External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G12770 162 / 9e-45
MEF22
mitochondrial editing factor 22 (.1)
AT4G35130 142 / 3e-37
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G05240 136 / 1e-35
MEF19
mitochondrial editing factor 19 (.1)
AT3G11460 135 / 2e-35
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G15510 136 / 3e-35
VAC1, ATECB2
VANILLA CREAM 1, ARABIDOPSIS EARLY CHLOROPLAST BIOGENESIS2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G03580 135 / 3e-35
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G33760 134 / 6e-35
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G08070 134 / 9e-35
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 128 / 6e-33
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G11290 128 / 1e-32
CRR22
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040648
564 / 0
AT4G18750 376 / 1e-119
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10024876
155 / 1e-41
AT3G12770 886 / 0.0
mitochondrial editing factor 22 (.1)
Lus10033858
150 / 1e-40
AT1G08070 422 / 4e-139
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014211
150 / 2e-40
AT3G11460 791 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10022702
145 / 3e-39
AT3G11460 641 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10037362
145 / 2e-38
AT3G57430 477 / 9e-158
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10000706
141 / 4e-37
AT3G12770 878 / 0.0
mitochondrial editing factor 22 (.1)
Lus10031028
139 / 2e-36
AT4G19191 727 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10018978
137 / 2e-35
AT4G13650 1258 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G152300
300 / 4e-96
AT4G33990 489 / 1e-162
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G215500
149 / 6e-40
AT3G63370 922 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 86, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G211400
147 / 2e-39
AT3G11460 772 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G152000
146 / 8e-39
AT3G12770 931 / 0.0
mitochondrial editing factor 22 (.1)
Potri.012G108000
145 / 1e-38
AT2G34400 640 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.012G111900
144 / 2e-38
AT4G33990 518 / 1e-173
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G071800
143 / 6e-38
AT1G08070 606 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G040100
140 / 8e-37
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G086200
139 / 2e-36
AT4G14820 840 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.009G044700
138 / 4e-36
AT2G29760 1013 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10018264 pacid=23179347 polypeptide=Lus10018264 locus=Lus10018264.g ID=Lus10018264.BGIv1.0 annot-version=v1.0
ATGAGTAACAAGTTGTTGTCTATACTCAACTCCAGAAGGCCATCACTGTATCTCTGGAATTTGAAAATACGAGACGCTGCAAATCGCCGCCTCTTCAACC
AAACTTTTCAACTCTACCGTTCCCTGTTGAGCTTCGGCATCCGCGGCGGCAGTAGCTTCACTTACCCATTACTCCTCAAATCCGCCGCCTCCTGCTCCAT
CCCCGACGCATTCCATCTCGGTAAAACGATCCATTCCCACGTTCTGCATCTAGGTTTCCAGTCGGACGTGTACGTGCAGACATCACTCCTCGACATGTAC
TCGAAATGTCGCGACTTGGTTTCTTCCCGCCAAGTGTTGGACGAAATGTCTGAAAGAAATGTTGTGTCTTGGAACTCCATGATCTCTGCCTATTGTAGTT
GCTCTCTGTTTGACCAAGCCATTTCTTTGCTGAACGAAATGAGGGGTCTTGGTTTGGGACCAAATTCGACAACTTTCGTCGTGCTGCTTTCTCCTTGTGG
AGGCTCTGGTTCACTTCTGCAGGGCCTTTCAGCTCAGTGCTGCGTCTTCAAGATGGGGCTTCTGGATTTAGACAATGTGCCATTAACTAACGCGGTGATG
GGAATGCTCGTTAATCACGGACAGACTCGTGAAGCTCGTGCCATTCTCGACAAGATGCACGAGCCTTCTCTAATCTCTTGGACAACACTACTAAACGGTT
ACGTAGAATCTGGCAACGTAACCGAAGCTTTCTCTGTATTCAATCAGACGAGAAGAAGAGAAAGCGTCGATTTCGTAGCTTTCCTCACCATGATCACTTG
TTGTGGCCAACACGGTTACCTGTCGATATCCTCCTCGCTCCATTCGTTGATAATCAAAACCGGGTGTGACGAGGAGAACGCCATTGCTAATCTTCTTGTG
AGCATGTATGCGAAATGTGGTGATCTTAGCTCAGCTCGACTTGTGTTTGATTTCACAGGAGAGAAAACTATCTCGGTATGGACATCAATAATGGCTGCAC
TCTGA
AA sequence
>Lus10018264 pacid=23179347 polypeptide=Lus10018264 locus=Lus10018264.g ID=Lus10018264.BGIv1.0 annot-version=v1.0
MSNKLLSILNSRRPSLYLWNLKIRDAANRRLFNQTFQLYRSLLSFGIRGGSSFTYPLLLKSAASCSIPDAFHLGKTIHSHVLHLGFQSDVYVQTSLLDMY
SKCRDLVSSRQVLDEMSERNVVSWNSMISAYCSCSLFDQAISLLNEMRGLGLGPNSTTFVVLLSPCGGSGSLLQGLSAQCCVFKMGLLDLDNVPLTNAVM
GMLVNHGQTREARAILDKMHEPSLISWTTLLNGYVESGNVTEAFSVFNQTRRRESVDFVAFLTMITCCGQHGYLSISSSLHSLIIKTGCDEENAIANLLV
SMYAKCGDLSSARLVFDFTGEKTISVWTSIMAAL
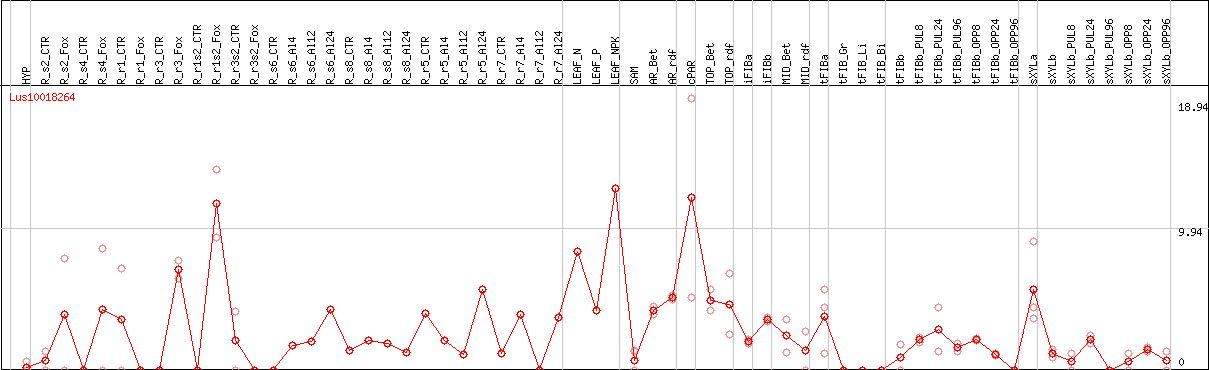
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018264 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.