External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G38870 90 / 1e-25
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
AT5G43580 76 / 1e-19
UPI
UNUSUAL SERINE PROTEASE INHIBITOR, Serine protease inhibitor, potato inhibitor I-type family protein (.1)
AT2G38900 75 / 1e-19
Serine protease inhibitor, potato inhibitor I-type family protein (.1.2)
AT3G46860 57 / 2e-12
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
AT5G43570 49 / 2e-09
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018783
108 / 4e-33
AT2G38870 89 / 2e-25
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Lus10018786
99 / 3e-29
AT2G38870 93 / 6e-27
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Lus10024870
96 / 4e-28
AT2G38870 94 / 2e-27
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Lus10014740
83 / 6e-23
AT2G38870 77 / 2e-20
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Lus10002732
76 / 4e-20
AT2G38870 76 / 4e-20
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Lus10032655
57 / 2e-11
AT1G52870 499 / 3e-176
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10043097
56 / 6e-11
AT1G52870 505 / 2e-179
Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.1), Peroxisomal membrane 22 kDa (Mpv17/PMP22) family protein (.2)
Lus10024370
46 / 4e-08
AT2G38870 47 / 3e-08
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Lus10010859
46 / 5e-08
AT2G38900 47 / 3e-08
Serine protease inhibitor, potato inhibitor I-type family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G075600
95 / 1e-27
AT5G43580 80 / 3e-21
UNUSUAL SERINE PROTEASE INHIBITOR, Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.010G075200
91 / 5e-26
AT2G38870 76 / 4e-20
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.006G212000
91 / 6e-26
AT2G38870 81 / 5e-22
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.010G075400
90 / 1e-25
AT2G38870 84 / 3e-23
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.016G078800
88 / 8e-25
AT2G38870 79 / 3e-21
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.016G079050
86 / 4e-24
AT2G38870 81 / 2e-22
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.006G212200
84 / 1e-23
AT2G38870 81 / 6e-22
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.016G078900
84 / 2e-23
AT2G38870 84 / 2e-23
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.009G028300
84 / 3e-23
AT5G43580 76 / 7e-20
UNUSUAL SERINE PROTEASE INHIBITOR, Serine protease inhibitor, potato inhibitor I-type family protein (.1)
Potri.010G075800
84 / 3e-23
AT2G38870 76 / 3e-20
Serine protease inhibitor, potato inhibitor I-type family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0367
CI-2
PF00280
potato_inhibit
Potato inhibitor I family
Representative CDS sequence
>Lus10018784 pacid=23176292 polypeptide=Lus10018784 locus=Lus10018784.g ID=Lus10018784.BGIv1.0 annot-version=v1.0
ATGTCGACTGACTGTTCAGGTAAGAACTCATGGCCGGAGCTGTTGGCGACCAACGGGGACGCGGCGGCGGCAGTCGTTGAGAGGGAGAACGACAACGTTG
ATGCGATCGTGGTGCTGGAAGGGACGATCGTTCCCCTAGATTTCAGGTGTAACCGGGTTTGGGTTTGGGTTGATGCCAATAGGGTTGTCACCCGTGTTCC
TACCGTTGGTTGA
AA sequence
>Lus10018784 pacid=23176292 polypeptide=Lus10018784 locus=Lus10018784.g ID=Lus10018784.BGIv1.0 annot-version=v1.0
MSTDCSGKNSWPELLATNGDAAAAVVERENDNVDAIVVLEGTIVPLDFRCNRVWVWVDANRVVTRVPTVG
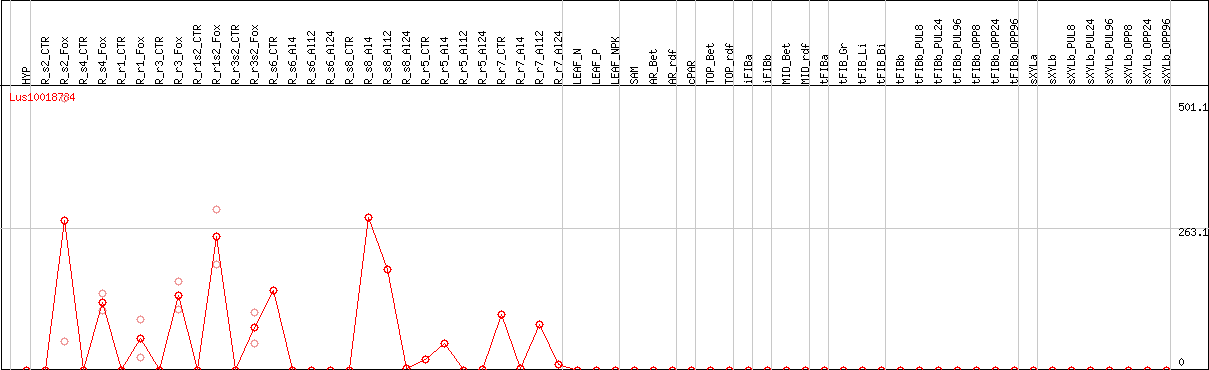
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018784 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.