External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G61105 225 / 2e-75
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT1G52900 210 / 2e-69
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT5G48770 80 / 2e-17
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G17615 74 / 2e-15
Disease resistance protein (TIR-NBS class) (.1)
AT1G72840 73 / 4e-15
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT1G72860 70 / 7e-14
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G17600 69 / 2e-13
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G72950 68 / 2e-13
Disease resistance protein (TIR-NBS class) (.1)
AT4G09430 65 / 3e-12
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G40100 64 / 5e-12
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011216
395 / 3e-142
AT1G61105 228 / 2e-76
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Lus10007823
76 / 4e-16
AT1G27170 322 / 3e-92
transmembrane receptors;ATP binding (.1.2)
Lus10011741
76 / 5e-16
AT5G36930 540 / 4e-172
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10007829
75 / 2e-15
AT5G36930 330 / 2e-94
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10042020
74 / 3e-15
AT1G72890 147 / 3e-39
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
Lus10012311
74 / 5e-15
AT5G36930 354 / 6e-103
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10013729
72 / 8e-15
AT5G36930 310 / 2e-92
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10000423
72 / 1e-14
AT1G27180 338 / 5e-95
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10004747
72 / 2e-14
AT5G36930 342 / 3e-99
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G038200
261 / 1e-89
AT1G61105 233 / 1e-78
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Potri.011G046800
251 / 9e-86
AT1G61105 230 / 1e-77
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Potri.001G403800
238 / 1e-80
AT1G52900 233 / 3e-78
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Potri.011G123100
238 / 2e-80
AT1G52900 229 / 2e-77
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Potri.015G043501
84 / 1e-18
AT1G27170 534 / 5e-176
transmembrane receptors;ATP binding (.1.2)
Potri.019G019053
84 / 1e-18
AT5G36930 655 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G017082
83 / 2e-18
AT5G36930 638 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.005G004500
79 / 9e-18
AT5G36930 152 / 3e-42
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G001971
80 / 3e-17
AT5G36930 427 / 3e-130
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.015G043600
80 / 3e-17
AT1G27170 1195 / 0.0
transmembrane receptors;ATP binding (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF01582
TIR
TIR domain
Representative CDS sequence
>Lus10018470 pacid=23162101 polypeptide=Lus10018470 locus=Lus10018470.g ID=Lus10018470.BGIv1.0 annot-version=v1.0
ATGCAACGTGCAACCTCAGCAATAGCCAAGGCATCTACCAAGATCCTGCGCAACGCACCGAAGGTCGACGTTTTCATCAACCACCGCGGGATAGACACCA
AGAGGACGATCTCGGGCTTGCTCTACGACCGTCTGGCCTCTCCCGGGTCACGGATCATTCCTTTCTTGGATAGCAAGAATATGAAGCCGGGGGATCGCCT
GTTTGACAACATAAATGCTGCCATTCAGGGGTGTAAAGTTGGGGTCGCAGTGTTCTCCCCGAGATATTGCGACTCGTACTTCTGTCTCCATGAGCTGGCG
ATGCTCATGGAGAGTAAAAAAAGGGTGATTCCCATCTTCTGTGACATGAAGCCCTCAGAGCTCCACGTTCCTGACAAGGGGGAGTGCTCGGAGGAGGAGA
TGCGACGGTTTAAGTATGCCCTCGAGGAGGCTAAGTACACCGTCGGATTAGCCTTCGACACCCGGGAAGGGAACTGGTCGGAGTTCATGAAGAAGGCAAC
CGACGCCGTTTTGAAGAACTTGATAGAGGTTGAAGGGGAAAAGTTCGGGTACTCATACGAGAACCCGAACTATACCGTGCAGAAGGCCACGACATATCGC
AGCCGTTGA
AA sequence
>Lus10018470 pacid=23162101 polypeptide=Lus10018470 locus=Lus10018470.g ID=Lus10018470.BGIv1.0 annot-version=v1.0
MQRATSAIAKASTKILRNAPKVDVFINHRGIDTKRTISGLLYDRLASPGSRIIPFLDSKNMKPGDRLFDNINAAIQGCKVGVAVFSPRYCDSYFCLHELA
MLMESKKRVIPIFCDMKPSELHVPDKGECSEEEMRRFKYALEEAKYTVGLAFDTREGNWSEFMKKATDAVLKNLIEVEGEKFGYSYENPNYTVQKATTYR
SR
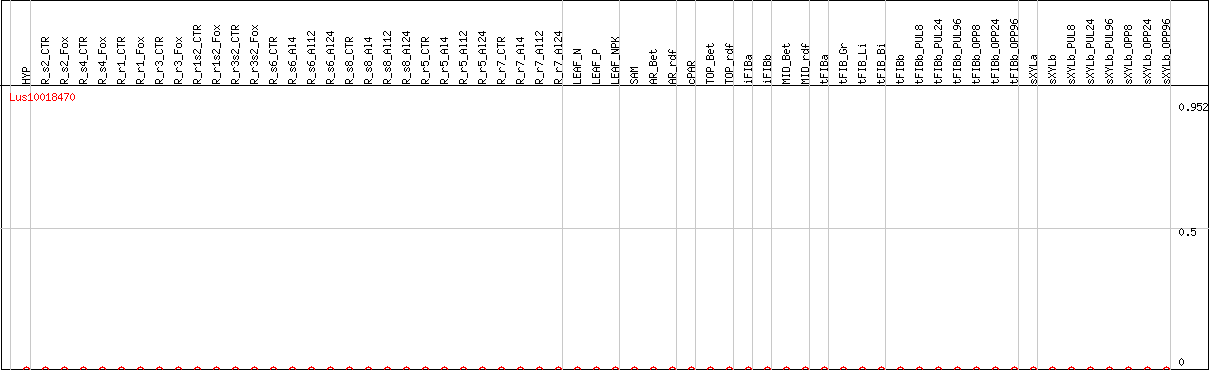
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018470 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.