External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G61660 187 / 2e-56
bHLH
bHLH112
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
AT3G20640 170 / 4e-49
bHLH
bHLH123
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G21340 134 / 5e-37
bHLH
bHLH103, B70
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G19500 129 / 4e-35
bHLH
bHLH113
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G20100 127 / 1e-33
bHLH
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT4G05170 123 / 2e-33
bHLH
bHLH114
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G27660 126 / 1e-32
bHLH
bHLH110
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G29100 122 / 2e-31
bHLH
bHLH068
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G31050 117 / 1e-29
bHLH
bHLH111
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G05710 109 / 4e-29
bHLH
bHLH153
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10007613
520 / 0
AT1G61660 194 / 1e-58
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Lus10038550
199 / 2e-59
AT3G20640 231 / 4e-70
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10014807
141 / 4e-38
AT3G20640 179 / 5e-51
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10025599
118 / 1e-32
AT1G05710 169 / 8e-55
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Lus10027064
118 / 1e-32
AT1G05710 174 / 9e-57
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Lus10023669
123 / 6e-32
AT2G20100 223 / 5e-69
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10010638
119 / 2e-30
AT1G27660 180 / 1e-51
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10013874
118 / 2e-30
AT3G19500 151 / 7e-44
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10014279
101 / 2e-23
AT5G44230 374 / 5e-120
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G029100
266 / 1e-85
AT1G61660 216 / 1e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.011G033000
259 / 2e-83
AT1G61660 216 / 2e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein
Potri.001G410600
225 / 2e-70
AT3G20640 274 / 2e-87
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.011G129500
221 / 9e-69
AT3G20640 289 / 3e-93
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.018G083700
137 / 9e-38
AT4G29100 279 / 3e-91
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.003G051600
130 / 4e-35
AT4G29100 197 / 4e-60
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G185900
129 / 4e-35
AT4G29100 212 / 5e-66
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G299300
127 / 8e-35
AT3G19500 152 / 4e-45
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.009G094300
124 / 2e-33
AT3G19500 161 / 2e-48
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.005G230800
127 / 3e-33
AT1G27660 247 / 3e-77
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10018400 pacid=23162196 polypeptide=Lus10018400 locus=Lus10018400.g ID=Lus10018400.BGIv1.0 annot-version=v1.0
ATGGGCTTTGGCGTCGGTGTATCGTCTTCTTCGCCGTCGTCCGAGGATTGGAATCAGCGACAGCATATTGGGAGTGGTAACTACAACAGTACTTCAATGC
TGGAAGATGAAATGAACAGCCGCGGTCGTCCACTCAACCATAATCTAAGTAGGTTGACCAATGATTTCTCATCGAATCCGATTCATGGCTACCCGATTCA
AGGCTTGTTCGAATCCGATTCACAACAAAACCCGATATTCGTCAACAACTTCCCGTCCCCTTCTGCTACTTACGGGATCGGAAACAGCTTGTTGAATGGA
TTGTCTCCTTCTTGGTCGAAATTCCAAAACCCTAATCATCAAAACGACAATCACCACCAAGATCACCTCCGGTTCTCGAATGGTGTGATGCCATTTTGGA
ACAATCAAACAGCGACCCAGACCGATAGTATCCAAACCGGTTATGCACCGACACAAACCCGGTTCGTTGTCCCGGCTTTCAACGAGAAACCAACCAGTTG
CCCCAGTAGAAGCTCCACTAAGTCTAAGAAGGAAGAAGCTACTAATAATACTACCAAGGAGTGGAAACCGAAAACCAAGAAGGAAACCGGTTCCGAGTCG
TCTTTCAAACGACCTCGAATCGAGACTCCATCATCGTTGCCAACTTTTAAGGTTCGGAAAGAGAAATTGGGCGACCGAATAACCGCTCTACAACAATTAG
TTTCGCCCTTCGGGAAGACGGATACAGCATCAGTGCTCCATGAAGCCATTGAGTACATCAAATTCCTCCATGACCAAGTCGGTGTTTTAAGCACTCCATA
CATGAAGAATGGAAACCCTATTCAGCACCACTTACAGGGTTGTGACAAATTGGACCAAAACGAACAAGACCTAAGAAGCCAAGGGTTATGTTTGGTCCCA
ATCTCAAGCACTTTCCCAGTTGCTCATGAGACCACAGCAGATTTCTGGACTCCAACTTTTGGTGGCTCCTTCAGGTAG
AA sequence
>Lus10018400 pacid=23162196 polypeptide=Lus10018400 locus=Lus10018400.g ID=Lus10018400.BGIv1.0 annot-version=v1.0
MGFGVGVSSSSPSSEDWNQRQHIGSGNYNSTSMLEDEMNSRGRPLNHNLSRLTNDFSSNPIHGYPIQGLFESDSQQNPIFVNNFPSPSATYGIGNSLLNG
LSPSWSKFQNPNHQNDNHHQDHLRFSNGVMPFWNNQTATQTDSIQTGYAPTQTRFVVPAFNEKPTSCPSRSSTKSKKEEATNNTTKEWKPKTKKETGSES
SFKRPRIETPSSLPTFKVRKEKLGDRITALQQLVSPFGKTDTASVLHEAIEYIKFLHDQVGVLSTPYMKNGNPIQHHLQGCDKLDQNEQDLRSQGLCLVP
ISSTFPVAHETTADFWTPTFGGSFR
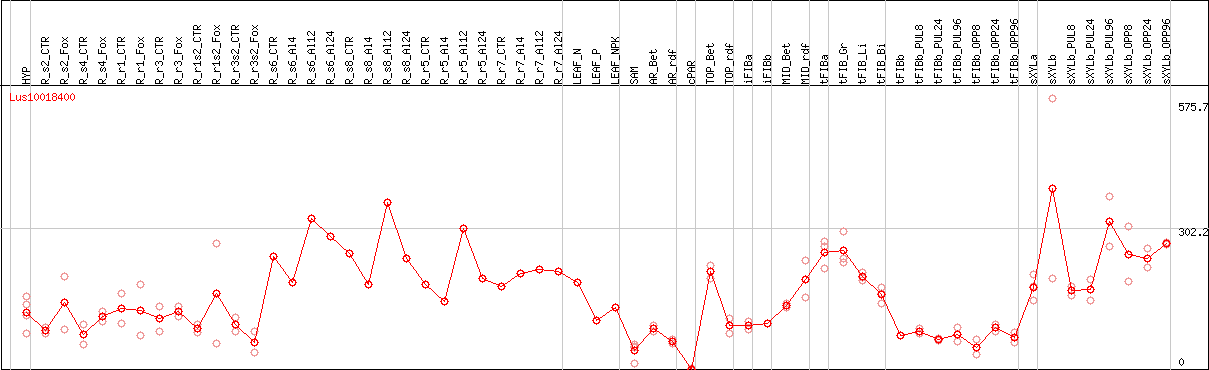
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.