Lus10018349 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10018349 pacid=23162185 polypeptide=Lus10018349 locus=Lus10018349.g ID=Lus10018349.BGIv1.0 annot-version=v1.0
ATGTCTTGGGGATTAGGATGGAAGCGGCCGTCGGAGATCTTCCGCTTAAGTCTCAACTATGGCACGGAGGAGTCCTGGGAGGATCCAATCCGTACATCGA
GCTCCTCATCGTGTTCTTCTTCGTCAACGATACAGCCGCTGGATCAGGATGGTGGGTTCCGCGTCGATTTGGATTGGACGGCCGGAGAAGATGAGGATCA
GTTGGCCCTGAGGTTGCAGTCGCAACTCACGGTGGCGCTCCCGATGCCGCAGGATTGTGTGACGGTGGAATTGGCGACGCAAGATGAGGAAGGAACGGAG
GAAGGCAATGTCAGAGTGGAGATGAATGTGGTGAAGAGGAGGGAGCCGTTGAGAGGGATCACGATCTCGAAGGCCGGCTCTGGACAGCAGAGCGATGGCA
TCGGCGTTTTGACGAGATTACTGAGGTCTAATTTAGCTTCTGCTGGAATTGGGGATGGTTCTGATTTTTCTTGTGCCGATCACTGGAAGACTGTCACCTT
GCTTAACCTCTGCGGCTGTGGTTTGGCGACATTGCCGCCTGAGCTTCTGCAATTGCCAACCCTCGAGAAACTCCTTCTTGACAACAATAGACTTACTTTT
TTGCCACCTGAGCTTGGGGAGCTGAAAACTTTGAAAGTTCTCAGTGTTGATTACAACTTGTTGGTTTCTGTTCCTGTGGAACTGAGGCAGTGTGTTCAAC
TCGTTGAGTTGTCATTGGAACATAACAAGCTTGTCCGTCCTCTTCTTGATTTCAGGGCCATGGCCGAGTTACAAATTCTTAGGCTATTTGGCAACCCTCT
GGAATTTCTTCCCGAGATTTTACCCCTCCACAAACTTCGCCACTTGACCCTTGCAAATATCAGGATTGTTTCTGATGAAAATTTAAGATCTGTGACTGTG
CAGATAGAGGGGGAGAACAATTCTTACTTTGGAGCATCTAGACATAAGTTAAGTGCGTTTTTTTCTCTTATATTCCGCTTCTCATCATGCCATCACTCTC
TCCTAGCATCTGCACTGGCAAAGATAGTGCAAGACCAAGGTAACCGTGCAGTAATTGGTAAAGATGAGAATGCAGTGCGACAGCTCATTAGCATGATCAG
TAGCGACGACCGTTATGTGGTTGAGCAAGCATGTTCTGCACTGTCCACTCTTGCTGGAGATGTTTCTATGGCAATGCAGTTGATTAAATGCGATATCATG
CAGCCTATTGAAACAGTGCTAAGATCTGTTTCCCAGGAAGAATTGCAAAAGTCGGCTTTGTTTGTGGTGGGTAATCTGGCGTTCTGCTTGGAAAATAGGC
GTATTTTGGTTACTTCTGAAAGTCTGCGGGACTTGCTATTGCGGTTGACAGTTACATCTGAACCTCGTGTGAACAAAGCTGCTGCTCGTGCCTTGGCCAT
TCTTGGAGAGAATGAAAATTTGAGACGTGCCGTAAAGGGAAGACCTGTGACCAAGCAAGGACTGCGTATTCTATCAATGGATGGAGGTGGAATGAAAGGT
CTGGCTACCGTACAGATTCTCAAGGCAATTGAGAAGGGAACTGGAAAACGAATACATGAGCTGTTTGACCTTATATGCGGCACATCAACGGGTGGAATGT
TGGCTGTTGCTCTTGGTATAAAGCTAATGTCCCTGGATCAATGCGAGGAAATATACAAAAATCTTGGGAAACTTGTGTTTGCTGAACCTGTGCCCAAGGA
TAACGAAGCTGCATCATGGAGAGAAAAGTTAGATCAGCTTTACAAAAGTTCGTCCCAGAGTTTTAGAGTTGTCGTACACGGATCCAAACACAGCGCAGAT
GAGTTTGAGAGACTGTTAAAGGAAATGTGTGCTGATGGGGATGGAGATCTATTGATAGACTCAGCAGTGAAATGCATTCCAAAAGTTTTTACCGTCTCAA
CTTTGGTTAGCACAATGCCAGCTCAGCCATTTATATTTCGTAACTATCAGTACCCAGTTGGGACACCAGAAGTGCCAACCACTCCTACAGAGGGATCTGG
TATCACTGTGTTAGGATCACCTTTTGCTGGTAGCCAAATTGGATACAGGCGCAGTGCTTCTGTTGGGAGTTGTAAACACCATGTTTGGCAAGCAATTAGA
GCATCGTCTGCTGCACCGTATTATCTTGATGACTATTCTGATGACACAAACCGCTGGCAAGATGGTGCTATTGTTGCGAATAATCCTACCATATTTGCCA
TAAGAGAAGCACAGCTTCTTTGGCCTGACACAAGAATCGACTGTCTTGTTTCGGTTGGGTGCGGTTCTGTTCCAACAAAGCCCCGGAAAGGTGGTTGGCG
CTATCTAGACACTGGGCAAGTTCTTATCGAAAGTGCATGCTCTGTGGACCGAGTGGAAGAAGCCTTAAGTACATTACTCCCCATGCTTCCGGAGATACAG
TATTTTCGTTTCAACCCAGTTGATGAACGCTGCGATATGGAACTGGATGAGACTGACCCTGCTGTATGGCTAAAGTTGGAAGCATCAGTTGAAGAATATA
TCCTAGAGAGTGCTGAAGCTTTCAAGAATGTCTGTGAGAGACTGGTGTTGCCATTCCAAAATGATGACAAGCTCTCAGACCACGTGAAAACTCAGCAACT
TCCTAGAGTAAAGACGTTCACTACTGGTGGGAGTGGCCCCACTCTCGGTTGGAGGTGTAATGTTCTACTTGTTGAAGCTCTTCACAGTCCTGATTCAGGA
AGAGTTATGCACCACGCCCGGGCACTCGAGTCGTTTTGTGCACGCAACAGCATAAAAATATCTCATGTTCATAGTACATCTGGAATTTCAAAGACGGTAC
CTCCCACAATCCCATCACCCTTCACCTCACCGCTAATCACCGGAAGCTTTCCTTCAAGCCCACTTCTATATAGTCCCGACTTTGCCTCCCAAAGAATGGG
TCGAATCGAAATGGTTCCACCTTTAAGCTTGGATGGAACTCAGTCTCTAAGAGGGGGATCATCTCCACCAATGTCCCCTGCCACACACAGACAGTTCTCT
TTACCCGTTCGATCATTGCATGAAAAGCTGCAGCATGCACCACAAGTGGGGATCGTGCACTTAGCCCTTCAGAATGATTCATCTGGCTCGATTTTAAGTT
GGCAGAATGACGTATTTGTGGTTGCTGAACCTGGAGAACTCGCTGACAAGTTTCTACAGAGTGTGAAGCTAAGTTTATTGTCCATGATGCGGTGCCACCA
ACGGAAGGTAACATCGTTCCTTTCGAAGATATCAACAGTTGCTGATCTCGTTAGCTCCAGACCGTACTTCCAAGTTGGAAATGTCGTCCACCGTTATATT
GGCCGGCAGACACAGGTGATGGAAGATGACCAAGAAATTGCAGCATACATGTTTCGAAGAACGGTTCCTTCAATTCATTTGACACCTGATGATGTCCGAT
GGATGGTCGGAGCTTGGAGAGACCGGATCATTATCTGCACGGGAACTTATGGTCCTTCCCAGACGCTAATCAAAGCCTTCATGGACTCAGGTGCAAAAGC
CGTCATATGTCCTTCCGCAGACCCAATGGACATTCCGGTGACAGCCGCCAGCTTGTCAGGTGACTTCAACTTCATGGAGAACGGTGGGAAGTTCGAGATC
GGAGAAGAGGAAGCAGAAGACGAAGATATCGAGCCTCCCAGTCCGGCAAGCGACTGGGAGGACAGCGATCCAGAGAAGGGGTGTGGGGGGTCAAACTCGA
TGGGGTTCTGGGACGACGACGAAGAGAAGCTAGCTCAGTTTCTAAGTCAGGTGTATGATTCGTTGTTCCGGGAAGGTGCCAGCGTGGATGTAGCCTTACG
TACTGCTCTGGCTTCGCATCGTAGACTGAGGTATTCATGCCATCTCCCCGGTTTGAAACCAACAGCCTGCAGCAGCTCCACGTCAGTACTGAATCTCCCT
GACCAGATGATGTCTGCTGCAGCAGCAGCTCCAGTTGCGGTCGGGACACGTGGCACGGTTGGTTCCCTTGTAAGGAAAGAGATCGACTACTTTACCAACT
TGGACACGGCCGAAAGCACTAGTGCTCGGAAATTTGCTGCTCCAGGGAGCATTAATGGTGGTGGTACTGCTACGTCTTCTTCTAGAAGATTTTGGCCGTG
GAGAATTAGCTGGAGAAGAAAGAAATGCGGCGGCAGTGGAGGCGGGAGCAAGTTGCTACATCCGGGGATGTGCGCCACCGTGGATGTTGGTGATTCTGGT
GTCGGGATCTCCGGCCGAAAGAAGGAAAAGAATGCGCCATCAATGGTGATCTCCAGATCGTTTAGCTCTCTGACGAGGAAGCACAACCCTAACCACCACC
TCCTTCCCTTCCTCGTCACCCACGCCGGCGATTCTCCAAGAACCTTCATCACAGCTTCAACATCTACTTTCCCTCCTTCTCCCTACGCTCTCCTTCCTTT
CCCGTCTTCTCCTTTTCCCCAACCTCACCGGTTCCTCTTCCCGCTCTCCAAATGGATCGCAGCCGCTCCGCTTCAAGGCCCTCTGTTCCTCTCATCGCCG
CCGTGGAAGCTGCTGCAGTCCTCGACTCCGCTTCATCTGCGCGGGAAGGGCGCCATTGCGCGGAAGCGAGCCGAGGCGTTGAAAAGCTTGAATTTGAGGC
TCAATTTGCTTCGGAATCGGATGGCTGCCACTGCGGACACGACGTGTGGCGGTGGAGTGACGGTGAATCGTGCAGCGGAGGCGGAGGAGGAGGAAGGCGG
TGTTTGGGAGAGCTTCGTGAATTTGCCTAATTTTATCTCTGTCAGTCGTTTGGTGTCCGGGCCTTTCCTTGGATGGATGATTGCTAATGAGATGTATTCC
GCTGCCTTTGTTGGCTTGGCAATCTCTGGAGCTAGTGATTGGCTTGATGGTTATGCTGCAAGAAAGATGGGGATCAATTCTGTAGTTGGTTCTTATCTTG
ATCCTCTTGCTGACAAGGTTCTTATAGGATCAGTTGCTGTAGCCATGGTTCATATGGATCTTTTACACCCTGGACTAGTCGGATTAGTTTTATTTCGTGA
TGTCGGACTGGTTGCTGCATCACTATATCATAGAGCTAACAGCTTGGGTTGGAAGTCGAATAGCTGGTCTGAATTCATAAACCTCAACGGATCCGGTGCC
GAGAAGATCCAGCCTCTATTCATAAGCAAGGTGAATACAGTGTTCCAGCTGCTTTTGGTCGCGGCAGCCCTTCTTCAACCAGAGTTTGGAACAGAGCAGA
CGCTAACATACGTAACATACTTGAGCTGGCTGGTGGCTTCGACGACAGCAGGTTCCACTGCAGCATACGGATTGAAATACTTCAACAACAGATCCTCGTT
GCTGGCTAGGAAGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10018349 pacid=23162185 polypeptide=Lus10018349 locus=Lus10018349.g ID=Lus10018349.BGIv1.0 annot-version=v1.0
MSWGLGWKRPSEIFRLSLNYGTEESWEDPIRTSSSSSCSSSSTIQPLDQDGGFRVDLDWTAGEDEDQLALRLQSQLTVALPMPQDCVTVELATQDEEGTE
EGNVRVEMNVVKRREPLRGITISKAGSGQQSDGIGVLTRLLRSNLASAGIGDGSDFSCADHWKTVTLLNLCGCGLATLPPELLQLPTLEKLLLDNNRLTF
LPPELGELKTLKVLSVDYNLLVSVPVELRQCVQLVELSLEHNKLVRPLLDFRAMAELQILRLFGNPLEFLPEILPLHKLRHLTLANIRIVSDENLRSVTV
QIEGENNSYFGASRHKLSAFFSLIFRFSSCHHSLLASALAKIVQDQGNRAVIGKDENAVRQLISMISSDDRYVVEQACSALSTLAGDVSMAMQLIKCDIM
QPIETVLRSVSQEELQKSALFVVGNLAFCLENRRILVTSESLRDLLLRLTVTSEPRVNKAAARALAILGENENLRRAVKGRPVTKQGLRILSMDGGGMKG
LATVQILKAIEKGTGKRIHELFDLICGTSTGGMLAVALGIKLMSLDQCEEIYKNLGKLVFAEPVPKDNEAASWREKLDQLYKSSSQSFRVVVHGSKHSAD
EFERLLKEMCADGDGDLLIDSAVKCIPKVFTVSTLVSTMPAQPFIFRNYQYPVGTPEVPTTPTEGSGITVLGSPFAGSQIGYRRSASVGSCKHHVWQAIR
ASSAAPYYLDDYSDDTNRWQDGAIVANNPTIFAIREAQLLWPDTRIDCLVSVGCGSVPTKPRKGGWRYLDTGQVLIESACSVDRVEEALSTLLPMLPEIQ
YFRFNPVDERCDMELDETDPAVWLKLEASVEEYILESAEAFKNVCERLVLPFQNDDKLSDHVKTQQLPRVKTFTTGGSGPTLGWRCNVLLVEALHSPDSG
RVMHHARALESFCARNSIKISHVHSTSGISKTVPPTIPSPFTSPLITGSFPSSPLLYSPDFASQRMGRIEMVPPLSLDGTQSLRGGSSPPMSPATHRQFS
LPVRSLHEKLQHAPQVGIVHLALQNDSSGSILSWQNDVFVVAEPGELADKFLQSVKLSLLSMMRCHQRKVTSFLSKISTVADLVSSRPYFQVGNVVHRYI
GRQTQVMEDDQEIAAYMFRRTVPSIHLTPDDVRWMVGAWRDRIIICTGTYGPSQTLIKAFMDSGAKAVICPSADPMDIPVTAASLSGDFNFMENGGKFEI
GEEEAEDEDIEPPSPASDWEDSDPEKGCGGSNSMGFWDDDEEKLAQFLSQVYDSLFREGASVDVALRTALASHRRLRYSCHLPGLKPTACSSSTSVLNLP
DQMMSAAAAAPVAVGTRGTVGSLVRKEIDYFTNLDTAESTSARKFAAPGSINGGGTATSSSRRFWPWRISWRRKKCGGSGGGSKLLHPGMCATVDVGDSG
VGISGRKKEKNAPSMVISRSFSSLTRKHNPNHHLLPFLVTHAGDSPRTFITASTSTFPPSPYALLPFPSSPFPQPHRFLFPLSKWIAAAPLQGPLFLSSP
PWKLLQSSTPLHLRGKGAIARKRAEALKSLNLRLNLLRNRMAATADTTCGGGVTVNRAAEAEEEEGGVWESFVNLPNFISVSRLVSGPFLGWMIANEMYS
AAFVGLAISGASDWLDGYAARKMGINSVVGSYLDPLADKVLIGSVAVAMVHMDLLHPGLVGLVLFRDVGLVAASLYHRANSLGWKSNSWSEFINLNGSGA
EKIQPLFISKVNTVFQLLLVAAALLQPEFGTEQTLTYVTYLSWLVASTTAGSTAAYGLKYFNNRSSLLARK
|
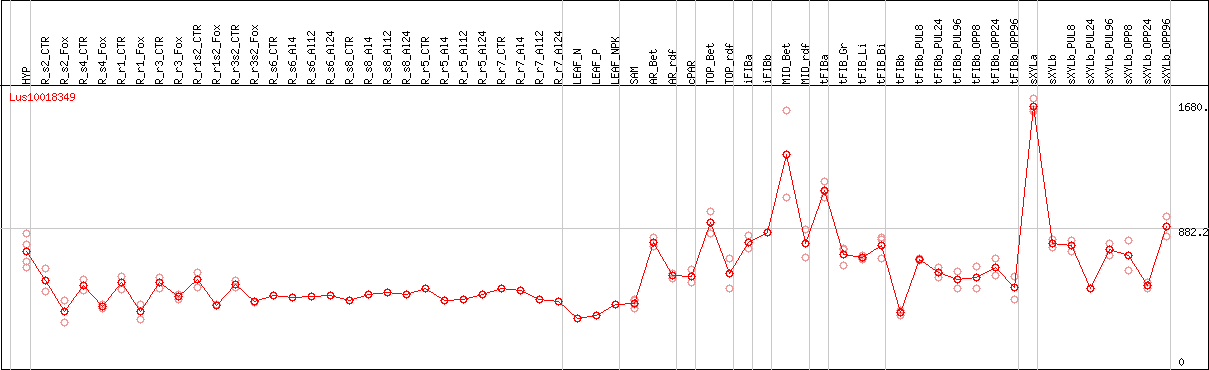
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018349 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
