Lus10018512 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10018512 pacid=23180376 polypeptide=Lus10018512 locus=Lus10018512.g ID=Lus10018512.BGIv1.0 annot-version=v1.0
ATGGAGTTTAACAAAGTCTTCACCTTTCTCACACTACTACTATTCCTCTGTTTCCTCCCTTCCCCTGTTTCCTCATCCATAACATTCACCTCCGCTACTT
CTTCTTCCGCTGTCCTCGTACGTGACAGCGCCGGAGAATCTCTAATCTCCGACGGCGGAGTATTCCAGCTCGGCTTCTTCACTCCGAATCGAACCAAACT
TCCATCTCGGTATTTAGGAATATGGTACAGGAATCACCCGTCGGTAATCGTCTGGGTAGCTAATCGCGAAATGGACCTCCCCGATACGACAGGGTTCCTC
TGTTTCACACCGGACGGCAACCTCCAAGTCCTCGACGGAACAACAACAACCAACGATTCTTCTTACTGGTCAACTGATTTGAAACAGTTGACCGTTAATA
ATTACACACTGAGTCTCCTGAATTCCGGGAACTTGATCCTGTACGCAGGAAATTCAATGGATGTGCTCTGGCAGAGCTTTGACAATCCAACGGACACTTT
CCTTCCCGGGATGAAAATGGTTAGTCATCTGAATTTAAGTTCCTGGAACTCTCCCGACGATCCTTCCACCGGAGATTACCTCTTCCTGACCGACGAAACG
ATGTCGCAGATCCTGAACAACTCTATCCCGTTCTGGAATTCAGCTGACCAAATTGGATCGTGTCAACTCCCGAACTTCGTAGTCGAATTCCTCTCCAATT
CGTCGTCCAATCCGTTGAAATTAGCTGATATAAATCCTCCACGACTTGTGATGGGTTTCGACGGCAGGATCCAGTTTTTTGAAACGGCGAACCAAACCTC
ACCATCCTGGATCACCCCTACCGACATTTGCAGCTTGGTAGCTCCCTGTGGGAGCTTCGGAAGCTGCAACGGAGGGTTCCGCGTCCCTTGCAAGTGTCTT
CCTGGATTCGTGCCGAAATCCGTCGAGAATTGGGAAACAGGGGATTTCTCGGAAGGGTGCAAGAGGGTTTCTCAGACGTGTCAGAACAAGAAAGACGGGA
GTACTAGTTTCTTCTTGAAGCTGAAGGTGGTGAAAGTGCAGAGGGAGACCGGTTTGAATGGAGATTTTGACGAGGAATATGAATGTAGAGAGTGGTGCTT
GGGGAACTGTAATTGTCAGGGTTACTCGTTTGATCATACGACTGCTTGCAGGTTTTGGACCGATGACTTGAAGGACATTGTGGAGATTAATGGGAGTAGT
GCTGCTCAGGGTACTGTCGATCTCAACGTTCGAGTCTCCATTTCGGACATAGAGTTGACAAGAAGAGATTGCAAGCCATGCGGGACTAACACCATCCCTT
ATCCACTAAGCACCGGACCAACCTGTGGAGATCCAATGTACTCAAGCTTCACTTGTAACAACAGCACCGGCTTGCTTACTTTCCACGCAATCAATGCAAG
CTACCCAGTGACTCGAATTGACAGATGGTTGCACAAGTTCTTCATCCAGGTCAACGATGAGTACTGCTCAGAACCAAACGTCATAGAGTTGGATCGGTCG
TTACCGTTTACTACCAACGATGGTAAATGCAATGCCGAACCAGTCAATCTCAAGTTAGCCCGATTCCAGGGAGGAAAAGTGTCCAAAGAAATGGAGATCG
TGTGGGAACCGCCTTCAGAGCCACTCTGTAGCTCTGAAGAGGACTGCCAGGGCTGGCTGAACTCGAGCTGTCGCGAAGTTAACGACGGGACTGGAGATTA
TAAGAGGTGTTTGTGTAGTGGGAATTCGCGCTGGGTTGGGGTCAATGCTACTTGCCAGACAAATGTTTTGTCTAACCACGTCGGATCATCATCTTCAAAG
AGAAGAAGACTATTGCTGTATTCCATTGTGTTTGGAATTCTTGGAGGTGTTCTGATAATGAGCTTAGCTACTGTAATACTGCAAATGAAAAGAAAAAGAA
GAAAGGATGTTCAAGCTTTGACAGAGAGCAAGTATGAGAAGGAGGAGAATCTGGCATTCAGACTCTACGACGGCGAAAAGCGGGTGAAGGAGTTCATGAA
CTCGACCGAGTTCGGAGAAGACGATAAGAAAGACATAGACGTTCCGTTCTTTGATTTGGAATGCATTCTTTCTGCTACAAACTTCTTCTGCCACACCAAC
AAGCTCGGCCAGGGCGGTTTTGGCCCTGTTTTCAAGGGGAAGTTTCCGGGGGGACAAGAGATAGCTGTGAAGAGGCTTTCAAGTGGTTCAACGCAAGGGC
TAGAAGAGTTTAAGAACGAGGTCATTTTGATCGCGAAGCTGCAGCACAGGAACCTGGTCAGGTTGCTGGGATTCTGTGTTGAAGGAAGTGAAAAGATGTT
GTTGTATGAGTATATGCCTAACAAGAGCTTGGATTCCTTCATCTTCGATCGAAGTCGAAGTGTGTTGCTGAATTGGGAGGTGAGGTTCAACATCATAATG
GGGATAGCTAGAGGGATGCTCTACCTCCATCATGATTCGAGGCTGACGATTGTCCATAGAGACTTGAAGACGAGCAATGTACTTCTTGACGAGGACATGA
ACCCCAAGATATCGGATTTTGGCATGGCTAGAATAGTGGCGGCCAAGCAAACTGAAGTCGACACTGTTCGGAACTCTCGTGTCGGCCCATTTTTTGGTTG
CGGATACATGTCTCCAGAGTATGCGCTAGATGGGAAGTTCTCAACGAAATCCGACGTTTTCAGCTTTGGAATAGTTGTTCTTGAGATCCTCATAGGCAGG
AAGAACACTGGATTTTACAAGTCTGAAGATGGCATTTCCCTTGCAAGCCATGCGTGGGGTAAGTGGAGAGAGGGCAATTTAATGGAAGTGATGGAGCAGT
CATTACTCGAGACATGCGATTCCAACGAGTTCACGAGGTGCGCGATCGTGGCGTTGTTGTGCGTGCAGGAGGATCCTAGCGACCGACCCACCATGGCCGA
TGTCGTATTCATGCTTGGAAGCGATGCGTCGAGCCTTACTCATCCTAAAGAGCCTGCATTTTCGGTTAGGAACCATGCAGCCACGGCTCCTTCTACCAGC
GTTGGGACTATGGAGATCACAACCGAATTACAAGGAAGATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10018512 pacid=23180376 polypeptide=Lus10018512 locus=Lus10018512.g ID=Lus10018512.BGIv1.0 annot-version=v1.0
MEFNKVFTFLTLLLFLCFLPSPVSSSITFTSATSSSAVLVRDSAGESLISDGGVFQLGFFTPNRTKLPSRYLGIWYRNHPSVIVWVANREMDLPDTTGFL
CFTPDGNLQVLDGTTTTNDSSYWSTDLKQLTVNNYTLSLLNSGNLILYAGNSMDVLWQSFDNPTDTFLPGMKMVSHLNLSSWNSPDDPSTGDYLFLTDET
MSQILNNSIPFWNSADQIGSCQLPNFVVEFLSNSSSNPLKLADINPPRLVMGFDGRIQFFETANQTSPSWITPTDICSLVAPCGSFGSCNGGFRVPCKCL
PGFVPKSVENWETGDFSEGCKRVSQTCQNKKDGSTSFFLKLKVVKVQRETGLNGDFDEEYECREWCLGNCNCQGYSFDHTTACRFWTDDLKDIVEINGSS
AAQGTVDLNVRVSISDIELTRRDCKPCGTNTIPYPLSTGPTCGDPMYSSFTCNNSTGLLTFHAINASYPVTRIDRWLHKFFIQVNDEYCSEPNVIELDRS
LPFTTNDGKCNAEPVNLKLARFQGGKVSKEMEIVWEPPSEPLCSSEEDCQGWLNSSCREVNDGTGDYKRCLCSGNSRWVGVNATCQTNVLSNHVGSSSSK
RRRLLLYSIVFGILGGVLIMSLATVILQMKRKRRKDVQALTESKYEKEENLAFRLYDGEKRVKEFMNSTEFGEDDKKDIDVPFFDLECILSATNFFCHTN
KLGQGGFGPVFKGKFPGGQEIAVKRLSSGSTQGLEEFKNEVILIAKLQHRNLVRLLGFCVEGSEKMLLYEYMPNKSLDSFIFDRSRSVLLNWEVRFNIIM
GIARGMLYLHHDSRLTIVHRDLKTSNVLLDEDMNPKISDFGMARIVAAKQTEVDTVRNSRVGPFFGCGYMSPEYALDGKFSTKSDVFSFGIVVLEILIGR
KNTGFYKSEDGISLASHAWGKWREGNLMEVMEQSLLETCDSNEFTRCAIVALLCVQEDPSDRPTMADVVFMLGSDASSLTHPKEPAFSVRNHAATAPSTS
VGTMEITTELQGR
|
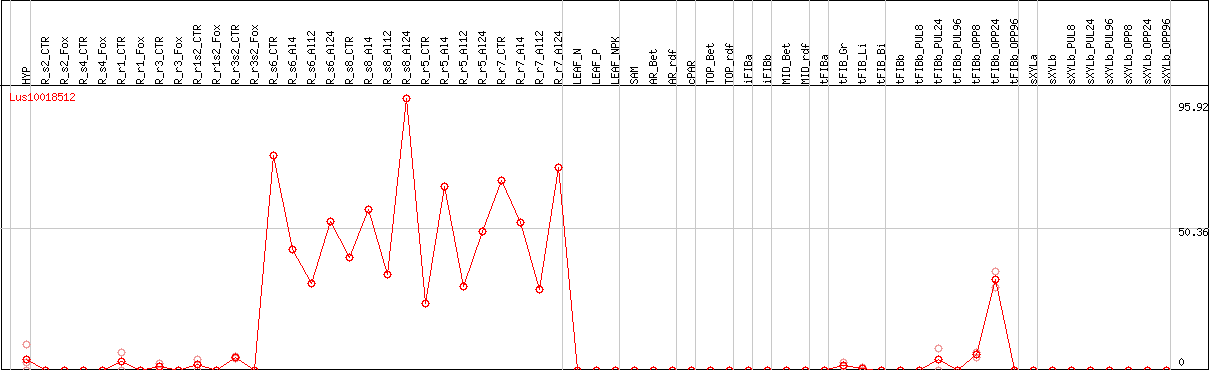
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018512 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
