Lus10019220 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10019220 pacid=23141417 polypeptide=Lus10019220 locus=Lus10019220.g ID=Lus10019220.BGIv1.0 annot-version=v1.0
ATGGGCTCTGCATCAAAGTTGATCTCGACAGCGCCTGATCCGGAAGCGGAGGATCCGTCATTGGCCGAATCAGGGAAGGGGGCAATTCCTTCGGATTCCA
AATCTAGAAAGGCAGTGGCAAGCACTAAAGGAAAGAAGAAGCGTGAAACTGCAAAGAGCATTGAACCTCGTGCTTCACCACCGAAGCGGCCGAGAAGGGC
TGCTGCATGTGTGAATCTGAAGGAGAAATCCGTGAAGCTGACTGATGAGTACACAGTTATCGAACCGAAGAAGGAGCAAGTTGTTCAAGATGAACTTCTC
GCTGTTCGCTTGACTGAGGGACAAGGAGAGGACGGTCGTCCGAATAGAAGACTGACTGATTTCATCGTCCATGATGCAGATTTCGCTCCACAGTATCTGG
AGATGATTGAAGTTAGTGATATGTTTATCTCTGGTGTCATATTACCGATCGATGATTGTATGGGGGATGACAAAGAGAAGCGGGTGAGGTGCGGCTACCG
GAAGTTTTATGATCGGTTTTTCGAGAAAGCTCGAGCTTGTATAGAAGTGTACAAGAAGCTGTCGAAATTTGCTGGTGGGAATCCTGACCTCAGCCTTGAT
GAGTTGCTTGCTGGGCTTGTTCGTTCTTTTGGCAGTAGCAAGTGCTTTTCTGGGGCATCATCTATTAAAGAGTTTCTTATCTCACAAGGTGAGTTCATAT
ACGATCAACTGAAAGGTTTAGATGAGACATCGAAGCAGAATGATATGAAGTTCTCAGAGCTGCCTGTGCTTTTGGCTTTAAGAGATAAGAGCAAGAAACA
TGAGAGTTTCATACTGGAAGGAGCAGTTGGCTATGGTGGTAGTGGGACGTTGGTTATTGAACCAAATGCTGATAAGAAAGACACAAAGATGGATGAAGTA
AATGGATCTGCTGGTAATGATGAGGACGAGGATGCAAAATTAGCCCGGATGCTGCAGCAGGAAGAATTATGGGCATCCATGAAGCAGAAGAAAAGAAAAG
GCTCATCCATGTCGAATGCTATCTACATTAAGGTCAATGAAAGAGATATTGCGAATGATTATCCTCTCCCTGCATACTACAAGCAAACCGTTGAAGAAAC
TGATGAGTATATAGCCGTTGACTGTGATGATGTGATCAACCCTGATGACCTGCCAAGAAAAGTGCTTCATAATTGGTCACTTTATGATTCAGATTTACGG
TTCATGTCCTTGGAGCTTCTGCCAATGAGACCGTGTGAAGATATTGACATCACCATCTTTGGGTCTGGTTTGATGACTGAAGATGACGGGGCAGGATTCT
GCCTTGAAGACGATCCTGATCAATCTTCTTCAGGTGCTTCGGCTTCGCCTGATGAGATGGGATTGCCAATTTTTTTGAGTGCTATAAAGGAATGGATGAT
TGAGTTCGGGGCTTCCATGGTTTTCGTATCAATTCGCACAGATATGGCCTGGTACAGACTTGGGAAGCCGTCAGCACAGTATGCACCATGGTTTAAGATT
GTTCTCAAGACAGCAAAACTCGGAATTAGCATTATCTCATTATTAAAGGAGCAAAGTCGAGTATCCAAACTTTCTTTTGCAGAAGTAGTTAAGAAAGTTT
CGGAGTTCAAGAAGGATAACGATGGCTACATTTCCTCTGACGTGGCCGTTGCTGAGAGATTTGCTGTTGTGCATGGGCAGATACTAGTGCAACTCTTTGC
TGAATTCCCTGATGCTAAAATCAAGAAAAGTGCATTTGTTACTGGCCTTACAAACAAAATGGAAGAGCGTAAACATACAAAATGGGTTGGTGGGAACAAG
AAGCAACTGATTTTGCAGAGGGGCCAAGCTAATTTGAACCCTCAAGCTAACATGGGACCTGTAGTTTCCAAGAGGAAGGCTATGCAGGCTACGACAACAA
GATTGATTAACAGGATCTGGGGAGAATACTATTCGAATTACTCGCCGGAGGATTTGCAAGAGGAAACAAATGAGAAGGTAAAAGAGGAACATGTCGTTGA
GGAGCAAGAGGAAAATGAGGATGATGCCGAAGAGGAGAAAGAGGAGACAATAGTTGTTCCAGAGGAAGTGCAAAAACCATCCCCTGGATCAGTGAAAGTC
CGGTCATCATGCAAGTCGAATGTGGAAGCAAGATGGGAAGGTAATCCCGGAAAGGTAATTCTTGGTAAAACTATTTACAGTCGAGCTATCATCCATGGAG
AGACCATTGAGGCTGGGAGTGCTGTCATGGTTGATGAAGATGATGATGATGACCTCCCTGCCATCTACTTCATAGAGTACATGTATGAAACGAAAAATGG
TGCCAAAATGGCTCATGGAAGAATGATGCAACGTGGTTCTGAGACTGTTCTTGGAAATGCATCCAATGAGAGAGAGGTCTTCTTAACAACCAACTGCTCA
GATTTCCAACTGCAGGATGTTAAGCAGCTCGTTGTTGTCGAAATCCGGAAAATGCCGTGGGGTTATTATCATAGGAAGGATCATTTGAATGCTGGCAGAC
AGGATAGAGCAAGGGCGGAAGAAAGGAAAAAGAAAGGATTGCCTTTGGAGTATTACTGTAAAAGTTTGTATTGGCCGGAGAAGGGTGCTTTCTTTAGTCT
TCCATCTGATGGCATGGGTCTTGGTTCTGGTATTTGCCAGTCGTGTAAGATTAATGAGGCTGATGAGGAGAAGTATACATTTAAAGTGAATTCTTCAGGC
ACAGGATTCTCGTGGGAAGGAGCTGAGTACTCCATCGACGACTTTGTTTATGTTCAGCCGGAGAACTTTGCCTTACGAAGAAAGGAAACTGGAACTTTTA
AAGGTGGAAGGAATGTTGGTCTAGAAGCTTATGCTATATGCCAATTGGTGCAGATCATTGCAGGAAAGCAAGAGATCGAAGAATCTACTCAGGTTGAAGT
CCGAAGATTCTTCAGACCAGAGGACATTGCTTCTGAGAAAGCATATGGTTCTGATATCCGCGAGATATACTACAGCGAGGAAACGCATACTTTACCTGTT
GAGATGATAGAAGGGAAATGTGAAGTCAGAAAGAAAATCGACGTTCCTGCTTGTAACGTTACCACAGTTTTGGATCATATCTTCTTTTGTGAGCGTCTCT
ACGATCCTTCTAATGGTTCACTTCGGCAGTTGCCAGCTAACACCAATCTAAGGTATTCCAGCCTTGGTGGCAATGGCGATGTTGCTTCTAGGAAGAGGAA
GGGTAAAGGTAAAGAGGTTGAAGTTGATGAAGCAGTGGAGAAACCAAAGGAAGAATCTCAAGGAAACCTTTTGGCCACATTGGATATTTTTGCTGGCTGT
GGTGGCTTATCAGAAGGGCTACGTCAAGCTGGTGTGTCGGTTACGAAGTGGGCTATCGAGTATGAGGAACCAGCTGGTGAAGCATTCAGATTGAACCATC
CTGAATCGCTGGTCTTCGTCAACAACTGTAATGTGATTCTGAAGGCTGTCATGGAGAAATGCGGTGATACAGATGACTGCATCTCAACTTCAGAGGCTGC
TGAATTGGCTGCTGCACTTGGTGAGAATGTCATCAAAGATTTACCTATGCCAGGACAAGTGGAATTCATTAACGGAGGGCCTCCATGTCAGGGATTCTCC
GGGATGAACAGGTTTAATCACAGCACATGGAGTAAGGTTCAATGCGAGATGATCTTAGCATTCCTATCCTTTGCCGATTACTTCAGACCTAAGTACTTCC
TCCTGGAGAATGTGAGGAACTTCGTCTCCTTCAACAAAGGACAGACCTTTCGCTTAACCTTGGCTTCGCTTCTCGAAATGGGTTATCAGGTGAGATTTGG
CATCTTAGAAGCTGGAACGTTTGGCGTTTCACAGTCACGGAAGCGTGCTTTCATATGGGCAGCTGCACCAGACGAGGCTCTCCCTGAATGGCCAGAGCCT
ATGTACGTCTTCGGCGGACCGGAATTGAAAGTCTCTCTGTCTAGCAAACTGCAGTATGCTGCTGCGAGAAGCACTGCTGGCGGAGCTCCTTTCCGAGCTG
TAACTGTCCGAGACACGATCGGAGATCTCCCTGCCGTGACCAATGGTGCCTCCAAAGCAACTCTAGAGTATTCGAATGATCCCGTGTCCTGGTACCAGAA
ACAAATCCGAACCAACATGGCTGTTCTCACCGATCACATATCTAAAGAAATGAACGAGCTGAATCTCATCCGGTGCAAGAAGATTCCGAAGCGGCCTGGT
GCTGATTGGCGTGATCTCCCAGAAGAAACAGTTACATTGTCGAACGGGCAGAAGGCCGACCTAATCCCATGGTGCTTGCCAAACACGGCAGCTCGCCACA
ACCAGTGGAAGGGACTGTTCGGGAGGCTGGACTGGGAAGGCAACTTCCCAACCTCCATCACCGATCCTCAACCCATGGGCAAAGTCGGGTTGTGCTTCCA
TCCTGATCAGGACAGGATCGTTACGGTCCGCGAGTGTGCTCGATCTCAAGGATTCCCTGATAGCTATCAGTTCGCTGGTAACATCATTAGCAAGCATCGT
CAGATTGGAAATGCAGTTCCTCCGCCGTTGGCTTATGCGTTGGGAAGGAAGCTGAAGGAAGCCGTTGATAGTAAGAAGGTGAAAGAGTTAGAGAGTTATG
AGTAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10019220 pacid=23141417 polypeptide=Lus10019220 locus=Lus10019220.g ID=Lus10019220.BGIv1.0 annot-version=v1.0
MGSASKLISTAPDPEAEDPSLAESGKGAIPSDSKSRKAVASTKGKKKRETAKSIEPRASPPKRPRRAAACVNLKEKSVKLTDEYTVIEPKKEQVVQDELL
AVRLTEGQGEDGRPNRRLTDFIVHDADFAPQYLEMIEVSDMFISGVILPIDDCMGDDKEKRVRCGYRKFYDRFFEKARACIEVYKKLSKFAGGNPDLSLD
ELLAGLVRSFGSSKCFSGASSIKEFLISQGEFIYDQLKGLDETSKQNDMKFSELPVLLALRDKSKKHESFILEGAVGYGGSGTLVIEPNADKKDTKMDEV
NGSAGNDEDEDAKLARMLQQEELWASMKQKKRKGSSMSNAIYIKVNERDIANDYPLPAYYKQTVEETDEYIAVDCDDVINPDDLPRKVLHNWSLYDSDLR
FMSLELLPMRPCEDIDITIFGSGLMTEDDGAGFCLEDDPDQSSSGASASPDEMGLPIFLSAIKEWMIEFGASMVFVSIRTDMAWYRLGKPSAQYAPWFKI
VLKTAKLGISIISLLKEQSRVSKLSFAEVVKKVSEFKKDNDGYISSDVAVAERFAVVHGQILVQLFAEFPDAKIKKSAFVTGLTNKMEERKHTKWVGGNK
KQLILQRGQANLNPQANMGPVVSKRKAMQATTTRLINRIWGEYYSNYSPEDLQEETNEKVKEEHVVEEQEENEDDAEEEKEETIVVPEEVQKPSPGSVKV
RSSCKSNVEARWEGNPGKVILGKTIYSRAIIHGETIEAGSAVMVDEDDDDDLPAIYFIEYMYETKNGAKMAHGRMMQRGSETVLGNASNEREVFLTTNCS
DFQLQDVKQLVVVEIRKMPWGYYHRKDHLNAGRQDRARAEERKKKGLPLEYYCKSLYWPEKGAFFSLPSDGMGLGSGICQSCKINEADEEKYTFKVNSSG
TGFSWEGAEYSIDDFVYVQPENFALRRKETGTFKGGRNVGLEAYAICQLVQIIAGKQEIEESTQVEVRRFFRPEDIASEKAYGSDIREIYYSEETHTLPV
EMIEGKCEVRKKIDVPACNVTTVLDHIFFCERLYDPSNGSLRQLPANTNLRYSSLGGNGDVASRKRKGKGKEVEVDEAVEKPKEESQGNLLATLDIFAGC
GGLSEGLRQAGVSVTKWAIEYEEPAGEAFRLNHPESLVFVNNCNVILKAVMEKCGDTDDCISTSEAAELAAALGENVIKDLPMPGQVEFINGGPPCQGFS
GMNRFNHSTWSKVQCEMILAFLSFADYFRPKYFLLENVRNFVSFNKGQTFRLTLASLLEMGYQVRFGILEAGTFGVSQSRKRAFIWAAAPDEALPEWPEP
MYVFGGPELKVSLSSKLQYAAARSTAGGAPFRAVTVRDTIGDLPAVTNGASKATLEYSNDPVSWYQKQIRTNMAVLTDHISKEMNELNLIRCKKIPKRPG
ADWRDLPEETVTLSNGQKADLIPWCLPNTAARHNQWKGLFGRLDWEGNFPTSITDPQPMGKVGLCFHPDQDRIVTVRECARSQGFPDSYQFAGNIISKHR
QIGNAVPPPLAYALGRKLKEAVDSKKVKELESYE
|
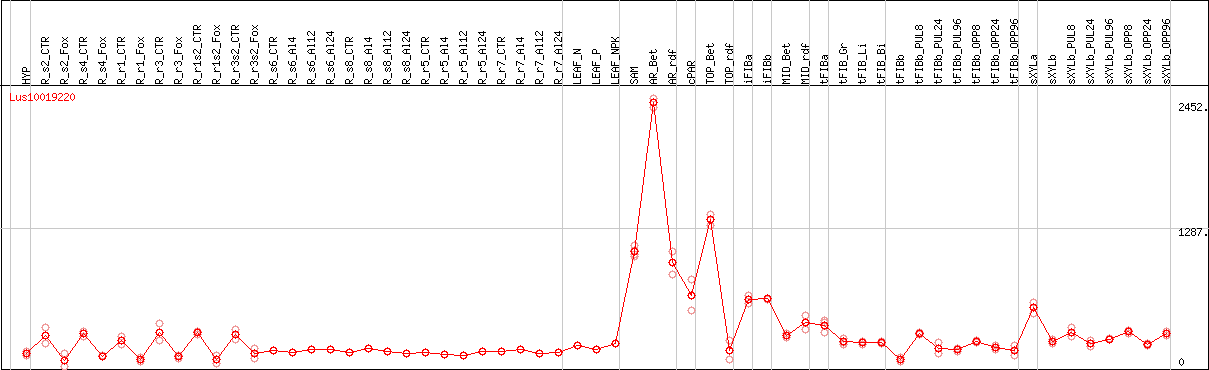
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10019220 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
