External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G19640 332 / 2e-117
ATRAB-F2B, ARA7, Ara-7, AtRABF2b, AtRab5B
ARABIDOPSIS RAB GTPASE HOMOLOG F2B, Ras-related small GTP-binding family protein (.1)
AT5G45130 331 / 3e-117
ATRAB-F2A, RHA1, AtRab5A, AtRABF2a
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
AT3G54840 211 / 8e-70
ARA6, AtRABF1, Ara-6, AtRab5C
Ras-related small GTP-binding family protein (.1.2)
AT1G06400 150 / 1e-45
ARA2, AtRABA1a, AtRab11E, Ara-2
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A1A, Ras-related small GTP-binding family protein (.1)
AT5G45750 150 / 1e-45
AtRABA1c
RAB GTPase homolog A1C (.1)
AT4G18800 149 / 4e-45
AthSGBP, AtRab11B, AtRABA1d
RAB GTPase homolog A1D (.1)
AT1G16920 149 / 4e-45
ATRABA4B, RAB11, ATRABA1B
RAB GTPase homolog A1B (.1)
AT4G18430 146 / 4e-44
AtRABA1e
RAB GTPase homolog A1E (.1)
AT5G60860 146 / 6e-44
AtRABA1f
RAB GTPase homolog A1F (.1)
AT3G15060 145 / 7e-44
AtRABA1g
RAB GTPase homolog A1G (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040625
372 / 2e-133
AT5G45130 355 / 6e-127
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
Lus10038334
335 / 2e-118
AT5G45130 349 / 3e-124
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
Lus10034903
229 / 1e-76
AT5G45130 258 / 2e-88
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
Lus10036196
219 / 4e-74
AT5G45130 214 / 2e-72
ARABIDOPSIS RAB HOMOLOG F2A, RAB homolog 1 (.1)
Lus10036198
216 / 2e-72
AT4G19640 216 / 8e-73
ARABIDOPSIS RAB GTPASE HOMOLOG F2B, Ras-related small GTP-binding family protein (.1)
Lus10028820
213 / 3e-70
AT3G54840 354 / 3e-126
Ras-related small GTP-binding family protein (.1.2)
Lus10017462
211 / 3e-67
AT3G54840 349 / 1e-121
Ras-related small GTP-binding family protein (.1.2)
Lus10040745
149 / 2e-45
AT1G07410 394 / 7e-142
ARABIDOPSIS RAB GTPASE HOMOLOG A2B, RAB GTPase homolog A2B (.1)
Lus10025432
149 / 3e-45
AT5G60860 422 / 1e-152
RAB GTPase homolog A1F (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF00071
Ras
Ras family
Representative CDS sequence
>Lus10018285 pacid=23179369 polypeptide=Lus10018285 locus=Lus10018285.g ID=Lus10018285.BGIv1.0 annot-version=v1.0
ATGGCCGCCACCGGGAACAAGAACATCAACGCTAAGTTGGTCCTCCTTGGCGACCCAGGTGCTGGAAAATCTAGCCTTGTTTTACGATTTGTCAAAGATC
AGTTTGTTGAGTTTCAGGAATCAACCATTGGTGCTGCCTTCTTTTCACAAACACTGGCTGTGAATGATGTCACCGTGAAATTTGAGATATGGGACACAGC
CGGTCAAGAGAGATATCACAGTCTTGCTCCTATGTACTATCGTGGGGCAGCAGCAGCTATTATTGTGTATGACATTACGAATGCAGCCTCATTTGATCGA
GCAAAGAAGTGGGTCCTGGAACTGCAAGCACAAGGCAACCCAAATATGGTCATGGCTCTAGCTGGGAACAAAGCCGATTTGCTTGATGCAAGGAAGGTAG
AAGCTGAGGAGGCAGAAGCATACGCAAAGGAGAATGGCCTTTTCTTTATGGAGACCTCAGCGAAAACTGCAACAAACGTCAATGACATTTTCAACGAAAT
AGCGAAGAGACTTCCCCGAGTTCAGCCGGCACAGAATCCCACAGGGATGAATCTCAATGATAGACCTGCGGAAAGAGCAGCTAGCGGGACTTGCTGTTCT
TAA
AA sequence
>Lus10018285 pacid=23179369 polypeptide=Lus10018285 locus=Lus10018285.g ID=Lus10018285.BGIv1.0 annot-version=v1.0
MAATGNKNINAKLVLLGDPGAGKSSLVLRFVKDQFVEFQESTIGAAFFSQTLAVNDVTVKFEIWDTAGQERYHSLAPMYYRGAAAAIIVYDITNAASFDR
AKKWVLELQAQGNPNMVMALAGNKADLLDARKVEAEEAEAYAKENGLFFMETSAKTATNVNDIFNEIAKRLPRVQPAQNPTGMNLNDRPAERAASGTCCS
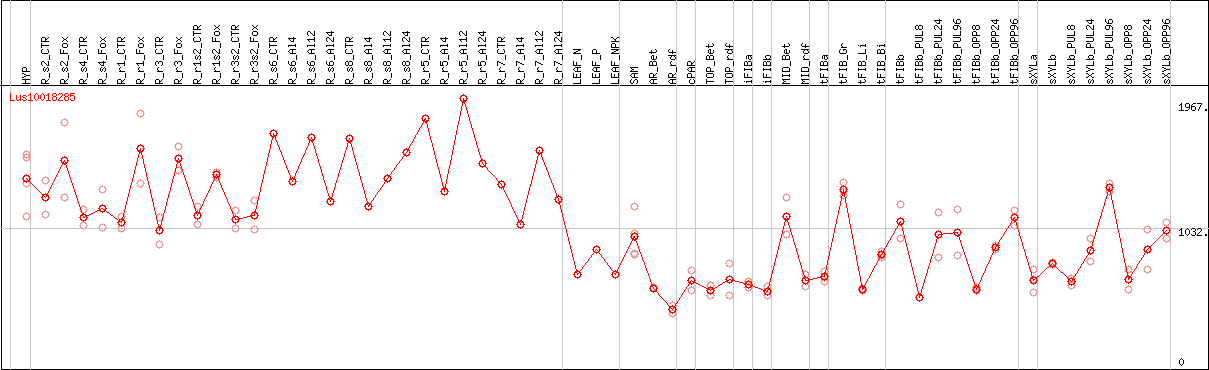
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018285 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.