External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G06450 47 / 9e-07
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G27820 44 / 1e-05
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G27890 43 / 2e-05
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G61470 42 / 5e-05
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT2G32070 40 / 0.0002
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G15920 40 / 0.0002
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2)
AT1G80780 40 / 0.0002
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017905
255 / 2e-86
AT1G06450 162 / 5e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10035077
245 / 2e-82
AT1G61470 162 / 6e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10000561
228 / 1e-75
AT1G61470 143 / 1e-40
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10001651
228 / 1e-74
AT1G61470 167 / 4e-49
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10008043
202 / 1e-65
AT1G06450 165 / 5e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10038112
197 / 2e-65
AT1G06450 52 / 4e-08
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10035078
120 / 1e-33
AT1G06450 168 / 2e-49
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10000327
110 / 6e-30
AT1G06450 174 / 1e-51
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10007771
74 / 4e-16
AT1G61470 170 / 5e-51
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G038700
48 / 4e-07
AT1G61470 182 / 8e-56
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G187200
44 / 8e-06
AT2G32070 403 / 2e-143
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.003G181100
42 / 6e-05
AT1G80780 468 / 1e-168
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2.3)
Potri.001G046700
40 / 0.0002
AT2G32070 472 / 2e-170
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G262500
39 / 0.0006
AT2G32070 443 / 5e-159
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.018G020900
38 / 0.001
AT2G32070 441 / 2e-158
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10017894 pacid=23148840 polypeptide=Lus10017894 locus=Lus10017894.g ID=Lus10017894.BGIv1.0 annot-version=v1.0
ATGGCTAAGTTCTATGAAGACACATTGAGTGAAATTGGGTTGCAGAAGTTAGCGGATGCTTTGGGGGTTAAGAGAAGAGGGGAAGCTCATACTGCCGCCT
CTGATAGTTTCTTGACGGCGCTGGTTTACTTTGAGTTGAAGAAGAAACTGATGCTGTTGGGTGTTGATGAAGAGTCTTACGTTGATTACGTCTATGGAAT
CAGCACCAAAATCTCCAGGAAGCAGAAAGAGCCCCTCTTTCTACCAACTGCTCGACCAGCAGGTCCTCCTGTGTATCATCATCATCCTTATCATCCTCAT
CTTTATCAGAGTAAGGTGCTGTATCCCATTCCATATCAGCAGAGAGTAGTACCCTTGCCTCCAAATGTGATGATGGGTCGTCGTTGGGTTGATTGCTCAT
TAGCTCCAACAGTATTAGTTCCTTCTTTCATCTACTGA
AA sequence
>Lus10017894 pacid=23148840 polypeptide=Lus10017894 locus=Lus10017894.g ID=Lus10017894.BGIv1.0 annot-version=v1.0
MAKFYEDTLSEIGLQKLADALGVKRRGEAHTAASDSFLTALVYFELKKKLMLLGVDEESYVDYVYGISTKISRKQKEPLFLPTARPAGPPVYHHHPYHPH
LYQSKVLYPIPYQQRVVPLPPNVMMGRRWVDCSLAPTVLVPSFIY
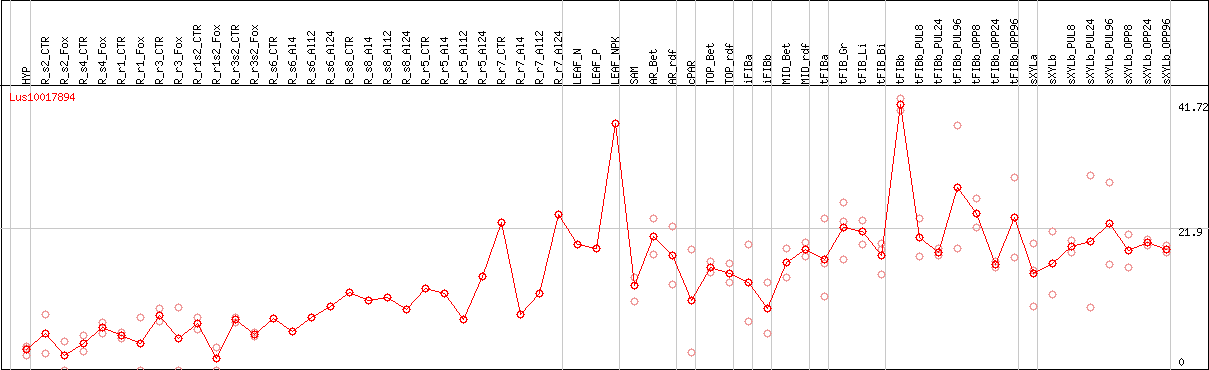
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10017894 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.