External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39050 75 / 2e-15
Kinesin motor family protein (.1)
AT1G68710 75 / 2e-15
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
AT1G26130 74 / 4e-15
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1.2)
AT1G13210 72 / 2e-14
ACA.L
autoinhibited Ca2+/ATPase II, autoinhibited Ca2+/ATPase II (.1)
AT3G25610 72 / 3e-14
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
AT3G27870 70 / 1e-13
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
AT1G17500 66 / 4e-12
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
AT1G72700 66 / 4e-12
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
AT3G13900 66 / 4e-12
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
AT1G54280 66 / 4e-12
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017969
195 / 8e-57
AT4G39050 1517 / 0.0
Kinesin motor family protein (.1)
Lus10017496
140 / 7e-38
AT4G39050 1235 / 0.0
Kinesin motor family protein (.1)
Lus10028790
135 / 6e-36
AT4G39050 1242 / 0.0
Kinesin motor family protein (.1)
Lus10041958
120 / 7e-31
AT4G39050 1202 / 0.0
Kinesin motor family protein (.1)
Lus10017906
89 / 6e-22
AT4G39050 105 / 7e-42
Kinesin motor family protein (.1)
Lus10035068
77 / 7e-16
AT1G68710 1677 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Lus10034404
74 / 5e-15
AT3G25610 969 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Lus10019152
74 / 6e-15
AT3G25610 1871 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Lus10008776
68 / 8e-13
AT3G27870 1540 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G161100
101 / 2e-24
AT4G39050 1411 / 0.0
Kinesin motor family protein (.1)
Potri.009G122100
100 / 8e-24
AT4G39050 1379 / 0.0
Kinesin motor family protein (.1)
Potri.012G058000
74 / 7e-15
AT1G68710 1764 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.010G132700
72 / 2e-14
AT1G68710 1753 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.008G111200
72 / 2e-14
AT1G68710 1767 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.001G349100
66 / 4e-12
AT3G27870 1687 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.001G081200
65 / 6e-12
AT1G17500 1461 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.003G043300
64 / 1e-11
AT1G17500 1891 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.001G197500
64 / 1e-11
AT1G17500 1902 / 0.0
ATPase E1-E2 type family protein / haloacid dehalogenase-like hydrolase family protein (.1)
Potri.006G109200
57 / 3e-09
AT5G04930 1321 / 0.0
aminophospholipid ATPase 1 (.1)
PFAM info
Representative CDS sequence
>Lus10017893 pacid=23148797 polypeptide=Lus10017893 locus=Lus10017893.g ID=Lus10017893.BGIv1.0 annot-version=v1.0
ATGGTTAAAGAAGGCATTGAAGATTGGCGTTGTAAACAACAGGCTGTCATGTCAAGTGACGTTGCAATTGCTCAATTTCGATACCTGGAGCGTTTGCTAC
TTGTACATGGACATTGGTGTTACAGAAGAATATCAGCAATGGTAATGCTAGATTTGCTACTTCTTTTACAAGAATGTTACATTTGGAATCTCGAGTCGTT
GCAATCTCGATCTGCCGTTGCACAAAATGGTAGCAGCTGCAAATATAATGACGGAATAAGATCCAGGAGGAAAGGCCGGTTCTCTGGCAGGGGAAACGAA
ATGCAAATGCACTCGGATGACTTTGAGTCGTGGGATCTTGATCCTAATGATATGAAGATGGAATTGCAGGCAAGGAAACAGCGAGAGGCAGCACTTGAGG
CTGCCTTGGCTGAAAAGGAACTCATAGAAGATGAGTACAGGAAGCGTGCTGAAGAAGCAAAACGAAGGGAAGAATCGTTGGAGAATGACTTGGCAAACAT
GTGGGTGCTTGTTGCTAAGTTGAAGAAGGAAGGAGGAAATGTCACCAAGTTTAACACCCACGAAAGGACCAGTACTGGAGCTGGGTATCACATGAGCGGA
CCAGAATTGGACGGGTTCGAGCATGGTTCTGTCTTGAGAGAAGACAAGTCAGTAGCGATGATTCGAAGCGAGCCGATGATATGCAAAAGGAAGAGCCGTT
GGTTGTTCGTTTGA
AA sequence
>Lus10017893 pacid=23148797 polypeptide=Lus10017893 locus=Lus10017893.g ID=Lus10017893.BGIv1.0 annot-version=v1.0
MVKEGIEDWRCKQQAVMSSDVAIAQFRYLERLLLVHGHWCYRRISAMVMLDLLLLLQECYIWNLESLQSRSAVAQNGSSCKYNDGIRSRRKGRFSGRGNE
MQMHSDDFESWDLDPNDMKMELQARKQREAALEAALAEKELIEDEYRKRAEEAKRREESLENDLANMWVLVAKLKKEGGNVTKFNTHERTSTGAGYHMSG
PELDGFEHGSVLREDKSVAMIRSEPMICKRKSRWLFV
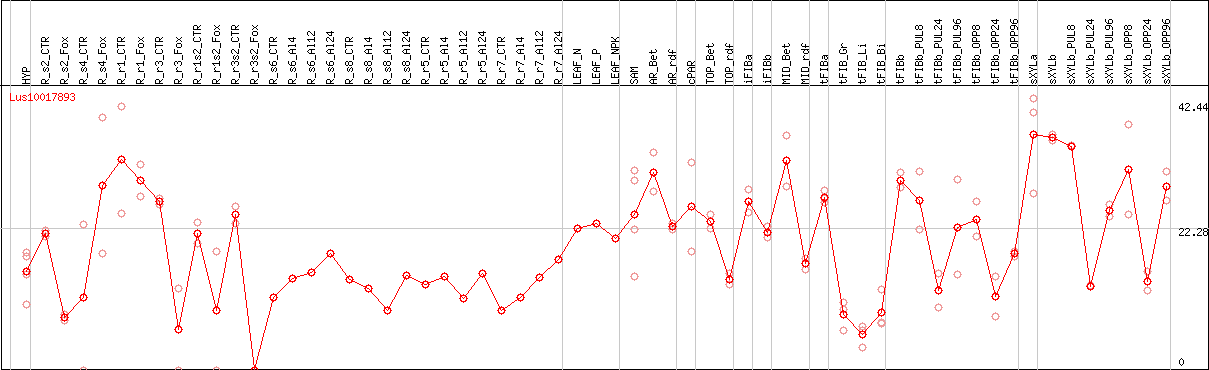
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017893 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.