Lus10018203 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10018203 pacid=23179325 polypeptide=Lus10018203 locus=Lus10018203.g ID=Lus10018203.BGIv1.0 annot-version=v1.0
ATGAGCTCTTCCTTCCTCTCGTTCCGTCCACCACTTCCATCATCTTCTCCTACCGCATCATCACAACACCGTAGAATCCTCAATTCCCAGCGTCTAATTT
CCTCCCCACTTGAGAATTTGCCGTCATGTTCTTCGCCAAGGAGTTCGATTCTACAATACAAGCTGCGCGGATTGCCCTCCTTGCAGCTCCGTTCAGTCTC
ATCTTCTTCTTCTTCTTCCTCGCGACTCAAAGCTTCGAAAGATGATGCTGAATTCGACTACTTACCGGACGGCAGTATCGAGCCGCTAAAGCCAAGTTTC
AGAGAGTTCATTACCTCGGAGAGAGTGAAAGTGGTGGCGATGCTTGCCCTGGCACTGGCGCTTTGTAATGCCGATCGAGTTGTGATGTCCGTGGCGATTG
TGCCGCTCTCGCTTTCTAACGGATGGAGCCGATCTTTCTCTGGTATCGTTCAGTCGTCTTTCTTGTGGGGATACTTAATTTCGCCGATACCTGGAGGAAC
CTTAGTGGATTACTACGGCGGCAAGGTGGTTATGGGATGGGGTGTAGCTTTATGGTCATTAGCTACATTTCTCACACCATGGGCGGCAGAAACCTCTTTA
TTTGCTCTCCTAGCAATGCGTGCTATGCTTGGCATTGCAGAAGGTGTAGCGCTTCCTTGCATGAATAATATGGTAGTAAGATGGTTCCCGCAAACAGAAC
GATCCAGGGCTGTTGGAATCGCCATGGCTGGATTTCAGCTTGGTTGTGCCATTGGGCTCACACTTTCCCCAATTCTCATGTCCAGAGGGGGTACATTCGG
GCCATTTGTTATATTCGGACTATCTGGATTTCTTTGGGTGTTAGTTTGGTATGCTGCAATATCAAGTTCTCCTGAGAGAAGTTCACAGATTTCAAATTTT
GAGTTGGGCTACATACAACGAGGAAGCAAAAAGCAGAAGTCCACTCCTGCTAAGGAGCGAACCAAGACGAGCAGGATGCTGCCTCCTTTTGGAAGGTTGC
TGTCTAAAATGCCAACGTGGTCATTGATTGTAGCCAATGCCATGCATAGCTGGGGGTTTTTTGTCATTCTTTCTTGGATGCCCATCTATTTTAATTATAT
ATATCACGTTGATCTCAAACATGCAGCTTGGTTTAGTGCTGTACCCTGGTCTGTGATGGCTTTCATGGGCTACTTTGGGGGCTTGTGGTCGGACGTGTTG
ATTCGGGCTGGTGTTACTGTTACTTTGACTCGGAAGATTATGCAGTCAATTGGCTTCCTGGGACCAGCGGTTGCTCTTGTTGGGTTAACGACTGCAAAAA
GTCCATCCATTGCTTCTGCTTGGCTAACTTTAGCTGTTGCGCTGAAATCTTTCAGCCACTCTGGCTTTCTTGTGAATCTTCAGGAAATTGCACCACACTA
CTCAGGCGTCTTACATGGCATGTCGAACACTGCAGGCACATTAGCAGCCATTTTAGGAACAGTTGGAGCTGGCTATTTTGTGGATCTGACTGGTTCTTTC
AGAGGATTCCTGCTACTGACATCGGCTTTATACTTCCTCTCTGCCATTTTCTATATCATCTTCTCAACTGGGGAGAGAGTGGATTTTGATGAAACGGCTC
AAAAAGTAAGCTCTTCCATGTTCATCTCAACTATGAAGAAAGGCAGCTCCCGGAAGAAACGAAACAATCATCCTCTTGACAAATTCAACGCCAAAGATTC
TCCTTCTCCGTTGATCAAATTCATGAAAGCTGCTGCCGGAAAAGTAGCTTCAGCCTTCACAATTTTCCGGCGGAGTAATCCTCCCGCCGCCGACGTCAAT
CTCGCCAGCAAGAGCATTAGCTCTGCTGCTGGATCGTACTTTTCAAGCGAGCTGTCAAGGGTAAGCAACAGCAGCAAGAGCACGTCGTCGAGGTTGAAGT
CTTCGAGCTCATACGGATCGTCGTCCACAAGCGGCGGAGGAGTAGTGTTCGGATCGACACAGCAGGGATACTCGCTTCAAGAGATTGTTAAGGCGACCGA
TAATTTCTCGTCGGGTACCAAGATTGGAGAAGGTAGTTTTGGGACTGTGTACAAGGGAAAGTTGAAAGATGGGTCTTTTATTGCAGTGAAGCGTGCCAGA
AAGAATTCACATGACAAGCGTTTGGCATTGGAGTTCAGGAACGAGATACTTACTCTGTCTAAGATTGAGCATTTGAATCTGGTGAGATTGTACGGCTATG
TCGAGCATCAAGACGAGAGGCTTATCGTGGTCGAATATGTCAACAATGGGAACCTCAGAGACCATCTTGACAATAGAAGAGGAACTATACTCGAATTCGC
GGAGCGTTTAGACATTGCCATTGATGTTGCACACGCTATTACCTATCTTCACATGTACACAGACCGAGCGATAATCCACAGAGACATAAAAGCATCAAAC
ATCCTTCTAAACGACAAGCTTCGAGCCAAAGTTGCAGACTTTGGTTTCGCAAGATTAGCCGCTGAAGATTGCAATGACACCCACATCTCGACTCAGATCA
AAGGCACAACTGGTTACTTAGACCCTGAATACCTCAGGACATATACACTCACCGAGAAGAGCGATGTCTTCTCCTTCGGTGTCCTACTCGTTGAACTCGT
TACTGGAAGGCACCCGATCGACCAGAATCGATCTATCAACGAGAAAGTTACCATCAAATGGGCAATGAAGAGACTCAAAGACGGAGATGCGATAATGGCG
ATGGATCCGATCCTGAGAAGGAGCCCTGCATCGAACAGAGCGACCGAGGATGTTCTGAGACTGGCTCAGAAGTGTCTGGCACCGATGAGACAATTGCGAC
CTTCCATGAAGGAATGCGCAGAGGTTCTGTGGAGAATACGAAAGGAGTTTCGAGAAAAGGGCGCCCTTCAGTCTTCCTCTGCGGCTTCAGCTTCTGCTGA
TTTTCCCAGGAGGAATCCTAATCAGACCCGGATTACTACATTCGGGTTGGAAGATATTGGTGAGAACCCCAGGTTCGTTTCCGCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10018203 pacid=23179325 polypeptide=Lus10018203 locus=Lus10018203.g ID=Lus10018203.BGIv1.0 annot-version=v1.0
MSSSFLSFRPPLPSSSPTASSQHRRILNSQRLISSPLENLPSCSSPRSSILQYKLRGLPSLQLRSVSSSSSSSSRLKASKDDAEFDYLPDGSIEPLKPSF
REFITSERVKVVAMLALALALCNADRVVMSVAIVPLSLSNGWSRSFSGIVQSSFLWGYLISPIPGGTLVDYYGGKVVMGWGVALWSLATFLTPWAAETSL
FALLAMRAMLGIAEGVALPCMNNMVVRWFPQTERSRAVGIAMAGFQLGCAIGLTLSPILMSRGGTFGPFVIFGLSGFLWVLVWYAAISSSPERSSQISNF
ELGYIQRGSKKQKSTPAKERTKTSRMLPPFGRLLSKMPTWSLIVANAMHSWGFFVILSWMPIYFNYIYHVDLKHAAWFSAVPWSVMAFMGYFGGLWSDVL
IRAGVTVTLTRKIMQSIGFLGPAVALVGLTTAKSPSIASAWLTLAVALKSFSHSGFLVNLQEIAPHYSGVLHGMSNTAGTLAAILGTVGAGYFVDLTGSF
RGFLLLTSALYFLSAIFYIIFSTGERVDFDETAQKVSSSMFISTMKKGSSRKKRNNHPLDKFNAKDSPSPLIKFMKAAAGKVASAFTIFRRSNPPAADVN
LASKSISSAAGSYFSSELSRVSNSSKSTSSRLKSSSSYGSSSTSGGGVVFGSTQQGYSLQEIVKATDNFSSGTKIGEGSFGTVYKGKLKDGSFIAVKRAR
KNSHDKRLALEFRNEILTLSKIEHLNLVRLYGYVEHQDERLIVVEYVNNGNLRDHLDNRRGTILEFAERLDIAIDVAHAITYLHMYTDRAIIHRDIKASN
ILLNDKLRAKVADFGFARLAAEDCNDTHISTQIKGTTGYLDPEYLRTYTLTEKSDVFSFGVLLVELVTGRHPIDQNRSINEKVTIKWAMKRLKDGDAIMA
MDPILRRSPASNRATEDVLRLAQKCLAPMRQLRPSMKECAEVLWRIRKEFREKGALQSSSAASASADFPRRNPNQTRITTFGLEDIGENPRFVSA
|
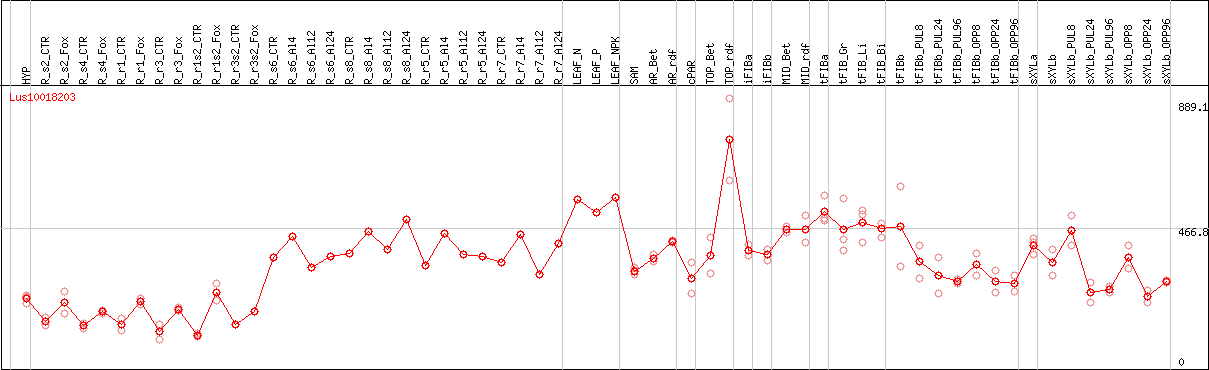
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018203 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
