External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G09310 194 / 2e-63
Protein of unknown function, DUF538 (.1)
AT1G56580 189 / 1e-61
SVB
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
AT5G46230 134 / 3e-40
Protein of unknown function, DUF538 (.1)
AT4G24130 115 / 9e-33
Protein of unknown function, DUF538 (.1)
AT5G49600 114 / 3e-32
Protein of unknown function, DUF538 (.1)
AT1G30020 105 / 5e-29
Protein of unknown function, DUF538 (.1)
AT1G02813 47 / 8e-07
Protein of unknown function, DUF538 (.1)
AT5G37070 44 / 1e-05
Protein of unknown function, DUF538 (.1)
AT3G08890 43 / 3e-05
Protein of unknown function, DUF538 (.1.2)
AT2G03350 42 / 9e-05
Protein of unknown function, DUF538 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039866
325 / 4e-115
AT1G09310 196 / 5e-64
Protein of unknown function, DUF538 (.1)
Lus10031428
215 / 5e-72
AT1G09310 185 / 2e-60
Protein of unknown function, DUF538 (.1)
Lus10010904
215 / 6e-72
AT1G09310 185 / 1e-60
Protein of unknown function, DUF538 (.1)
Lus10017330
129 / 7e-38
AT4G24130 209 / 3e-70
Protein of unknown function, DUF538 (.1)
Lus10001674
126 / 4e-37
AT4G24130 212 / 2e-71
Protein of unknown function, DUF538 (.1)
Lus10037804
114 / 4e-32
AT5G49600 144 / 2e-44
Protein of unknown function, DUF538 (.1)
Lus10017087
115 / 2e-31
AT5G49600 145 / 3e-43
Protein of unknown function, DUF538 (.1)
Lus10017655
107 / 1e-29
AT1G56580 115 / 1e-33
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Lus10000355
105 / 6e-29
AT5G46230 117 / 1e-34
Protein of unknown function, DUF538 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G011200
210 / 8e-70
AT1G56580 192 / 4e-63
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.013G007000
204 / 7e-68
AT1G56580 176 / 5e-57
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.004G064700
158 / 1e-49
AT5G46230 184 / 4e-61
Protein of unknown function, DUF538 (.1)
Potri.003G146400
133 / 1e-39
AT4G24130 211 / 5e-71
Protein of unknown function, DUF538 (.1)
Potri.001G084200
131 / 4e-39
AT4G24130 208 / 5e-70
Protein of unknown function, DUF538 (.1)
Potri.010G149100
120 / 2e-34
AT5G49600 152 / 3e-47
Protein of unknown function, DUF538 (.1)
Potri.004G056700
91 / 2e-23
AT1G09310 99 / 4e-27
Protein of unknown function, DUF538 (.1)
Potri.004G084600
82 / 4e-20
AT1G56580 96 / 9e-26
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.017G133300
81 / 1e-19
AT1G56580 69 / 1e-15
SMALLER WITH VARIABLE BRANCHES, Protein of unknown function, DUF538 (.1)
Potri.008G092500
48 / 4e-07
AT2G03350 281 / 4e-98
Protein of unknown function, DUF538 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04398
DUF538
Protein of unknown function, DUF538
Representative CDS sequence
>Lus10018630 pacid=23180472 polypeptide=Lus10018630 locus=Lus10018630.g ID=Lus10018630.BGIv1.0 annot-version=v1.0
ATGTCTCTAGTAACAGAAGCCATCCGAGCCAGCGCCACCGAGGTCTACACAGGCGACGAGATGTGCCAAGTCAAATCCAAGGAGCTCCTCAAGGAGATCG
AACTCCCCAACGGCCTCCTCCCCCTACACGACATCGAGGAGTGCGGGATCGTTCGCGAGACCGGGTTCGTGTGGCTGAAGCAGAAGAAGAGCATCACCCA
CAAGTTCGACAAGATCGGGAAGCTCGTCAGCTACGCCCCCGAGGTCACTGCCGTGGTCGAGAAGGGCAAGATCAAGAAGCTCACGGGCGTCAAGACCAAG
GAGCTTCTCATCTGGGTGTCGCTCTCCGACATCTACACCGCAGACGAGGGGAAAATTACCTTTAAGACCCCGACGGGTCTTTACAGGTCTTTTCCCGCGA
CCGCTTTTCAAGTTCCCGAAGAGGCAGTGAAGGAAGCGTCCTCGCANNNNNNNNNNCCACCACCGTCGGTGCAGCACCACCAAAAAAAGGGGCGGGCGAG
CCCGTGGCGGTTAAGGAAGTGTAAGTTTGGTTTGGGTTATGAGACACGAGTAAAGAAATCATGTCCATCGATCATGCAATACCAATGA
AA sequence
>Lus10018630 pacid=23180472 polypeptide=Lus10018630 locus=Lus10018630.g ID=Lus10018630.BGIv1.0 annot-version=v1.0
MSLVTEAIRASATEVYTGDEMCQVKSKELLKEIELPNGLLPLHDIEECGIVRETGFVWLKQKKSITHKFDKIGKLVSYAPEVTAVVEKGKIKKLTGVKTK
ELLIWVSLSDIYTADEGKITFKTPTGLYRSFPATAFQVPEEAVKEASSXXXXPPPSVQHHQKKGRASPWRLRKCKFGLGYETRVKKSCPSIMQYQ
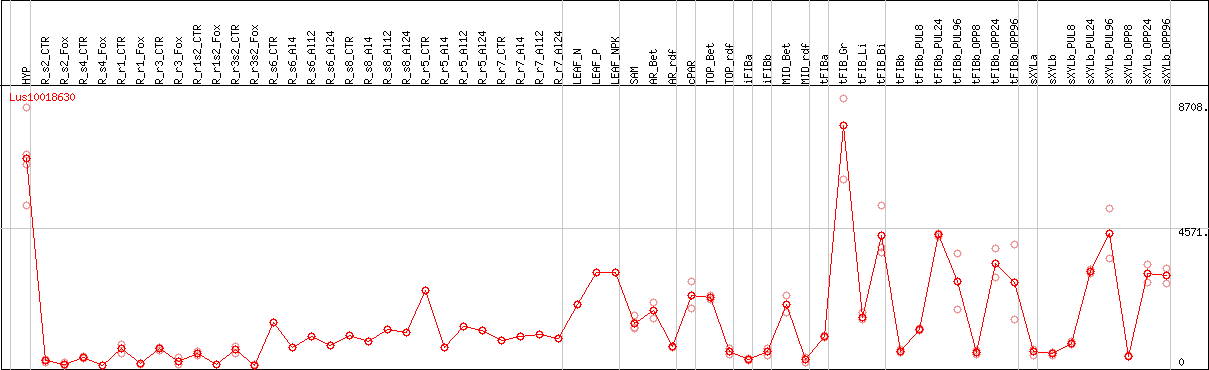
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018630 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.