External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G61400 159 / 5e-45
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G38730 75 / 3e-15
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G64320 73 / 2e-14
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G39710 71 / 9e-14
EMB2745
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G55840 68 / 6e-13
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G48810 68 / 7e-13
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G05670 67 / 2e-12
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
AT1G09680 66 / 4e-12
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G20090 64 / 1e-11
EMB1025
embryo defective 1025, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G61990 64 / 1e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015717
404 / 1e-144
AT5G61400 151 / 4e-42
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10008185
78 / 4e-16
AT1G19290 864 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10027974
76 / 2e-15
AT1G19290 480 / 1e-156
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10013077
74 / 7e-15
AT5G40400 587 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10032174
70 / 1e-13
AT5G38730 746 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10034421
69 / 3e-13
AT2G02150 706 / 0.0
EMBRYO DEFECTIVE 2794, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10038804
68 / 6e-13
AT5G64320 367 / 3e-115
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10039056
68 / 7e-13
AT5G64320 832 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10026465
67 / 2e-12
AT1G05670 197 / 6e-55
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G015900
73 / 1e-14
AT5G64320 878 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.014G040200
72 / 5e-14
AT1G19290 883 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.017G071600
69 / 2e-13
AT5G40400 634 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.001G075900
67 / 9e-13
AT4G20090 806 / 0.0
embryo defective 1025, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.004G074500
67 / 9e-13
AT1G62930 478 / 1e-162
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G035700
67 / 2e-12
AT1G05670 922 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Potri.003G204100
66 / 4e-12
AT1G07740 502 / 2e-176
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G368700
65 / 7e-12
AT5G55840 1272 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G234500
64 / 1e-11
AT1G74580 1034 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G154800
64 / 1e-11
AT4G20090 816 / 0.0
embryo defective 1025, Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10019080 pacid=23141818 polypeptide=Lus10019080 locus=Lus10019080.g ID=Lus10019080.BGIv1.0 annot-version=v1.0
ATGTATAAGCCGCTCATTCCAAAATGCTTTACGATCGTCAACGCTCCCCAAAACCCTCGTCTCTCCTTATTCCTATTCTCTTCTTCTTACTCCTCTCCTC
CCTCTTCGCCGCCTTCTGATCTCACATCCGCCATTCTCAACTCCAGAACCCCTGACCAAGCCCTCCAACTCTTCACCTCCGTCGTAAATCGCAACTCCAA
ATCCCCAGTGAAAAACCTTCCACATTTCTCCGCCGTGATTCACGTGTTAACCGGTGCTGGTGCGTACACGCCTGCCAGGTGCTTGATAAAAGGCCTGATC
CAGTCCTTCCTGAAATCCCACGACCCAAAACGTGCTAGCTCGTTGATTTTCGACGCTCTTAGTCGGCTGAAGAGCTCCAAATTCACTTCCGATGTGTTCG
GGACGTTGATTATTGCACTCTGTGAGGTGGGACTTGTTGATGAAGCCCTGTGGGTGCACCGCAAGATTGGTACTTTACCAGCGAGGCAAGCTTGTAATGC
CCTTATGAATGGGCTAGTTAAGAAAGGGAGATTCGAATCCATGTGGGAAACTTATGAAGAAATGATTTCCGGTGGGTTGGTGCCTAGTGTTGTCAGTTAT
GGCGTACTGATCAATGCTTGTTGCTGCCAAGGTGATCTAATAAAGGCGGCGGCTTTGATTAGCGAGATTATGGAGAAAGGGATTCAGCCCCACCGTTGTG
ATCTACACAACTCTCATTCGTGGGTTTTGCTGTGA
AA sequence
>Lus10019080 pacid=23141818 polypeptide=Lus10019080 locus=Lus10019080.g ID=Lus10019080.BGIv1.0 annot-version=v1.0
MYKPLIPKCFTIVNAPQNPRLSLFLFSSSYSSPPSSPPSDLTSAILNSRTPDQALQLFTSVVNRNSKSPVKNLPHFSAVIHVLTGAGAYTPARCLIKGLI
QSFLKSHDPKRASSLIFDALSRLKSSKFTSDVFGTLIIALCEVGLVDEALWVHRKIGTLPARQACNALMNGLVKKGRFESMWETYEEMISGGLVPSVVSY
GVLINACCCQGDLIKAAALISEIMEKGIQPHRCDLHNSHSWVLL
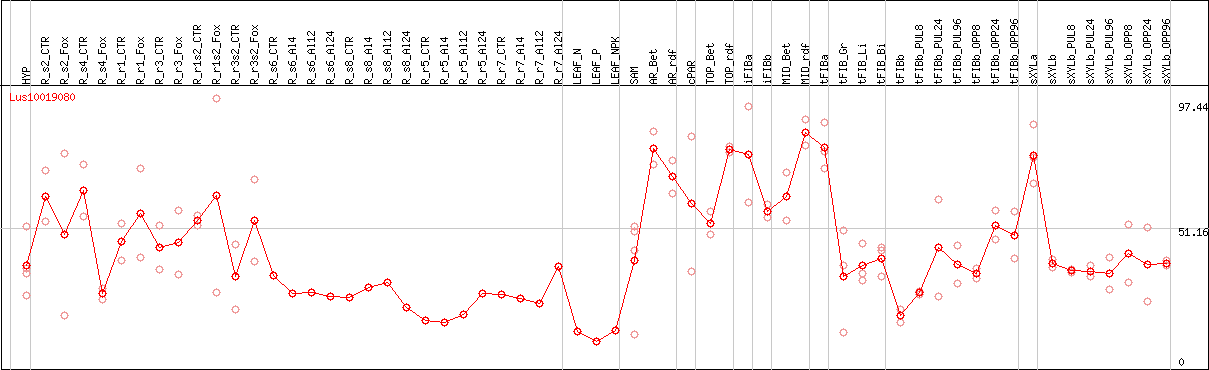
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019080 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.