Lus10019014 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10019014 pacid=23152652 polypeptide=Lus10019014 locus=Lus10019014.g ID=Lus10019014.BGIv1.0 annot-version=v1.0
ATGAATTATCCACCAGAGAAGTTTGTGAAGTTTCAGGATTGGAGATCAGACAAAGTCTCCGAATCAGATTATTCTTCCACGAATGGGAGGTTTAGAACGA
AGATAAACTCCTTCTCTGATAAGTTCAGGAGGAGGTTGGAATCTGGTTCAGAAAGCATCAGTAGGGTTAGAAAGTCGTGGAAATCATACTCCTTCAATGG
TGAAAGTACTAGTACTAGCAGTGGTATAGATGGTTCTTGTTCAAGCAAGAAGATCCTTGACCCTCAGGGAGCTTTCCTTCAGAAATGGAACAAGATATTT
GTGTTATCCTGTTTCATTGCTGTCTCCTTGGACCCATTGTTCTTCTACGTCCCAGTAATCGACGACAAAAACAAGTGCCTTGACCTGGACGATAAGATGC
AGATAACTGCCAGCGTACTTAGGTCCTTCACCGATATATTCTACATACTCCACATTGTTTTCCAGTTCCGCACCGGCTTCATTGCTCCTTCTTCCCGTGT
TTTTGGGAGAGGTGTCTTGGTTCAGGACAGTTGGGCGATTGCGAAAAGATACATGCTTTCGTACTTCTTGATCGACGTCCTTGCGTTGCTGCCATTGCCA
CAGATGGTGATTCTAATCATCATTCCAAGGCTGCGAGGGTCGAGAACACTCAACACCAAAAACCTGTTGAAGTTTGTTGTAGTCCTTCAGTATCTGCCAA
GGATTATTCGGGTTTATCCATTGTATATGGAAGTCACAAAGACCTCCGGAATCCTCACCGAAACTGCTTGGGCCGGAGCTGCATTCAATCTGTTCCTTTA
CATGCTTGCTAGTCACGTGCTGGGAGCATTTTGGTACTTGTTCTCTGTTGAAAGACTGACAGTATGCTGGAGGAAAGCTTGCGACAAAATCTCATGTGAT
CGGGCTATACTGTATTGTGATGGTCCTGGAGGCGACAAGAACTTCTTCAACGCTTCTTGTCCTATTTCTCCAGAGAATGCAGATGTATTCAATTTCGGTA
TATTCCTCGATGCTCTTCAGTCTGGAGTTGTGCAATCTCATGACTTCCCGAAGAAACTCTTTTACTGTTTCTGGTGGGGGCTTCGCAATCTTAGTTCTCT
TGGTCAGAATCTTGCAACTAGCACCTTTGTCTGGGAAATCTGTTTCTCCGTGGCCATTTCTATATTCGGACTAGTGTTGTTCTCGTTCCTCATCGGGAAT
ATGCAGACATACTTGCAATCCACAACTACTAGATTGGAAGAAATGAGAGTGAAGAGGCGAGACGCGGAGCAATGGATGTCCCATCGGATGCTTCCGGATA
ATCTGAGGGAGCGGATCAGGCGACACGAGCAGTACAAGTGGCAAGAAACTAGAGGAGTTGACGAAGAGAAGTTGGTACACAATCTTCCCAAGGACATTCG
GAGGGACATCAAACGTCATCTCTGCTTAGCGCTTCTTATGAGAGTGCCGATGTTCGAAAAGATGGACGAACAGCTGCTGGACGCAATGTGTGACCGTCTC
AAACCAGCGCTATACACGGAAGAAAGCTACATCGTTCGTGAGGGAGATCCAGTCGACGAGATGCTCTTCGTTATGCGAGGAAAGCTCCTAACAATGACCA
CGAACGGAGGAAGGACAGGGTTTTTCAACTCGGAGTATCTCAAGGCAGGCGATTTCTGCGGTGAGGAGCTGCTGACATGGGCGCTGGATCCCCACACTTC
GTCGAACCTTCCCATCTCGACCAGAACGGTTCGTACGATAACCGAAGTGGAAGCGTTTGCACTGACGGCCGAGGACTTGAAGTTCGTGGCTTCGCAGTTC
CGCCGGCTTCACAGCAAGCAGATACGCCACACGTTCAGATTCTATTCGCAGCAGTGGCGGACTTGGGCCGCTTGCTTCATACAAGCTGCTTGGCGACGGT
ACAGTAAGAAGAAGCTGGAGGAATCGTTGAGGCAAGAAGAGGACAGGCTGCAGGATGCGTTGGCTAAGGCGAGCGGGAACTCTCCTAGTCTCGGTGCGAC
TATATACGCGTCGAGGTTTGCTGCCAATGCGTTGAGGCTACTGAGGAGGAACTCGTCGAGAAAGATGCGGGTCCCGGAGAGGGTTCCTCCTATGTTGTTG
CAGAAGCCGGCTGAGCCAGATTTTACTGCCGATGAACGGACCTTCGACCCGAGGGAATCGCTGGTTCGGAGGTGGCACTTGGTATTGCTGGTAACAAGCG
GAGCTGCTGTGCTGATAGATCCTCTCTTTCTTTACATCCTACGAATCAACAACCACCGCACTAGTACTACTAAGAACGAGAAATTTATGATAGGGTTCGA
CAAAACTTTGCTGATTTCTGCAACCACGCTCCGCTCCACTGCTGACGTTTTCTGTGGCCTGCTGCACGTCTTTTTCAAATTCCGCACAGCTTTCCCTGCT
CCTTCTTCAAGAGTGTTCAGCCGTGTTGAACTAGTTCACGACTCATCTGCAATTGCTCGCAGATACTTCCCTTCCTTTCTCCTCGTTGACTTCCTTGCCC
TCCTTCCCCTGCCTCAGGTGGTTGTAGTTGTAGGTTTGATTTCCAGAGAGGCCATGAGAACCAAACTGTTGACACCCACTATAACTTCGCTGCACCTGGT
GCTTTCTTTGCAATTGGTGCCAAGGCTGCTGAGATTCTACCAGCTATACCAAGATGTGATCAAGTCTTCATACGCAATCCGAAAAGCTGCACTTGTGAGG
ACTGCCTTCACCGTCTTTCTTTACATCACTGCTGCTCATGCAGTGGGAGCATTCTGGTACTCATCTTCCATACGCAGAGAAATGGATTGCTGGCAATTAG
CCTGCAAAAATCATGCTAGATGTGATTACATTGCACTGTTGACCTTTGGCAATGACAGACCGCCTCCTCAGATGGATTACTCGTCATTTCTGAATCGGAC
TTGTCCCAAAGGAGGTGCGGAATCGAGTTTCGATTATGGGATGTTTGCTGATGCTGTTAAGTCTGGTGTGCTGGAATCTGCAAATTTCGACGAGAAGTTG
CTCTACAGCTTTTGGTGGGGTCTGCAAAGTCTGAGCACCTTGGGTCAGAACGTAAAACCAAGCATCAACAGTCTGGAGATATGTTTTGCACTGTGCATTT
TCATTGCTGGGTTGCTGTTATTTTCGCAAGTCATCGGTATCATACAGACAAGGATGCTGACATCGACGATTAGGATTGAGGAGATGAAAATGAAATGGGA
AGATGTGGAGGGATGGATGTCTTCCCATTTCATCCCACACGAACTAAAGAGGCGCCTGAGGCGACATCAAGAGTACAGATGGCGTACGGACCAAGGAGTC
GATTTCAACCACGTACTAGATAATGTTCCAAAGGACCTCCGAAGGGATGTCAAGCGCCATCTTTGGCTGCCGTTGCTCATGAGAGTGCCAATGTTCGAAG
AGATGGATGAACAGCTACTAGATGCAATGTGTGACCGTCTCAAACCAGCACTATACATAGAGGAAAGCCACATTGTTCGTGAGGGAGATCCAGTGGATGA
GATGCTCTTCGTTATGCGAGGAAAGCTCCTGACAACAACCACGGACGGAGGAAGGACAGGGTTTTTCAACTCGGTGTACCTCAAGGCAGGCGATTTCTGC
GGCGAGGAGCTTCTGACATGGGCGCTGGATCCTCACTCTTCGTCGTACCTCCCTAACTCGACCAGAACAGTTCGTACGATAACCGAAGTAGAAGCATTTG
CACTAACAGCGGAGGACTTGAAGTTCGTGGCTTTGCAGTTTCGTAGGCTTCACAGCAAGCAGATACAGCACACGTTCAGATTCTATTCGCAACAGTGGCG
GACTTGGGCTGCTTGTTTCATACAAGCTGCCTGGCGACGATACAGAAAGAAGAAGCTTGAAGAAGCAATGAGGCAAAAAGAGGACAGGCTACAGGATGCA
TTGGCTAAGGCGAGCAGTAGCGGGAACTCTCCTAGCCTTGGTGCAACAAAATACGCATCGAGGGTTGGTGTCAATAGGCAGGCGATTTCTGCGGCGAGGA
GCTTCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10019014 pacid=23152652 polypeptide=Lus10019014 locus=Lus10019014.g ID=Lus10019014.BGIv1.0 annot-version=v1.0
MNYPPEKFVKFQDWRSDKVSESDYSSTNGRFRTKINSFSDKFRRRLESGSESISRVRKSWKSYSFNGESTSTSSGIDGSCSSKKILDPQGAFLQKWNKIF
VLSCFIAVSLDPLFFYVPVIDDKNKCLDLDDKMQITASVLRSFTDIFYILHIVFQFRTGFIAPSSRVFGRGVLVQDSWAIAKRYMLSYFLIDVLALLPLP
QMVILIIIPRLRGSRTLNTKNLLKFVVVLQYLPRIIRVYPLYMEVTKTSGILTETAWAGAAFNLFLYMLASHVLGAFWYLFSVERLTVCWRKACDKISCD
RAILYCDGPGGDKNFFNASCPISPENADVFNFGIFLDALQSGVVQSHDFPKKLFYCFWWGLRNLSSLGQNLATSTFVWEICFSVAISIFGLVLFSFLIGN
MQTYLQSTTTRLEEMRVKRRDAEQWMSHRMLPDNLRERIRRHEQYKWQETRGVDEEKLVHNLPKDIRRDIKRHLCLALLMRVPMFEKMDEQLLDAMCDRL
KPALYTEESYIVREGDPVDEMLFVMRGKLLTMTTNGGRTGFFNSEYLKAGDFCGEELLTWALDPHTSSNLPISTRTVRTITEVEAFALTAEDLKFVASQF
RRLHSKQIRHTFRFYSQQWRTWAACFIQAAWRRYSKKKLEESLRQEEDRLQDALAKASGNSPSLGATIYASRFAANALRLLRRNSSRKMRVPERVPPMLL
QKPAEPDFTADERTFDPRESLVRRWHLVLLVTSGAAVLIDPLFLYILRINNHRTSTTKNEKFMIGFDKTLLISATTLRSTADVFCGLLHVFFKFRTAFPA
PSSRVFSRVELVHDSSAIARRYFPSFLLVDFLALLPLPQVVVVVGLISREAMRTKLLTPTITSLHLVLSLQLVPRLLRFYQLYQDVIKSSYAIRKAALVR
TAFTVFLYITAAHAVGAFWYSSSIRREMDCWQLACKNHARCDYIALLTFGNDRPPPQMDYSSFLNRTCPKGGAESSFDYGMFADAVKSGVLESANFDEKL
LYSFWWGLQSLSTLGQNVKPSINSLEICFALCIFIAGLLLFSQVIGIIQTRMLTSTIRIEEMKMKWEDVEGWMSSHFIPHELKRRLRRHQEYRWRTDQGV
DFNHVLDNVPKDLRRDVKRHLWLPLLMRVPMFEEMDEQLLDAMCDRLKPALYIEESHIVREGDPVDEMLFVMRGKLLTTTTDGGRTGFFNSVYLKAGDFC
GEELLTWALDPHSSSYLPNSTRTVRTITEVEAFALTAEDLKFVALQFRRLHSKQIQHTFRFYSQQWRTWAACFIQAAWRRYRKKKLEEAMRQKEDRLQDA
LAKASSSGNSPSLGATKYASRVGVNRQAISAARSF
|
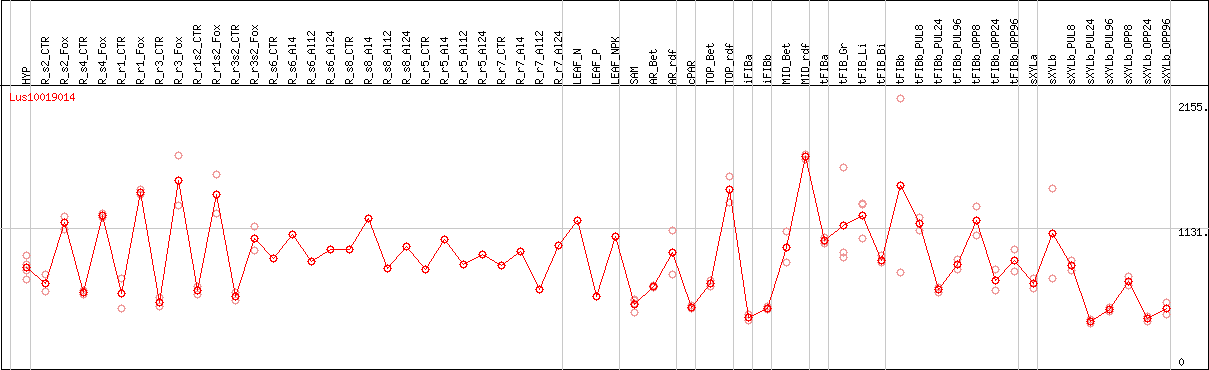
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019014 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
