External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G26550 219 / 1e-74
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
AT2G18040 62 / 2e-13
PIN1AT
"peptidylprolyl cis/trans isomerase, NIMA-interacting 1", peptidylprolyl cis/trans isomerase, NIMA-interacting 1 (.1)
AT5G19370 48 / 4e-07
rhodanese-like domain-containing protein / PPIC-type PPIASE domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015714
228 / 6e-78
AT1G26550 224 / 1e-76
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Lus10014266
61 / 8e-13
AT2G18040 199 / 9e-68
"peptidylprolyl cis/trans isomerase, NIMA-interacting 1", peptidylprolyl cis/trans isomerase, NIMA-interacting 1 (.1)
Lus10025967
62 / 2e-12
AT2G18040 200 / 8e-67
"peptidylprolyl cis/trans isomerase, NIMA-interacting 1", peptidylprolyl cis/trans isomerase, NIMA-interacting 1 (.1)
Lus10000413
50 / 5e-08
AT5G19370 240 / 1e-77
rhodanese-like domain-containing protein / PPIC-type PPIASE domain-containing protein (.1)
Lus10026105
50 / 5e-08
AT5G19370 233 / 8e-76
rhodanese-like domain-containing protein / PPIC-type PPIASE domain-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G089900
212 / 6e-72
AT1G26550 214 / 1e-72
FKBP-like peptidyl-prolyl cis-trans isomerase family protein (.1)
Potri.007G013400
66 / 1e-14
AT2G18040 198 / 4e-67
"peptidylprolyl cis/trans isomerase, NIMA-interacting 1", peptidylprolyl cis/trans isomerase, NIMA-interacting 1 (.1)
Potri.009G069300
50 / 1e-07
AT5G19370 281 / 3e-94
rhodanese-like domain-containing protein / PPIC-type PPIASE domain-containing protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0487
FKBP
PF00639
Rotamase
PPIC-type PPIASE domain
Representative CDS sequence
>Lus10019084 pacid=23141742 polypeptide=Lus10019084 locus=Lus10019084.g ID=Lus10019084.BGIv1.0 annot-version=v1.0
ATGGGGAAGAAAGATGGGGCAAAAGGCAAAGGTAAGGCAGCTAGTAGCAGCGATGAAAATGCTTCAAAGGGTAAGGGAAAAGCTGGAAAGGCCGATGGAC
TTGGCACATGCACATATGTAAAAGCAAGGCATATTTTATGTGAGAAGCAAGGTAAGATTAACGAGGCATACAAGAAACTGCAAGACGGTTGGCTAAGCAA
TGGAGATAAAGTTCCCCCCGCTGAGTTTGCGAAGGTAGCGACCGAGTATTCAGAATGCCCTTCGGGAAAGAAGGGAGGAGACCTTGGATGGTTTCCGCGT
GGGAAGATGGCCGGTCCGTTCCAGGAGGTTGCTTTCAACACCCCTGTTGGAGCTACTAGTGCACCGTTTAAGTCAACGCATGGGTATCACATAATCCTAT
CTGAAGGCAGAAAGAATTGA
AA sequence
>Lus10019084 pacid=23141742 polypeptide=Lus10019084 locus=Lus10019084.g ID=Lus10019084.BGIv1.0 annot-version=v1.0
MGKKDGAKGKGKAASSSDENASKGKGKAGKADGLGTCTYVKARHILCEKQGKINEAYKKLQDGWLSNGDKVPPAEFAKVATEYSECPSGKKGGDLGWFPR
GKMAGPFQEVAFNTPVGATSAPFKSTHGYHIILSEGRKN
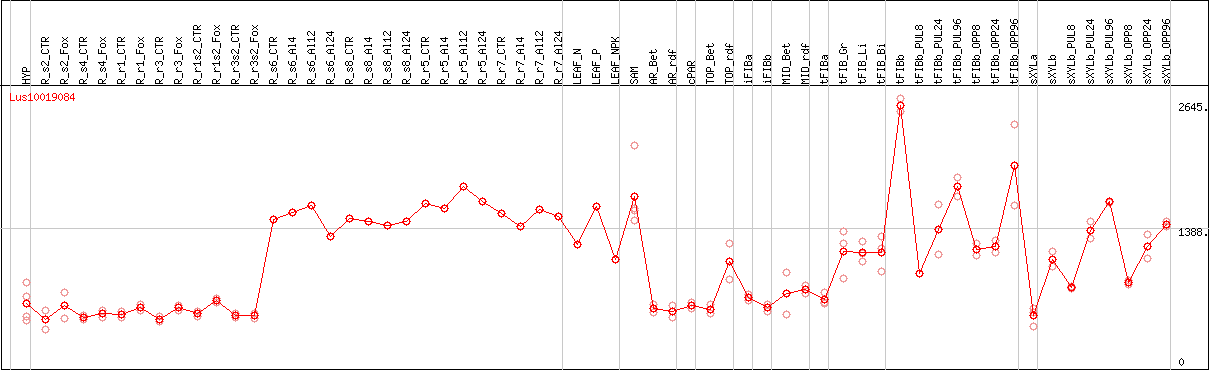
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019084 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.