External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69080 192 / 2e-61
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT2G03720 184 / 3e-59
MRH6
morphogenesis of root hair 6, Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT5G17390 162 / 2e-49
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT3G03290 155 / 1e-46
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT1G44760 110 / 7e-30
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT4G13450 80 / 4e-18
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT1G48960 60 / 8e-11
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT1G11360 49 / 6e-07
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
AT2G21620 48 / 7e-07
RD2
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT1G68300 48 / 8e-07
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036814
181 / 5e-59
AT1G69080 109 / 7e-32
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10034661
171 / 4e-53
AT5G17390 160 / 3e-48
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10007982
163 / 9e-50
AT5G17390 149 / 1e-43
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10025644
106 / 1e-28
AT1G44760 211 / 1e-69
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10018190
100 / 3e-26
AT1G44760 197 / 1e-64
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10029664
99 / 1e-25
AT1G44760 236 / 2e-79
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10042701
99 / 2e-25
AT1G44760 235 / 6e-79
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10014459
81 / 1e-18
AT4G13450 195 / 8e-63
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10023716
59 / 3e-11
AT4G13450 136 / 2e-41
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G140200
277 / 7e-95
AT1G69080 202 / 3e-65
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.008G109000
268 / 4e-91
AT1G69080 196 / 6e-63
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.T170801
267 / 7e-91
AT1G69080 196 / 3e-63
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.015G060700
201 / 6e-65
AT3G03290 198 / 2e-63
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.002G084600
120 / 1e-33
AT1G44760 200 / 5e-65
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.005G177100
114 / 3e-31
AT1G44760 226 / 3e-75
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.010G064800
89 / 1e-21
AT4G13450 194 / 4e-63
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.012G059100
51 / 1e-07
AT1G48960 175 / 3e-55
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.010G123200
50 / 2e-07
AT1G68300 173 / 5e-56
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.011G039800
50 / 3e-07
AT1G11360 269 / 3e-91
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0039
HUP
PF00582
Usp
Universal stress protein family
Representative CDS sequence
>Lus10019173 pacid=23141769 polypeptide=Lus10019173 locus=Lus10019173.g ID=Lus10019173.BGIv1.0 annot-version=v1.0
ATGGGTGCAAAGACAGCAGCAGCAGGGGGGTTCTGCCTCAACCGCATCCGCCCACACGTCAGAGTAGTCCGGTCCCCGCAGCAACCTGACCGGCCGAAAC
TAGATGATCATGACGGCGGAGGTAAAACTACGCCGACTGAGGCGGTGGCGATCGGAGGAGGAGGAAGGAAGGTGATGATAGTGGTGGATTCGAGCGCGGA
GGCTAAGGTTGCTCTGCAGTGGGCGCTTTCCCATACTGTTCAAGTGCAGGACACTCTGCTTCTTCTCCATGTTGCCAAGCCATCTAACAACCATGGCCAC
CAAGGGTTGATGAAAGGAAGAGGTTCAAGAGGTTATGAGCTGGCCAACTCTCTAAAGAAAATGAGCCAGATGAAGAGGCCAGAGGTGGAGATAGAGACAG
TAATAGTGGAAGGGAAGGAGAAAGGTCCAACAATAGTTGAAGAAGCAAAGAAGCAAGGCGTAGCTCTGCTGGTACTTGGCCAGAGGAAGAGATCTTCGAT
GACGTGGCGGCTGATCATGATGTGGGCCAGTAGTAAAGTCAATGCAGCCGGCGGCGGCGGGGTGGTGGAGTATTGTATCCAGAATGCAGAGTGCATGGCG
ATCGCCGTCCGGCGGAAGAGCAAGAAACATGGCGGCTACTTGATCACCACTAAACGCCATAAGGATTTCTGGCTCTTAGCCTAG
AA sequence
>Lus10019173 pacid=23141769 polypeptide=Lus10019173 locus=Lus10019173.g ID=Lus10019173.BGIv1.0 annot-version=v1.0
MGAKTAAAGGFCLNRIRPHVRVVRSPQQPDRPKLDDHDGGGKTTPTEAVAIGGGGRKVMIVVDSSAEAKVALQWALSHTVQVQDTLLLLHVAKPSNNHGH
QGLMKGRGSRGYELANSLKKMSQMKRPEVEIETVIVEGKEKGPTIVEEAKKQGVALLVLGQRKRSSMTWRLIMMWASSKVNAAGGGGVVEYCIQNAECMA
IAVRRKSKKHGGYLITTKRHKDFWLLA
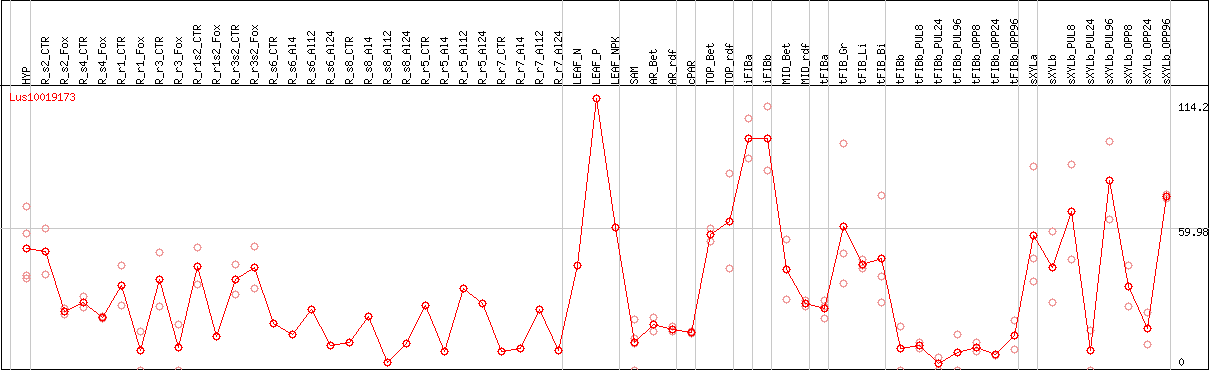
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019173 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.