External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G23340 329 / 3e-112
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT5G51310 328 / 5e-112
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G80330 125 / 3e-33
ATGA3OX4
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 4, gibberellin 3-oxidase 4 (.1)
AT3G13610 124 / 1e-32
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G15550 121 / 2e-31
ATGA3OX1, GA4
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
AT1G55290 120 / 5e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52820 118 / 1e-30
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G19010 118 / 2e-30
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.3)
AT1G06620 117 / 5e-30
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G60980 116 / 2e-29
ATGA20OX4
gibberellin 20-oxidase 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009663
326 / 3e-111
AT4G23340 358 / 6e-124
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10017799
168 / 3e-52
AT5G51310 108 / 1e-29
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10009025
154 / 8e-45
AT5G51310 131 / 9e-38
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10006759
125 / 5e-33
AT4G21200 419 / 2e-147
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10011476
124 / 2e-32
AT1G15550 412 / 1e-143
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Lus10021002
124 / 2e-32
AT3G19000 389 / 3e-135
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10015016
124 / 3e-32
AT4G25420 494 / 3e-176
GA REQUIRING 5, ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10007640
123 / 3e-32
AT4G21200 429 / 1e-151
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10031671
124 / 4e-32
AT5G07200 508 / 1e-180
ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 3, gibberellin 20-oxidase 3 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G159500
391 / 8e-137
AT4G23340 402 / 3e-141
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.003G128100
368 / 1e-127
AT4G23340 458 / 2e-163
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.003G106900
365 / 2e-126
AT4G23340 392 / 1e-137
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.011G026700
122 / 3e-32
AT4G21200 444 / 1e-157
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.005G184400
122 / 8e-32
AT5G51810 380 / 5e-131
gibberellin 20 oxidase 2 (.1)
Potri.005G184200
121 / 1e-31
AT5G51810 383 / 1e-132
gibberellin 20 oxidase 2 (.1)
Potri.018G033600
121 / 1e-31
AT1G15550 328 / 2e-111
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.001G176600
120 / 3e-31
AT1G15550 452 / 1e-159
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.006G247700
120 / 4e-31
AT1G15550 328 / 2e-111
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
Potri.004G146100
120 / 5e-31
AT3G19000 374 / 5e-129
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10018836 pacid=23144689 polypeptide=Lus10018836 locus=Lus10018836.g ID=Lus10018836.BGIv1.0 annot-version=v1.0
ATGGTATTAGGAGAAGGAAGCAGTAGCAGTAGCAGTACTAGTACAGTCCAAGTACTCCCAGTGGTGGACATATCTCAGCCGTTGAATCCATCATCACTGT
TATCTCTAGCTGATGCCTGCCAGAAATGGGGATTCTTCCATATCATCAACCATGGAATCCCCAGGAATCTTTACGACAGAATCTATTCCTTATCTCAGCA
TTTCTTCAGCTTGCCTTGTCATTCGAAACTTGAACACTTGGGTCTCCAAACTTACACTCCAAACTTCATAGCCTCCCCTTTCTTCGAGAGCCTCCGTGTG
TCTGGTCCGGACTTCTTCTCCTCTGCTCACTCCTCTGCCTCTCCCCTGCTCACTCCACCCATCTCTTCTGAATTCAGTGAGTTGGTTGAAGAGTACGGAA
GCAAGATGACAGAGCTATCAAAACGAATCATGGCGATCCTGCTAACAATCCTCGGAAAAGGGTTCGACGAGAAGTTCCACGAATCCGAATTCGACAAGTG
CCATGGATACCTGAGGGTGATCAACTACACCGTTCCTGCCGAAACTGACCCCGTTAACGAAGTAGAGGGGCTGGGGATGCACACGGATATGAGCTGCGTG
ACGATTGTGTACCAAGACCAAATTGGTGGCCTCCAAGTTAAGTCGGAATCACTAGGCGGTGGTACGAAATGGATGGACATAAGACCTTCCGAGGAGACGC
TTGTGGTGAACATTGGGGACATGTTGCAGGCTTGGAGCAACGACAAGTTCCGGTCGTCAGAGCACCGGGTGATGCTGTGCGGACGGCCACGGTTGTCTCT
TGCCTTCTTCTGGTGTTTTGAGGATGAGAAAGTGATTCGGGCACCTGATGAAGTTGTGGGAGATGAGGGAAGAAGGGCTTATAGGTCATTTGTCTGCTCT
GAATATCTGAGGTTTAGGGAGATCAAGAGTGAGAGAGGAAGGTTTGAGAAAGTTGGGTATACTGTCAGAGATTTTGCTAATGCTGATCTACTCACTGATT
CTTCATAA
AA sequence
>Lus10018836 pacid=23144689 polypeptide=Lus10018836 locus=Lus10018836.g ID=Lus10018836.BGIv1.0 annot-version=v1.0
MVLGEGSSSSSSTSTVQVLPVVDISQPLNPSSLLSLADACQKWGFFHIINHGIPRNLYDRIYSLSQHFFSLPCHSKLEHLGLQTYTPNFIASPFFESLRV
SGPDFFSSAHSSASPLLTPPISSEFSELVEEYGSKMTELSKRIMAILLTILGKGFDEKFHESEFDKCHGYLRVINYTVPAETDPVNEVEGLGMHTDMSCV
TIVYQDQIGGLQVKSESLGGGTKWMDIRPSEETLVVNIGDMLQAWSNDKFRSSEHRVMLCGRPRLSLAFFWCFEDEKVIRAPDEVVGDEGRRAYRSFVCS
EYLRFREIKSERGRFEKVGYTVRDFANADLLTDSS
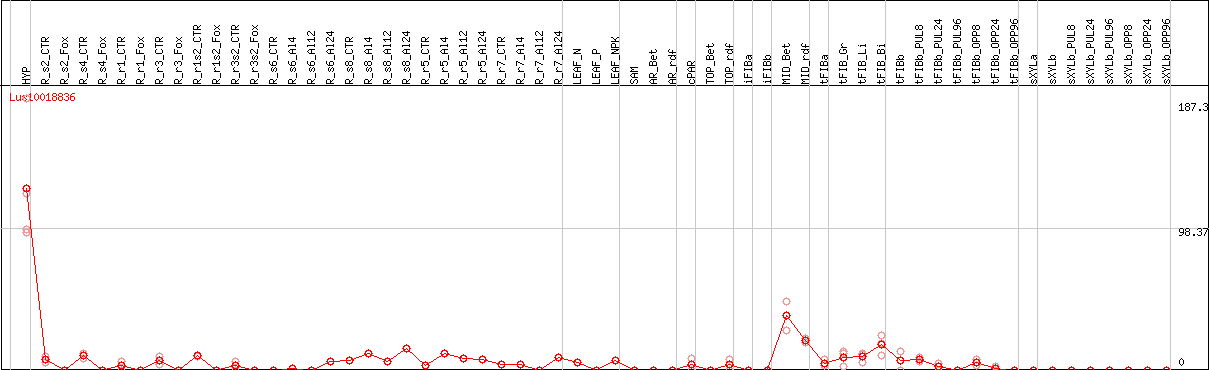
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10018836 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.