External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G16860 194 / 1e-57
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G39350 192 / 1e-57
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G16835 187 / 5e-56
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G18750 189 / 6e-56
DOT4
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G13600 186 / 4e-55
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G18485 187 / 8e-55
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G24000 184 / 1e-54
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G66520 182 / 2e-54
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G03880 181 / 7e-54
REME1
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G02750 182 / 2e-53
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040648
366 / 4e-125
AT4G18750 376 / 1e-119
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10035788
206 / 6e-62
AT3G16610 636 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10037362
203 / 1e-61
AT3G57430 477 / 9e-158
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10028593
190 / 1e-56
AT5G16860 879 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10031989
188 / 7e-56
AT3G26782 865 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10018897
190 / 8e-56
AT5G16860 970 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10006628
181 / 1e-55
AT4G21065 399 / 7e-137
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10005987
183 / 2e-55
AT5G08510 602 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10030225
179 / 3e-55
AT5G08510 472 / 2e-165
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10018265 pacid=23179272 polypeptide=Lus10018265 locus=Lus10018265.g ID=Lus10018265.BGIv1.0 annot-version=v1.0
ATGTTTAACAGCATATTGGTTAAGGACTTGACAGCTTGGAGCTCCATGATTAACGGATACGCGATTCACGGGATGGGAAATGAAGCTCTCGGTTTATTCC
ACGAGATGCGAAAAGAAGGGATAGTGCCGGATAGTGTTACTTACACGAGTGTATTACTTGCGTGTAGTCATTCGGGACTCGTTGAGGATGGAGTGAATTA
TTTTCTGAGCATGCAGAAGGATTTCGGAATTGAACCTGGGATAGAACACTATACCTGCTTGGTAGATCTTCTTGGGAGAGCTGGTCGGCTCGACTTGGCG
ATGAAACCGACGTTGGAAATGCCCGGAGAGATTCAAGCGCGAATGTGGCCTTCGATTCTGAGTTCGTGCAGGAAATATCAAAATGTAGAGCTGGGCGAAT
GTGCTGCTAAGAAGCTGTTAGAACTGAGTCCTGATCAGACTGGGAATTATGTTCTGATGGCTAATCTGTACATGTCAGAAGGTAAGTGGAAGGAGGCTGC
CAAGATGAGAAGCTCGATGGATGGTCGAGGATTAGTAAAGAAACCTGCTTGGAGCCAGGTCGAGATTAATGGTTCGGTCAACGACTCCATTGCCAGAGTT
GGATGA
AA sequence
>Lus10018265 pacid=23179272 polypeptide=Lus10018265 locus=Lus10018265.g ID=Lus10018265.BGIv1.0 annot-version=v1.0
MFNSILVKDLTAWSSMINGYAIHGMGNEALGLFHEMRKEGIVPDSVTYTSVLLACSHSGLVEDGVNYFLSMQKDFGIEPGIEHYTCLVDLLGRAGRLDLA
MKPTLEMPGEIQARMWPSILSSCRKYQNVELGECAAKKLLELSPDQTGNYVLMANLYMSEGKWKEAAKMRSSMDGRGLVKKPAWSQVEINGSVNDSIARV
G
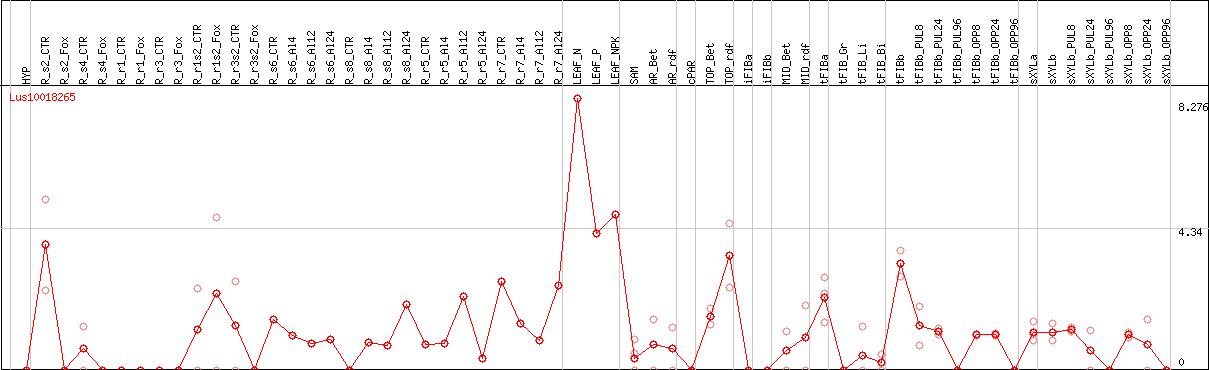
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018265 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.