External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G60060 209 / 7e-65
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1)
AT5G53900 119 / 6e-31
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
AT3G15240 115 / 2e-29
Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
AT2G31280 76 / 3e-15
bHLH
bHLH155 ,CPuORF7
conserved peptide upstream open reading frame 7 (.1.2.3.4)
AT1G06150 69 / 6e-13
bHLH
bHLH089, EMB1444
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT2G27230 66 / 1e-11
bHLH
LHW, bHLH156
LONESOME HIGHWAY, transcription factor-related (.1.2)
AT5G24352 45 / 2e-06
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
AT1G64625 46 / 2e-05
bHLH
bHLH157
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2), Serine/threonine-protein kinase WNK (With No Lysine)-related (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024800
553 / 0
AT1G60060 221 / 8e-70
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1)
Lus10012986
209 / 8e-65
AT1G60060 428 / 8e-150
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1)
Lus10029179
208 / 1e-64
AT1G60060 429 / 6e-150
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1)
Lus10041936
120 / 5e-31
AT5G53900 419 / 7e-146
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
Lus10005549
108 / 1e-26
AT5G53900 410 / 7e-142
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
Lus10033880
65 / 3e-11
AT2G31280 481 / 7e-160
conserved peptide upstream open reading frame 7 (.1.2.3.4)
Lus10013348
52 / 6e-07
AT2G27230 355 / 6e-112
LONESOME HIGHWAY, transcription factor-related (.1.2)
Lus10000609
47 / 1e-05
AT2G31280 483 / 2e-161
conserved peptide upstream open reading frame 7 (.1.2.3.4)
Lus10033222
44 / 0.0001
AT1G64625 211 / 2e-60
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2), Serine/threonine-protein kinase WNK (With No Lysine)-related (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G024000
379 / 1e-131
AT1G60060 243 / 4e-77
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1)
Potri.010G096400
215 / 2e-67
AT1G60060 459 / 4e-162
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1)
Potri.001G397800
120 / 7e-31
AT5G53900 439 / 1e-153
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
Potri.011G116400
117 / 5e-30
AT5G53900 438 / 3e-153
Serine/threonine-protein kinase WNK (With No Lysine)-related (.1), Serine/threonine-protein kinase WNK (With No Lysine)-related (.2)
Potri.005G222500
76 / 5e-15
AT2G31280 568 / 0.0
conserved peptide upstream open reading frame 7 (.1.2.3.4)
Potri.002G040600
68 / 2e-12
AT2G31280 437 / 7e-144
conserved peptide upstream open reading frame 7 (.1.2.3.4)
Potri.001G216900
61 / 4e-10
AT2G27230 363 / 3e-113
LONESOME HIGHWAY, transcription factor-related (.1.2)
Potri.003G144900
60 / 9e-10
AT2G27230 233 / 2e-66
LONESOME HIGHWAY, transcription factor-related (.1.2)
Potri.001G086000
59 / 1e-09
AT2G27230 248 / 7e-72
LONESOME HIGHWAY, transcription factor-related (.1.2)
Potri.009G017700
53 / 2e-07
AT2G27230 355 / 2e-110
LONESOME HIGHWAY, transcription factor-related (.1.2)
PFAM info
Representative CDS sequence
>Lus10018736 pacid=23176247 polypeptide=Lus10018736 locus=Lus10018736.g ID=Lus10018736.BGIv1.0 annot-version=v1.0
ATGGAAGGTGGATTTCCACTGCTTAACTGCCTTCTGCATTACACATTGAGGAGTTTGTGTTCAAGTTCATCAGATTCCAACTCATCCAAATGGGTATATG
CTGTGTTCTGGAGAATGTTGCCCAGGAATTACCCACCTCCTAAGTGGGATCATTGTGGAACAGCTCTTGATCGCTCCAAAGGAAACAAAAGAAACTGGAT
GCTAGTCTGGGAAGATGGCTTCTGTGATTTCCTGGAATGTGAGAGGCAAGGAAATGGGTACAAGTTTGGAGCTGATGTCTTCTTCAAATTGTCTCATGAA
GTGTATAGCTATGGAGAAGGATTATTAGGGAAAGTTGCAGCAGAGAGGAGTCAGAAATGGGTTTTCAGTGAATCAACCAATGAGCATGAACCAAATCCCA
CCTCCTCCTGGAATATGGCACTTGAACCTCAACCAAGACCTTGGGAATGCCAGTTCCAGTCAGGCATTCAGACAATTGCAATCATTTCAGTCAGAGAAGG
CATCATTCAACTGGGTTCATTTCACAAGATTGTGGAAGATGCAAATCTGGTGATCAACGTTCAGAGAACATTCAGCTATCTTCAGAGCATACCTGGAACA
TTTTCCATTCATAGAACAAACTTACCGATCCAACATACAAACGTTGACAGGAAAGAATCGGCAAAGAGACAGACGATCGGAATGAAGAGAGCGTTTGATC
AAAGAAGGGATGATGAGGATTTCCCTTTCAAGTCCATCAACTTAGGATGGAACAGCCCACAATGTGGAGGTGGCCCTGAGAAATTCATCTTCTCAATACC
GCCTCTCTTGCCCACCATGCCTTTTAGCTGTAGACCAAAGCTCCATTATTCTCGTGACCATACTGGAGATGTAGTAAACTGCTGCAACAATGGCATACCT
ACTAGCAACACAATCAAGTCTGAAGATGAGGGTAACTAA
AA sequence
>Lus10018736 pacid=23176247 polypeptide=Lus10018736 locus=Lus10018736.g ID=Lus10018736.BGIv1.0 annot-version=v1.0
MEGGFPLLNCLLHYTLRSLCSSSSDSNSSKWVYAVFWRMLPRNYPPPKWDHCGTALDRSKGNKRNWMLVWEDGFCDFLECERQGNGYKFGADVFFKLSHE
VYSYGEGLLGKVAAERSQKWVFSESTNEHEPNPTSSWNMALEPQPRPWECQFQSGIQTIAIISVREGIIQLGSFHKIVEDANLVINVQRTFSYLQSIPGT
FSIHRTNLPIQHTNVDRKESAKRQTIGMKRAFDQRRDDEDFPFKSINLGWNSPQCGGGPEKFIFSIPPLLPTMPFSCRPKLHYSRDHTGDVVNCCNNGIP
TSNTIKSEDEGN
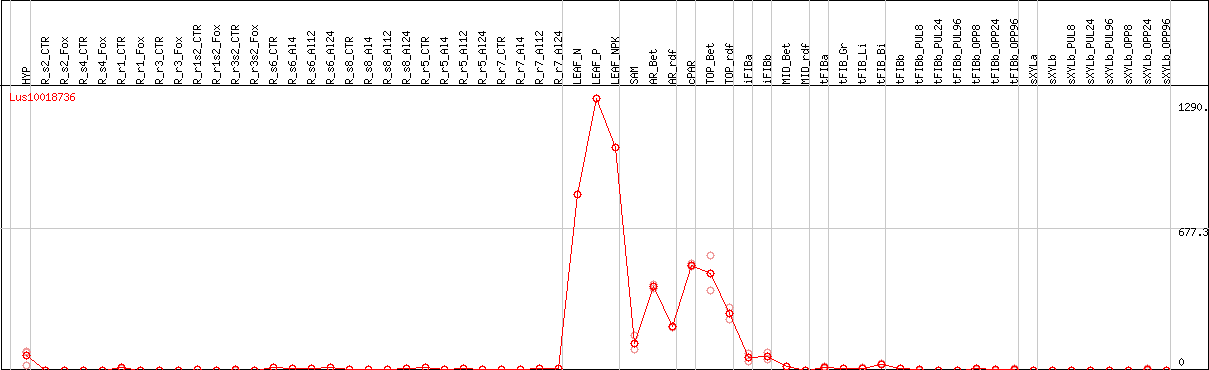
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018736 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.