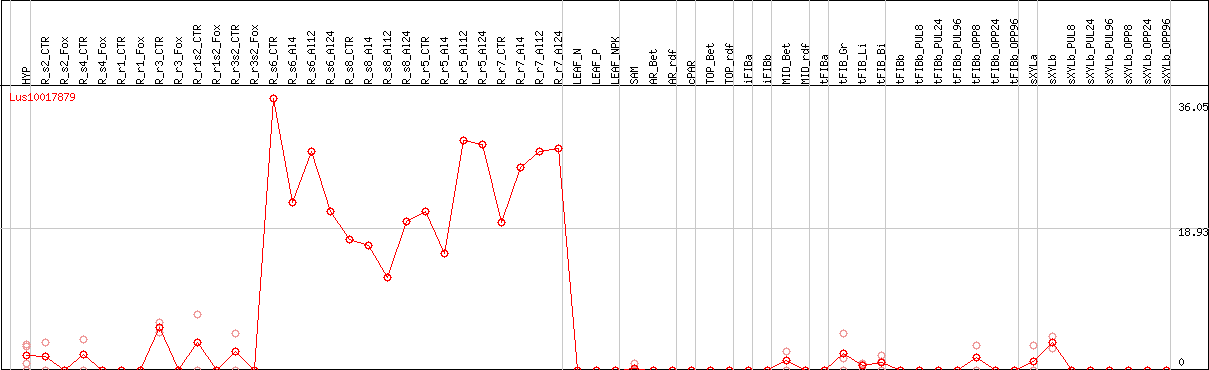
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017879 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.